Abstract
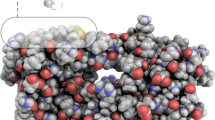
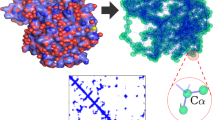
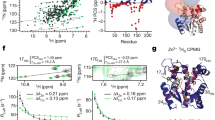
A unique feature of chemical catalysis mediated by enzymes is that the catalytically reactive atoms are embedded within a folded protein. Although current understanding of enzyme function has been focused on the chemical reactions and static three-dimensional structures, the dynamic nature of proteins has been proposed to have a function in catalysis1,2,3,4,5. The concept of conformational substates has been described6; however, the challenge is to unravel the intimate linkage between protein flexibility and enzymatic function. Here we show that the intrinsic plasticity of the protein is a key characteristic of catalysis. The dynamics of the prolyl cis–trans isomerase cyclophilin A (CypA) in its substrate-free state and during catalysis were characterized with NMR relaxation experiments. The characteristic enzyme motions detected during catalysis are already present in the free enzyme with frequencies corresponding to the catalytic turnover rates. This correlation suggests that the protein motions necessary for catalysis are an intrinsic property of the enzyme and may even limit the overall turnover rate. Motion is localized not only to the active site but also to a wider dynamic network. Whereas coupled networks in proteins have been proposed previously3,7,8,9,10, we experimentally measured the collective nature of motions with the use of mutant forms of CypA. We propose that the pre-existence of collective dynamics in enzymes before catalysis is a common feature of biocatalysts and that proteins have evolved under synergistic pressure between structure and dynamics.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Jencks, W. P. Catalysis in Chemistry and Enzymology (Dover, New York, 1987)
Hammes, G. Multiple conformational changes in enzyme catalysis. Biochemistry 41, 8221–8228 (2002)
Benkovic, S. J. & Hammes-Schiffer, S. A perspective on enzyme catalysis. Science 301, 1196–1202 (2003)
Garcia-Viloca, M., Gao, J., Karplus, M. & Truhlar, D. G. How enzymes work: analysis by modern rate theory and computer simulations. Science 303, 186–195 (2004)
Eisenmesser, E. Z., Akke, M., Bosco, D. A. & Kern, D. Enzyme dynamics during catalysis. Science 295, 1520–1523 (2002)
Austin, R. H. et al. Dynamics of ligand binding to myoglobin. Biochemistry 14, 5355–5373 (1975)
Berendsen, H. J. C. & Hayward, S. Collective protein dynamics in relation to function. Curr. Opin. Struct. Biol. 10, 165–169 (2000)
Rod, T. H., Radkiewicz, J. L. & Brooks, C. L. Correlated motion and the effect of distal mutations in dihydrofolate reductase. Proc. Natl Acad. Sci. USA 100, 6980–6985 (2003)
Suel, G. M., Lockless, S. W., Wall, M. A. & Ranganathan, R. Evolutionarily conserved networks of residues mediate allosteric communication in proteins. Nature Struct. Biol. 10, 59–69 (2003)
Agarwal, P. K., Geist, A. & Gorin, A. Protein dynamics and enzymatic catalysis: Investigating the peptidyl-prolyl cis-trans isomerization activity of cyclophilin A. Biochemistry 43, 10605–10618 (2004)
Schmid, F. X. Prolyl isomerases. Adv. Protein Chem. 59, 243–282 (2001)
Goff, S. P. Genetic control of retrovirus susceptibility in mammalian cells. Annu. Rev. Genet. 38, 61–85 (2004)
Mulder, F. A. A., Mittermaier, A., Hon, B., Kahlquist, F. W. & Kay, L. E. Studying excited states of proteins by NMR spectroscopy. Nature Struct. Biol. 8, 932–935 (2001)
Palmer, A. G. NMR characterization of the dynamics of biomacromolecules. Chem. Rev. 104, 3623–3640 (2004)
Kern, D., Kern, G., Scherer, G., Fischer, G. & Drakenberg, T. Kinetic analysis of cyclophilin-catalyzed prolyl cis/trans isomerization by dynamic NMR spectroscopy. Biochemistry 34, 13594–13602 (1995)
Zhao, Y. & Ke, H. Crystal structure implies that cyclophilin predominantly catalyzes the trans to cis isomerization. Biochemistry 35, 7356–7361 (1996)
Fischer, G., Bang, H. & Mech, C. Determination of enzymatic catalysis for the cis-trans-isomerization of peptide binding in proline-containing peptides. Biomed. Biochim. Acta 43, 1101–1111 (1984)
Ottiger, M., Zerbe, O., Guntert, P. & Wuthrich, K. The NMR solution conformation of unligated human cyclophilin A. J. Mol. Biol. 272, 64–81 (1997)
Skrynnikov, N. R., Dahlquist, F. W. & Kay, L. E. Reconstructing NMR spectra of ‘invisible’ excited protein states using HSQC and HMQC experiments. J. Am. Chem. Soc. 124, 12352–12360 (2002)
Wishart, D. S., Sykes, B. D. & Richards, F. M. Relationship between nuclear magnetic resonance chemical shift and protein secondary structure. J. Mol. Biol. 222, 311–333 (1991)
Rozovsky, S., Jogl, G., Tong, L. & McKermott, A. E. Solution-state NMR investigations of triosephosphate isomerase active site loop motion: ligand release in relation to active site loop dynamics. J. Mol. Biol. 310, 271–280 (2001)
Williams, J. C. & McDermott, A. E. Dynamics of the flexible loop of triosephosphate isomerase: the loop motion is not ligand gated. Biochemistry 34, 8309–8319 (1995)
Schnell, J. R., Dyson, H. J. & Wright, P. E. Structure, dynamics, and catalytic function of dihydrofolate reductase. Annu. Rev. Biophys. Biomol. Struct. 33, 119–140 (2004)
McElheny, D., Schnell, J. R., Lansing, J. C., Dyson, H. J. & Wright, P. E. Defining the role of active-site loop fluctuations in dihydrofolate reductase catalysis. Proc. Natl Acad. Sci. USA 102, 5032–5037 (2005)
Ishima, R., Freedberg, D. I., Wang, Y. X., Louis, J. M. & Torchia, D. A. Flap opening and dimer-interface flexibility in the free and inhibitor-bound HIV protease, and their implications for function. Struct. Fold. Des. 7, 1047–1055 (1999)
Cole, R. & Loria, J. P. Evidence for flexibility in the function of ribonuclease A. Biochemistry 41, 6072–6081 (2002)
Volkman, B. F., Lipson, D., Wemmer, D. E. & Kern, D. Two-state allosteric behaviour in a single-domain signalling protein. Science 291, 2429–2433 (2001)
Rosen, M. K. et al. Selective methyl group protonation of perdeuterated proteins. J. Mol. Biol. 263, 627–636 (1996)
Loria, J. P., Rance, M. & Palmer, A. G. A TROSY CPMG sequence for characterizing chemical exchange in large proteins. J. Biomol. NMR 15, 151–155 (1999)
Carver, J. P. & Richards, R. E. A general two-site solution for the chemical exchange produced dependence of T2 upon the Carr-Purcell pulse separation. J. Magn. Reson. 6, 89–105 (1972)
Acknowledgements
We thank K. H. Wildman for discussions. This work was supported by NIH grants to D.K., by a grant from the Canadian Institutes of Health Research to L.E.K., and by a grant from the Swedish Research Council to M.W.W. Part of the NMR studies was performed at the NHMFL at Florida with support from the NSF.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Figure 1
Concentration dependence of relaxation dispersion data for CypA. (PDF 216 kb)
Supplementary Figure 2
Quantitative analysis of protein dynamics of free CypA at 25°C. (PDF 197 kb)
Supplementary Figure 3
Pre-existing motions within the active site of CypA. (PDF 210 kb)
Supplementary Figure 4
Backbone amide chemical shift differences of CypA mutants. (PDF 258 kb)
Supplementary Figure Legends
Text to accompany the above Supplementary Figures. (DOC 30 kb)
Supplementary Tables
Supplementary Tables 1–4. (DOC 157 kb)
Supplementary Methods
Additional descriptions of methods used in this study. (DOC 29 kb)
Rights and permissions
About this article
Cite this article
Eisenmesser, E., Millet, O., Labeikovsky, W. et al. Intrinsic dynamics of an enzyme underlies catalysis. Nature 438, 117–121 (2005). https://doi.org/10.1038/nature04105
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature04105
This article is cited by
-
Acceleration of enzymatic catalysis by active hydrodynamic fluctuations
Communications Physics (2022)
-
Functional control of a 0.5 MDa TET aminopeptidase by a flexible loop revealed by MAS NMR
Nature Communications (2022)
-
Cross-scale analysis of temperature compensation in the cyanobacterial circadian clock system
Communications Physics (2022)
-
Listening to the sound of a bacterium
Nature Nanotechnology (2020)
-
Structure-based designing efficient peptides based on p53 binding site residues to disrupt p53-MDM2/X interaction
Scientific Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.