Abstract
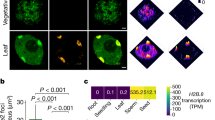
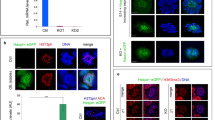
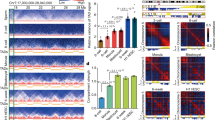
In sexually reproducing animals, a crucial step in zygote formation is the decondensation of the fertilizing sperm nucleus into a DNA replication-competent male pronucleus. Genome-wide nucleosome assembly on paternal DNA implies the replacement of sperm chromosomal proteins, such as protamines, by maternally provided histones1,2. This fundamental process is specifically impaired in sésame (ssm), a unique Drosophila maternal effect mutant that prevents male pronucleus formation3. Here we show that ssm is a point mutation in the Hira gene, thus demonstrating that the histone chaperone protein HIRA is required for nucleosome assembly during sperm nucleus decondensation. In vertebrates, HIRA has recently been shown to be critical for a nucleosome assembly pathway independent of DNA synthesis that specifically involves the H3.3 histone variant4,5. We also show that nucleosomes containing H3.3, and not H3, are specifically assembled in paternal Drosophila chromatin before the first round of DNA replication. The exclusive marking of paternal chromosomes with H3.3 represents a primary epigenetic distinction between parental genomes in the zygote, and underlines an important consequence of the critical and highly specialized function of HIRA at fertilization.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Wright, S. J. Sperm nuclear activation during fertilization. Curr. Top. Dev. Biol. 46, 133–178 (1999)
McLay, D. W. & Clarke, H. J. Remodelling the paternal chromatin at fertilization in mammals. Reproduction 125, 625–633 (2003)
Loppin, B., Docquier, M., Bonneton, F. & Couble, P. The maternal effect mutation sésame affects the formation of the male pronucleus in Drosophila melanogaster. Dev. Biol. 222, 392–404 (2000)
Ray-Gallet, D. et al. HIRA is critical for a nucleosome assembly pathway independent of DNA synthesis. Mol. Cell 9, 1091–1100 (2002)
Tagami, H., Ray-Gallet, D., Almouzni, G. & Nakatani, Y. Histone H3.1 and H3.3 complexes mediate nucleosome assembly pathways dependent or independent of DNA synthesis. Cell 116, 51–61 (2004)
Loppin, B., Berger, F. & Couble, P. The Drosophila maternal gene sésame is required for sperm chromatin remodeling at fertilization. Chromosoma 110, 430–440 (2001)
Jayaramaiah Raja, S. & Renkawitz-Pohl, R. Replacement by Drosophila melanogaster protamines and Mst77F of histones during chromatin condensation in late spermatids and role of Sésame in the removal of these proteins from the male pronucleus. Mol. Cell. Biol. 25, 6165–6177 (2005)
Loppin, B. & Karr, T. L. in Comprehensive Molecular Insect Science (eds Gilbert, L. B. & Iatrou, K.) 213–236 (Elsevier, Oxford, 2004)
Llevadot, R. et al. Cloning, chromosome mapping and expression analysis of the HIRA gene from Drosophila melanogaster. Biochem. Biophys. Res. Commun. 249, 486–491 (1998)
Kirov, N., Shtilbans, A. & Rushlow, C. Isolation and characterization of a new gene encoding a member of the HIRA family of proteins from Drosophila melanogaster. Gene 212, 323–332 (1998)
Smith, T. F., Gaitatzes, C., Saxena, K. & Neer, E. J. The WD repeat: a common architecture for diverse functions. Trends Biochem. Sci. 24, 181–185 (1999)
Foe, V. E., Odell, G. M. & Edgar, B. A. in The Development of Drosophila melanogaster (eds Bates, M. & Martinez Arias, A.) 149–300 (Cold Spring Harbor Press, Cold Spring Harbor, New York, 1993)
Ahmad, K. & Henikoff, S. Histone H3 variants specify modes of chromatin assembly. Proc. Natl Acad. Sci. USA 99 (suppl.), 16477–16484 (2002)
Ahmad, K. & Henikoff, S. The histone variant H3.3 marks active chromatin by replication-independent nucleosome assembly. Mol. Cell 9, 1191–1200 (2002)
Janicki, S. M. et al. From silencing to gene expression: real-time analysis in single cells. Cell 116, 683–698 (2004)
Schwartz, B. E. & Ahmad, K. Transcriptional activation triggers deposition and removal of the histone variant H3.3. Genes Dev. 19, 804–814 (2005)
Henikoff, S., Furuyama, T. & Ahmad, K. Histone variants, nucleosome assembly and epigenetic inheritance. Trends Genet. 20, 320–326 (2004)
Akhmanova, A. S. et al. Structure and expression of histone H3.3 genes in Drosophila melanogaster and Drosophila hydei. Genome 38, 586–600 (1995)
Yamaguchi, M., Date, T. & Matsukage, A. Distribution of PCNA in Drosophila embryo during nuclear division cycles. J. Cell Sci. 100, 729–733 (1991)
Roberts, C. et al. Targeted mutagenesis of the Hira gene results in gastrulation defects and patterning abnormalities of mesoendodermal derivatives prior to early embryonic lethality. Mol. Cell. Biol. 22, 2318–2328 (2002)
van der Heijden, G. W. et al. Asymmetry in Histone H3 variants and lysine methylation between paternal and maternal chromatin of the early mouse zygote. Mech. Dev. 122, 1008–1022 (2005)
Cowell, I. G. et al. Heterochromatin, HP1 and methylation at lysine 9 of histone H3 in animals. Chromosoma 111, 22–36 (2002)
Arney, K. L., Bao, S., Bannister, A. J., Kouzarides, T. & Surani, M. A. Histone methylation defines epigenetic asymmetry in the mouse zygote. Int. J. Dev. Biol. 46, 317–320 (2002)
Lepikhov, K. & Walter, J. Differential dynamics of histone H3 methylation at positions K4 and K9 in the mouse zygote. BMC Dev. Biol. 4, 12 (2004)
Mayer, W., Niveleau, A., Walter, J., Fundele, R. & Haaf, T. Demethylation of the zygotic paternal genome. Nature 403, 501–502 (2000)
Huynh, K. D. & Lee, J. T. Inheritance of a pre-inactivated paternal X chromosome in early mouse embryos. Nature 426, 857–862 (2003)
Okamoto, I., Otte, A. P., Allis, C. D., Reinberg, D. & Heard, E. Epigenetic dynamics of imprinted X inactivation during early mouse development. Science 303, 644–649 (2004)
Klemenz, R., Weber, U. & Gehring, W. J. The white gene as a marker in a new P-element vector for gene transfer in Drosophila. Nucleic Acids Res. 15, 3947–3959 (1987)
Drysdale, R. A., Crosby, M. A. & The FlyBase Consortium. FlyBase: genes and gene models. Nucleic Acids Res. 33 (database issue), D390–D395 (2005).
Acknowledgements
We thank P. Fisher for anti-PCNA antibodies and the European Drosophila Genome Project for cosmid clones. We are grateful to B. Durand and B. Horard for helpful discussions. We also thank J. Schmitt and the CTµ microscopy center for technical assistance. This work was supported by the C.N.R.S. and the French Ministry of Research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Figure 1
RT-PCR analysis of Hira expression (PDF 735 kb)
Supplementary Figure 2
Western Blot analysis of HIRA-FLAG, H3.3-FLAG and H3-FLAG proteins. (PDF 283 kb)
Supplementary Figure 3
Pronuclei replicate their DNA by the time they appose. (PDF 532 kb)
Supplementary Table 1
Complementation of the ssm185b phenotype with Hira transgenes. (PDF 783 kb)
Rights and permissions
About this article
Cite this article
Loppin, B., Bonnefoy, E., Anselme, C. et al. The histone H3.3 chaperone HIRA is essential for chromatin assembly in the male pronucleus. Nature 437, 1386–1390 (2005). https://doi.org/10.1038/nature04059
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature04059
This article is cited by
-
Insights into epigenetic patterns in mammalian early embryos
Protein & Cell (2021)
-
Rad9a is involved in chromatin decondensation and post-zygotic embryo development in mice
Cell Death & Differentiation (2019)
-
Nuclear formation induced by DNA-conjugated beads in living fertilised mouse egg
Scientific Reports (2019)
-
Chromatin-mediated regulators of meiotic recombination revealed by proteomics of a recombination hotspot
Epigenetics & Chromatin (2018)
-
ASF1 is required to load histones on the HIRA complex in preparation of paternal chromatin assembly at fertilization
Epigenetics & Chromatin (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.