Abstract
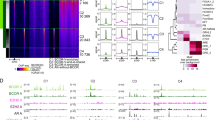
Gene regulation in eukaryotes requires the coordinate interaction of chromatin-modulating proteins with specific transcription factors such as the androgen receptor1. Gene activation and repression is specifically regulated by histone methylation status at distinct lysine residues2. Here we show that lysine-specific demethylase 1 (LSD1; also known as BHC110)3 co-localizes with the androgen receptor in normal human prostate and prostate tumour. LSD1 interacts with androgen receptor in vitro and in vivo, and stimulates androgen-receptor-dependent transcription. Conversely, knockdown of LSD1 protein levels abrogates androgen-induced transcriptional activation and cell proliferation. Chromatin immunoprecipitation analyses demonstrate that androgen receptor and LSD1 form chromatin-associated complexes in a ligand-dependent manner. LSD1 relieves repressive histone marks by demethylation of histone H3 at lysine 9 (H3-K9), thereby leading to de-repression of androgen receptor target genes. Furthermore, we identify pargyline as an inhibitor of LSD1. Pargyline blocks demethylation of H3-K9 by LSD1 and consequently androgen-receptor-dependent transcription. Thus, modulation of LSD1 activity offers a new strategy to regulate androgen receptor functions. Here, we link demethylation of a repressive histone mark with androgen-receptor-dependent gene activation, thus providing a mechanism by which demethylases control specific gene expression.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Glass, C. K. & Rosenfeld, M. G. The coregulator exchange in transcriptional function of nuclear receptors. Genes Dev. 14, 121–141 (2000)
Strahl, B. D. & Allis, C. D. The language of covalent histone modifications. Nature 403, 41–45 (2000)
Shi, Y. et al. Histone demethylation mediated by the nuclear amine oxidase homolog LSD1. Cell 119, 941–953 (2004)
Shi, Y. et al. Coordinated histone modifications mediated by a CtBP co-repressor complex. Nature 422, 735–738 (2003)
Hakimi, M. A. et al. A candidate X-linked mental retardation gene is a component of a new family of histone deacetylase-containing complexes. J. Biol. Chem. 278, 7234–7239 (2003)
Hakimi, M. A. et al. A core-BRAF35 complex containing histone deacetylase mediates repression of neuronal-specific genes. Proc. Natl Acad. Sci. USA 99, 7420–7425 (2002)
Eimer, S. et al. Loss of spr-5 bypasses the requirement for the C. elegans presenilin sel-12 by derepressing hop-1. EMBO J. 21, 5787–5796 (2002)
Müller, J. M. et al. FHL2, a novel tissue-specific coactivator of the androgen receptor. EMBO J. 19, 359–369 (2000)
Müller, J. M. et al. The transcriptional coactivator FHL2 transmits Rho signals from the cell membrane into the nucleus. EMBO J. 21, 736–748 (2002)
Shang, Y., Myers, M. & Brown, M. Formation of the androgen receptor transcription complex. Mol. Cell 9, 601–610 (2002)
Shatkina, L. et al. The cochaperone Bag-1L enhances androgen receptor action via interaction with the NH2-terminal region of the receptor. Mol. Cell. Biol. 23, 7189–7197 (2003)
Kang, Z., Pirskanen, A., Jänne, O. A. & Palvimo, J. J. Involvement of proteasome in the dynamic assembly of the androgen receptor transcription complex. J. Biol. Chem. 277, 48366–48371 (2002)
Metzger, E. et al. A novel inducible transactivation domain in the androgen receptor: implications for PRK in prostate cancer. EMBO J. 22, 270–280 (2003)
Wiznerowicz, M. & Trono, D. Conditional suppression of cellular genes: lentivirus vector-mediated drug-inducible RNA interference. J. Virol. 77, 8957–8961 (2003)
Pfaffl, M. W., Horgan, G. W. & Dempfle, L. Relative expression software tool (REST) for group-wise comparison and statistical analysis of relative expression results in real-time PCR. Nucleic Acids Res. 30, e36 (2002)
O'Neill, T. E., Roberge, M. & Bradbury, E. M. Nucleosome arrays inhibit both initiation and elongation of transcripts by bacteriophage T7 RNA polymerase. J. Mol. Biol. 223, 67–78 (1992)
Acknowledgements
We thank T. Benzing, F. Claessens, T. Jenuwein, Z. Sun and D. Trono for providing reagents. We are obliged to the members of the Schüle laboratory for discussions. We thank K. Fischer, P. Kahl and L. Heukamp for technical assistance. This work was supported by grants from the Deutsche Forschungsgemeinschaft and Deutsche Krebshilfe to R.S.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Notes
Contains Supplementary Methods (plasmids, immunofluorescence, in vitro pull-down assays, mRNAanalyses, tandem affinity purification) and the legends for the Supplementary Figures. (DOC 41 kb)
Supplementary Figure S1
a, Coomassie blue staining reveals that LSD1 (arrow) is co-purified with TAP-FHL2 (arrow) during tandem affinity purification. b, Western blot analysis using -LSD1 antibody. (PDF 113 kb)
Supplementary Figure S2
LSD1 expression analyses. (PDF 88 kb)
Supplementary Figure S3
LSD1 interacts with chromatin. (PDF 82 kb)
Supplementary Figure S4
LNCaP cells were incubated with or without R1881 and treated with or without pargyline. (PDF 229 kb)
Supplementary Figure S5
siRNA mediated knockdown of LSD1. (PDF 52 kb)
Supplementary Figure S6a-h
Specificity of LSD1 in the control of AR-induced transcriptional activity. (PDF 15 kb)
Supplementary Figure S6i-k
Specificity of LSD1 in the control of AR-induced transcriptional activity. (PDF 8 kb)
Rights and permissions
About this article
Cite this article
Metzger, E., Wissmann, M., Yin, N. et al. LSD1 demethylates repressive histone marks to promote androgen-receptor-dependent transcription. Nature 437, 436–439 (2005). https://doi.org/10.1038/nature04020
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature04020
This article is cited by
-
Kdm1a safeguards the topological boundaries of PRC2-repressed genes and prevents aging-related euchromatinization in neurons
Nature Communications (2024)
-
LSD1 modulates the bone metastasis of breast cancer cells through hnRNPA2B1-mediated sorting of exosomal miRNAs
Cell Death Discovery (2024)
-
Exploring epigenetic strategies for the treatment of osteoporosis
Molecular Biology Reports (2024)
-
Lysin (K)-specific demethylase 1 inhibition enhances proteasome inhibitor response and overcomes drug resistance in multiple myeloma
Experimental Hematology & Oncology (2023)
-
LSD1: an emerging face in altering the tumor microenvironment and enhancing immune checkpoint therapy
Journal of Biomedical Science (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.