Abstract
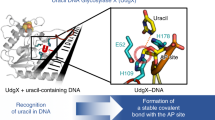
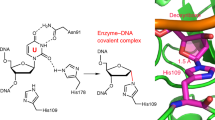
Members of the small ubiquitin-like modifier (SUMO) family can be covalently attached to the lysine residue of a target protein through an enzymatic pathway similar to that used in ubiquitin conjugation1, and are involved in various cellular events that do not rely on degradative signalling via the proteasome or lysosome2,3,4,5. However, little is known about the molecular mechanisms of SUMO-modification-induced protein functional transfer. During DNA mismatch repair, SUMO conjugation of the uracil/thymine DNA glycosylase TDG promotes the release of TDG from the abasic (AP) site created after base excision, and coordinates its transfer to AP endonuclease 1, which catalyses the next step in the repair pathway6. Here we report the crystal structure of the central region of human TDG conjugated to SUMO-1 at 2.1 Å resolution. The structure reveals a helix protruding from the protein surface, which presumably interferes with the product DNA and thus promotes the dissociation of TDG from the DNA molecule. This helix is formed by covalent and non-covalent contacts between TDG and SUMO-1. The non-covalent contacts are also essential for release from the product DNA, as verified by mutagenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Johnson, E. S. Protein modification by SUMO. Annu. Rev. Biochem. 73, 355–382 (2004)
Gill, G. SUMO and ubiquitin in the nucleus: different functions, similar mechanisms? Genes Dev. 18, 2046–2059 (2004)
Seeler, J. S. & Dejean, A. Nuclear and unclear functions of SUMO. Nature Rev. Mol. Cell Biol. 4, 690–699 (2003)
Kim, K. I., Baek, S. H. & Chung, C. H. Versatile protein tag, SUMO: its enzymology and biological function. J. Cell. Physiol. 191, 257–268 (2002)
Mahajan, R., Delphin, C., Guan, T., Gerace, L. & Melchior, F. A small ubiquitin-related polypeptide involved in targeting RanGAP1 to nuclear pore complex protein RanBP2. Cell 88, 97–107 (1997)
Hardeland, U. et al. Thymine DNA glycosylase. Prog. Nucleic Acid Res. Mol. Biol. 68, 235–253 (2001)
Waters, T. R. & Swann, P. F. Kinetics of the action of thymine DNA glycosylase. J. Biol. Chem. 273, 20007–20014 (1998)
Waters, T. R., Gallinari, P., Jiricny, J. & Swann, P. F. Human thymine DNA glycosylase binds to apurinic sites in DNA but is displaced by human apurinic endonuclease 1. J. Biol. Chem. 274, 67–74 (1999)
Hardeland, U., Steinacher, R., Jiricny, J. & Schar, P. Modification of the human thymine-DNA glycosylase by ubiquitin-like proteins facilitates enzymatic turnover. EMBO J. 21, 1456–1464 (2002)
Barrett, T. E. et al. Crystal structure of a G:T/U mismatch-specific DNA glycosylase: mismatch recognition by complementary-strand interactions. Cell 92, 117–129 (1998)
Hardeland, U., Bentele, M., Jiricny, J. & Schar, P. Separating substrate recognition from base hydrolysis in human thymine DNA glycosylase by mutational analysis. J. Biol. Chem. 275, 33449–33456 (2000)
Roberts, R. J. On base flipping. Cell 82, 9–12 (1995)
Mol, C. D. et al. Crystal structure of human uracil-DNA glycosylase in complex with a protein inhibitor: protein mimicry of DNA. Cell 82, 701–708 (1995)
Mol, C. D. et al. Crystal structure and mutational analysis of human uracil-DNA glycosylase: structural basis for specificity and catalysis. Cell 80, 869–878 (1995)
Bayer, P. et al. Structure determination of the small ubiquitin-related modifier SUMO-1. J. Mol. Biol. 280, 275–286 (1998)
Song, J., Durrin, L. K., Wilkinson, T. A., Krontiris, T. G. & Chen, Y. Identification of a SUMO-binding motif that recognizes SUMO-modified proteins. Proc. Natl Acad. Sci. USA 101, 14373–14378 (2004)
Uchimura, Y., Nakamura, M., Sugasawa, K., Nakao, M. & Saitoh, H. Overproduction of eukaryotic SUMO-1- and SUMO-2-conjugated proteins in Escherichia coli. Anal. Biochem. 331, 204–206 (2004)
Leslie, A. G. Integration of macromolecular diffraction data. Acta Crystallogr. D 55, 1696–1702 (1999)
Collaborative Computational Project No. 4, The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Vagin, A. & Teplyakov, A. MOLREP: an automated program for molecular replacement. J. Appl. Crystallogr. 30, 1022–1025 (1997)
Mossessova, E. & Lima, C. D. Ulp1-SUMO crystal structure and genetic analysis reveal conserved interactions and a regulatory element essential for cell growth in yeast. Mol. Cell 5, 865–876 (2000)
Jones, T. A., Zou, J. Y., Cowan, S. W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991)
Brunger, A. T. et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
DeLano, W. L. PyMOLhttp://www.pymol.org (2002).
Barrett, T. E. et al. Crystal structure of a thwarted mismatch glycosylase DNA repair complex. EMBO J. 18, 6599–6609 (1999)
Acknowledgements
This work was supported by grants to M.S. from the Japanese Ministry of Education, Science, Sports and Culture, and Japan Science and Technology Agency.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Atomic coordinates of SUMO-1–TDG have been deposited in the Protein Data Bank under the accession number 1WYW. Reprints and permissions information is available at npg.nature.com/reprintsandpermissions. The authors declare no competing financial interests.
Supplementary information
Supplementary Figure S1
Electron density map of SUMO1-TDG (DOC 745 kb)
Supplementary Figure S2
Limited proteolysis of TDG and SUMO1-TDG (DOC 539 kb)
Supplementary Figure S3
DNA and SUMO-1 binding activity of the full-length TDG (DOC 1491 kb)
Supplementary Table S1
Data collection and model refinement statistics (DOC 45 kb)
Supplementary Methods
Protein expression and purification (DOC 26 kb)
Rights and permissions
About this article
Cite this article
Baba, D., Maita, N., Jee, JG. et al. Crystal structure of thymine DNA glycosylase conjugated to SUMO-1. Nature 435, 979–982 (2005). https://doi.org/10.1038/nature03634
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature03634
This article is cited by
-
An expanded lexicon for the ubiquitin code
Nature Reviews Molecular Cell Biology (2023)
-
CUL5-ARIH2 E3-E3 ubiquitin ligase structure reveals cullin-specific NEDD8 activation
Nature Chemical Biology (2021)
-
Characterization of thymine microcrystals by CARS and SHG microscopy
Scientific Reports (2020)
-
Two-way communications between ubiquitin-like modifiers and DNA
Nature Structural & Molecular Biology (2014)
-
Structure of the human ATG12~ATG5 conjugate required for LC3 lipidation in autophagy
Nature Structural & Molecular Biology (2013)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.