Abstract
F1-ATPase is the smallest known rotary motor, and it rotates in an anticlockwise direction as it hydrolyses ATP1,2,3,4,5. Single-molecule experiments6,7,8,9 point towards three catalytic events per turn, in agreement with the molecular structure of the complex10. The physiological function of F1 is ATP synthesis. In the ubiquitous F0F1 complex, this energetically uphill reaction is driven by F0, the partner motor of F1, which forces the backward (clockwise) rotation of F1, leading to ATP synthesis11,12,13. Here, we have devised an experiment combining single-molecule manipulation and microfabrication techniques to measure the yield of this mechanochemical transformation. Single F1 molecules were enclosed in femtolitre-sized hermetic chambers and rotated in a clockwise direction using magnetic tweezers. When the magnetic field was switched off, the F1 molecule underwent anticlockwise rotation at a speed proportional to the amount of synthesized ATP. At 10 Hz, the mechanochemical coupling efficiency was low for the α3β3γ subcomplex (F1-ɛ), but reached up to 77% after reconstitution with the ɛ-subunit (F1+ɛ). We provide here direct evidence that F1 is designed to tightly couple its catalytic reactions with the mechanical rotation. Our results suggest that the ɛ-subunit has an essential function during ATP synthesis.
Similar content being viewed by others
Main
Genetically engineered, single F1 molecules can be grafted on to a glass slide, and the ATP-driven anticlockwise rotation has been observed directly under an optical microscope by coupling the γ-subunit to a fluorescent filament or a submicrometre-sized plastic bead9,14. Our experiment used a similar procedure with some alternations: the magnetic bead was attached to the γ-subunit to rotate it relative to the glass-bound α3β3 stator part of the F1 molecule using magnetic tweezers15. We also developed a microfabrication technique to confine this system to a 6-fl transparent container16 (Fig. 1b). Owing to the extremely small volume, the number of ATP molecules produced by a single F1 enzyme reached a detectable micromolar concentration within minutes of forced clockwise rotation. After the magnetic field was switched off, this ATP concentration could be readily determined by monitoring the speed of the enzyme spontaneously rotating in an anticlockwise direction. By comparing the number of turns in both directions (mechanical input/output) with the changes in ATP concentration (chemical input/output), we were able to assess directly the mechanochemical coupling efficiencies of F1.
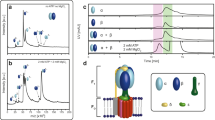
a, Schematic view of the membrane-embedded F0F1-ATP synthase in which the proton-driven F0 rotates F1 in a clockwise direction for ATP synthesis. Isolated F1 only hydrolyses ATP, and hence is called F1-ATPase. F1-ATPase comprises five different subunits, α3β3γδɛ, of which the α3β3γ subcomplex is the minimal functional part of the ATPase30. ATP hydrolysis occurs sequentially on the three β-subunits, which generate power strokes driving the anticlockwise rotation of γɛ (or γ if ɛ is absent). b, To force the ATP synthesis of F1 against the chemical potential, a magnetic bead was attached to the γ-subunit of an immobilized F1, and rotated clockwise using magnetic tweezers. A single bead–enzyme complex was enclosed in a PDMS container in order to trap synthesized ATP molecules. c, ATP synthesis was detectable via the enzyme itself: after the magnetic field was released, the enzyme resumed its ATP-hydrolysing anticlockwise rotation at a speed proportional to the ATP concentration in the chamber.
As a first step, we focused on the enzyme acting as a motor; that is, hydrolysing ATP. F1 molecules were immobilized on a glass plate at a density of less than one per 4 µm2 (the surface area of a chamber). A polydimethylsiloxane (PDMS) sheet with microchambers on its surface (Fig. 2a, left) was suspended above the glass plate and partially pressed down onto it with a small, flat-tipped glass needle to enclose a rotating F1 molecule in a chamber (Fig. 2b). The chamber size was large enough to avoid physical conflict with the rotating bead (Fig. 2a, right), and could contain only 1,800 ± 200 ATP molecules at 500 nM ATP. (The error number was derived from the error of the chamber volume, around 10%.) Because the motor consumed ATP molecules in the chamber, the rotation speed decreased (Fig. 2c). Under these conditions of low ATP concentration, where ATP binding is the rate-limiting step, the speed decreases linearly with ATP concentration and the expected first-order decay was observed (Fig. 2d). We re-opened the chamber when the speed approached zero, thus taking back the system to its initial condition (Fig. 2b). The enzyme immediately resumed its initial speed, showing that the motor was still active. Extrapolating the rotation speed versus the number of accumulated rotations (Fig. 2d, inset) yielded the maximum number of turns that the single enzyme could perform by exhausting the 1,800 available ATP molecules. We determined the value to be 580 ± 100 turns (mean ± s.d.), corresponding to a coupling ratio of 3.15 ± 0.5 ATPs per turn. A similar result was obtained for F1+ɛ (data not shown). Such a high coupling ratio in the motor direction had been proposed previously8,17, but only indirect evidence had been provided so far, because the mechanical output and the real ATP consumption could not be monitored simultaneously. In particular, the possibility that the motor might ‘slip’ and hydrolyse ATP without rotation was a lingering question. Back steps8 could also affect the coupling ratio; however, this was such a rare event (less than 1% in our case) that it can be considered to be negligible. The present experiment directly demonstrates that F1+ɛ and F1-ɛ do not waste ATP, and work as motors with nearly perfect mechanochemical coupling efficiencies.
a, Scanning electron microscopy image of a PDMS sheet with cylindrical cavities of 1.1-µm radius and 1.5-µm height (left), and a bright-field image under an optical microscope after enclosing a rotating bead (right), with successive images of rotation. b, Elastic properties of PDMS allow the reversible closing/opening of the chamber, by partially pressing a suspended sheet against the bottom glass plate. c, The chamber volume was small enough to ensure that a single enzyme exhausted most of the available ATP (1,800 molecules) within a few minutes. d, Time course of accumulated revolutions showing an exponential decay after closing the chamber, and the recovery of the initial velocity after opening the chamber. The inset shows the rotation speeds as a function of the accumulated revolutions for six different F1 molecules. The linear decreases were extrapolated to give the maximum number of possible turns and the coupling ratio between ATP hydrolysis and mechanical rotation.
In a second series of experiments, we turned to the more challenging question of ATP synthesis. The buffer now included physiological concentrations of ADP and inorganic phosphate Pi, as well as 200 nM ATP, in order to identify active rotating molecules. As before, the chamber was closed around a single enzyme, but its rotation was now forced into the clockwise direction using a rotating magnetic field and the magnetic bead as a handle. The maximum rate of ATP synthesis for this thermophilic F0F1 enzyme was biochemically determined to be 30 ATPs s-1 at room temperature (R. Iino and M. Yoshida, personal communication). Therefore, we selected a rotating speed of 10 rotations per second (r.p.s.). The magnetic field was applied in repetitive sequences of 1 min duration, with an interval of ∼1 min (Fig. 3a, b). When it was turned off, the bead resumed its anticlockwise rotation at a speed proportional to the ATP concentration.
a, b, An F1-ɛ/+ɛ enzyme bound to a magnetic bead was rotated in a backward (clockwise) direction with magnetic tweezers within a microchamber in the presence of ADP, Pi and a small amount of ATP. Synthesized ATP accumulated in the chamber. Before and after a forced clockwise rotation, spontaneous anticlockwise rotation was recorded as a means of detecting ATP synthesis through the increased rotational speed (see inset). c, The anticlockwise rotation speeds at each experimental step in a are shown. Between the second and third point, the motor was stalled without rotation in order to confirm the absence of background ATP synthesis. d, e, Anticlockwise rotation speeds were converted into the total number of synthesized ATPs for F1-ɛ (d) and F1+ɛ (e). Each trace belongs to an individual F1 molecule. Dotted lines indicate ATP yields expected for mechanochemical coupling ratios: 3 for 100% efficiency; 1.5 for 50%; and 0 for 0%. The standard error deviation from the rotational velocity determination (30%) and the chamber volume (10%) is indicated at each end of the traces only, for the sake of clarity.
If perfect mechanical coupling is assumed, the enzyme should produce three ATPs per clockwise turn. The concentration increment after 10 Hz rotation for 1 min corresponds to 500 nM ATP in 6 fl. This is a significantly higher concentration than the initial ATP concentration of 200 nM, thus one would expect a marked acceleration of the anticlockwise rotation after release of the magnetic field. Notably, this was not the case for F1-ɛ: the beads hardly increased their speed. Analysis of these experiments (Fig. 3d) gave a coupling ratio of 0.5 ± 0.4 (mean ± s.d.) ATPs produced per turn. This is in agreement with a previous bulk experiment13, indicating a very low mechanochemical coupling efficiency of F1-ɛ at 10 Hz.
The results were different for F1+ɛ, reconstituted from F1-ɛ and the ɛ-subunit. In this case, the forced backward rotations led, in most experiments, to a large increase in the rotation speed (Fig. 3b, c). These reproducible patterns confirmed the production of ATP inside the microchamber. For computation of the ATP yield from the rotation speed, calibration curves of the speed versus ATP concentration (ranging from 200 nM to 2 mM) were determined in independent experiments in the same buffer (see Supplementary Information). It followed Michaelis–Menten kinetics. We then calculated the number of ATP molecules produced from the increment of ATP concentration and the chamber volume to compare with the number of forced clockwise rotations (Fig. 3e). Most traces were found between dotted lines for coupling ratios of 3 and 1.5. We obtained an average value of 2.3 ± 1.6 (mean ± s.d.) ATPs per turn (77%) after eliminating obviously defective traces (blue and grey traces in Fig. 3e). This is probably a lower limit, because most artefacts, such as transient leaks in the chamber, defective connections in the glass–F1–streptavidin-bead complex, or contamination by other free motors, should decrease this value. Therefore our data point to an excellent mechanochemical coupling efficiency. In the best cases, we observed the postulated value of three ATPs synthesized per turn.
Efficiency of molecular machines is generally difficult to assess because it requires precise mechanical information—available only at the single-molecule level—while simultaneously measuring minute amounts of chemical output17. Moreover, it is easier to detect a mechanical output from a chemical input than to do the contrary. The mechanical synthesis of ATP by reversing F1 molecules was previously reported13, but it was not a quantitative measurement. As a consequence, a good understanding of the ATP-hydrolysing function of F1 has been accumulated1,3,4, but very little was known about its biologically more relevant ATP synthesis ability2. In this approach we have taken advantage of transparent containers so small that a single enzyme's chemical activity quickly results in detectable concentration changes16. Our experiments show that both F1-ɛ and F1+ɛ consume strictly three ATPs per anticlockwise turn, whereas only F1+ɛ is efficient enough to produce ATP when forced to rotate in the backward (clockwise) direction. These results are consistent with the ubiquity of this strategic enzyme that fuels most of the energy-consuming biological processes.
The present work reveals the unexpected importance of the ɛ-subunit in the synthesis of ATP. Its regulatory role for ATP hydrolysis is well recognized18,19,20 but its exact function in synthesis is still a matter for discussion. Our findings suggest that the ɛ-subunit is necessary for high-yield and high-rate ATP synthesis. It was already known that the absence of the ɛ-subunit suppresses the synthesizing ability of chloroplast F0F121, even though the enzyme is structurally intact. Recent biochemical studies have suggested that ɛ can insert its helical domain into the α3β3 ring, in a cleft along the γ-subunit22,23,24. This could be the structural basis for a switch of the enzyme into the ATP synthesis mode23, and/or a shift in the ATP/ADP affinity balance2. Alternatively, as shown in the crystal structure25 of F1+ɛ, the ɛ-subunit may stabilize the protruding part of the γ-subunit where the mechanical torque is applied. In this case, it would possess mostly a structural function. The precise role of the ɛ-subunit in ATP synthesis remains to be established. The experimental set-up described here can also be used to investigate the parameters of the original mechanical-to-chemical biological transduction performed by F1, such as maximum velocity, torque, energy barriers and affinities for substrates.
Methods
Materials
All buffers were prepared with 0.05 mg ml-1 heat-shock-purified BSA, 50 mM MOPS pH 7, 50 mM KCl and 2 mM MgCl2. For the ATP-driven rotation assay, 500 nM ATP was added, whereas for ATP synthesis the buffer contained 200 nM ATP, 10 mM Pi and 100 µM ADP, which was purified as previously described13. In buffers containing both BSA and ADP, spontaneous ATP synthesis activity was detected owing to contaminating adenylate kinase in BSA. However, the activity was negligible, less than 2 nM h-1. A mutant F1-ɛ (His6-αC193S, His10-β and γS107C/I210C), derived from the thermophilic Bacillus PS3, was prepared and biotinylated as described26. Six His tags were used to strengthen the binding of F1 to the glass plate, but did not alter significantly the rotary and catalytic properties of F1 as compared to previous mutants9. Recombinant ɛ-subunit was prepared as described7 and mixed at 7.5 µM with 0.5 µM F1-ɛ, in order to reconstitute F1+ɛ. PDMS sheets with microchambers were prepared by conventional moulding procedures27 with a slight modification16. The dimensions of the chambers, determined from electron microscope images, were 1.13 µm radius and 1.50 µm height, with an average volume of 6.0 ± 0.6 fl (mean ± s.d.).
Equipment
The magnetic field was generated by custom-made magnetic tweezers15 mounted on the stage of a microscope (Olympus, IX71). The bead rotation was recorded using a CCD camera (MTI, RC300) at the video rate, and the trajectories of rotation were analysed using custom-made software (R. Yasuda). During the time intervals when the magnetic field was cut off, the speed of ATP-driven rotation was calculated from more than five successive turns28.
ATP-driven rotation assay in a chamber
A 2 × 20 mm2 flow cell was constructed by suspending a PDMS sheet on top of a Ni-NTA-modified cover plate with two 40-µm spacer sheets. F1 molecules were immobilized on the cover plate at a very low concentration (4.7 pM), such that the density of immobilized F1 was less than one enzyme per 4 µm2 (the surface of a chamber). Streptavidin-coated magnetic beads (Seradyn; ∼0.47-µm diameter) were then attached to the biotinylated γ-subunit. When a rotating bead was found, it was enclosed in a microchamber by pressing down on the PDMS sheet using a glass needle (30-µm diameter) controlled by a z-axis micromanipulator (Narishige). After visual confirmation of contact between the sheet and the cover plate, the glass needle was pressed down a further 2–3 µm to seal the chambers tightly. This value is less than 10% of the value required for the complete compression (more than 30 µm). Thus, taking the elastomeric properties of PDMS29 into account, the chamber volume was not strongly modified under these conditions; this is also supported by the ATP-driven rotation experiments. On rare occasions, small focus drifts and transient leakage of the chamber were observed, in which case the entire experiment was rejected.
Mechanical ATP synthesis in a chamber
The set-up was similar to previous experiments, but magnetic tweezers and synthesis buffer were used. Under these high ADP conditions, more than 90% of F1-ɛ or F1+ɛ molecules were in the Mg-ADP-inhibited form (data not shown). Therefore, most of the magnetic beads did not show rotation, and they had to be pushed into the active state by mechanical, 180° clockwise rotation15. A 10 Hz rotating magnetic field was applied for 1 min to force 600 turns in the clockwise direction. Some beads (possibly with a lower magnetic content) did not perfectly follow the rotating field, and thus the actual number of clockwise turns had to be directly counted from the video analysis. Between rounds of the forced rotation, speed of the ATP-driven anticlockwise rotation was measured. The standard curves (see Supplementary Information) were built from independent experiments in regular flow cells. Michaelis–Menten parameters (F1-ɛ: Km = 3.3 µM, Vmax = 4.5 r.p.s.; F1+ɛ: Km = 2.1 µM, Vmax = 4.9 r.p.s.) and the chamber volumes were used to convert rotational speed into the total number of synthesized ATP molecules.
References
Boyer, P. D. The ATP synthase–a splendid molecular machine. Annu. Rev. Biochem. 66, 717–749 (1997)
Senior, A. E., Nadanaciva, S. & Weber, J. The molecular mechanism of ATP synthesis by F1F0-ATP synthase. Biochim. Biophys. Acta 1553, 188–211 (2002)
Yoshida, M., Muneyuki, E. & Hisabori, T. ATP synthase–a marvellous rotary engine of the cell. Nature Rev. Mol. Cell Biol. 2, 669–677 (2001)
Kinosita, K. Jr, Yasuda, R. & Noji, H. F1-ATPase: a highly efficient rotary ATP machine. Essays Biochem. 35, 3–18 (2000)
Leslie, A. G. & Walker, J. E. Structural model of F1-ATPase and the implications for rotary catalysis. Phil. Trans R. Soc. Lond. B 355, 465–471 (2000)
Noji, H., Yasuda, R., Yoshida, M. & Kinosita, K. Jr Direct observation of the rotation of F1-ATPase. Nature 386, 299–302 (1997)
Kato-Yamada, Y., Noji, H., Yasuda, R., Kinosita, K. Jr & Yoshida, M. Direct observation of the rotation of ɛ subunit in F1-ATPase. J. Biol. Chem. 273, 19375–19377 (1998)
Yasuda, R., Noji, H., Kinosita, K. Jr & Yoshida, M. F1-ATPase is a highly efficient molecular motor that rotates with discrete 120 degree steps. Cell 93, 1117–1124 (1998)
Yasuda, R., Noji, H., Yoshida, M., Kinosita, K. Jr & Itoh, H. Resolution of distinct rotational substeps by submillisecond kinetic analysis of F1-ATPase. Nature 410, 898–904 (2001)
Abrahams, J. P., Leslie, A. G., Lutter, R. & Walker, J. E. Structure at 2.8 Å resolution of F1-ATPase from bovine heart mitochondria. Nature 370, 621–628 (1994)
Zhou, Y., Duncan, T. M. & Cross, R. L. Subunit rotation in Escherichia coli FoF1-ATP synthase during oxidative phosphorylation. Proc. Natl Acad. Sci. USA 94, 10583–10587 (1997)
Diez, M. et al. Proton-powered subunit rotation in single membrane-bound FoF1-ATP synthase. Nature Struct. Mol. Biol. 11, 135–141 (2004)
Itoh, H. et al. Mechanically driven ATP synthesis by F1-ATPase. Nature 427, 465–468 (2004)
Hirono-Hara, Y. et al. Pause and rotation of F1-ATPase during catalysis. Proc. Natl Acad. Sci. USA 98, 13649–13654 (2001)
Hirono-Hara, Y. et al. Activation of pausing F1-motor by external force. Proc. Natl Acad. Sci. USA.(in the press)
Rondelez, Y. et al. Microfabricated array of femtoliter chambers allow single molecule enzymology. Nature Biotechnol.(in the press)
Nishizaka, T. et al. Chemomechanical coupling in F1-ATPase revealed by simultaneous observation of nucleotide kinetics and rotation. Nature Struct. Mol. Biol. 11, 142–148 (2004)
Laget, P. P. & Smith, J. B. Inhibitory properties of endogenous subunit ɛ in the Escherichia coli F1 ATPase. Arch. Biochem. Biophys. 197, 83–89 (1979)
Dunn, S. D., Zadorozny, V. D., Tozer, R. G. & Orr, L. E. Epsilon subunit of Escherichia coli F1-ATPase: effects on affinity for aurovertin and inhibition of product release in unisite ATP hydrolysis. Biochemistry 26, 4488–4493 (1987)
Kato, Y. et al. Thermophilic F1-ATPase is activated without dissociation of an endogenous inhibitor, ɛ subunit. J. Biol. Chem. 272, 24906–24912 (1997)
Nowak, K. F., Tabidze, V. & McCarty, R. E. The C-terminal domain of the ɛ subunit of the chloroplast ATP synthase is not required for ATP synthesis. Biochemistry 41, 15130–15134 (2002)
Tsunoda, S. P. et al. Large conformational changes of the ɛ subunit in the bacterial F1Fo ATP synthase provide a ratchet action to regulate this rotary motor enzyme. Proc. Natl Acad. Sci. USA 98, 6560–6564 (2001)
Suzuki, T. et al. FoF1-ATPase/synthase is geared to the synthesis mode by conformational rearrangement of ɛ subunit in response to proton motive force and ADP/ATP balance. J. Biol. Chem. 278, 46840–46846 (2003)
Bulygin, V. V., Duncan, T. M. & Cross, R. L. Rotor/stator interactions of the ɛ subunit in Escherichia coli ATP synthase and implications for enzyme regulation. J. Biol. Chem. 279, 35616–35621 (2004)
Gibbons, C., Montgomery, M. G., Leslie, A. G. & Walker, J. E. The structure of the central stalk in bovine F1-ATPase at 2.4 Å resolution. Nature Struct. Biol. 7, 1055–1061 (2000)
Noji, H. et al. Purine but not pyrimidine nucleotides support rotation of F1-ATPase. J. Biol. Chem. 276, 25480–25486 (2001)
McDonald, J. C. & Whitesides, G. M. Poly(dimethylsiloxane) as a material for fabricating microfluidic devices. Acc. Chem. Res. 35, 491–499 (2002)
Adachi, K., Noji, H. & Kinosita, K. Jr Single-molecule imaging of rotation of F1-ATPase. Methods Enzymol. 361, 211–227 (2003)
Unger, M. A., Chou, H. P., Thorsen, T., Scherer, A. & Quake, S. R. Monolithic microfabricated valves and pumps by multilayer soft lithography. Science 288, 113–116 (2000)
Matsui, T. & Yoshida, M. Expression of the wild-type and the Cys-/Trp-less α3β3γ complex of thermophilic F1-ATPase in Escherichia coli. Biochim. Biophys. Acta 1231, 139–149 (1995)
Acknowledgements
We thank all members of the Noji and Takeuchi laboratories, and H. Arata and A. Tixier-Mita for discussion and experimental support; R. Yasuda for PC programming of image analysis (CREST image); and Central Workshop in IIS for an optical microscope stage. This work was performed in the framework of LIMMS/CNRS-IIS, and supported in part by Bio-oriented Technology Research Advancement Institution (H.N. and S.T.), and Grants-in-Aid from Ministry of Education, Science, Sports and Culture of Japan (H.N., H.F. and S.T.). Y.R. and G.T. are Research Fellows of the Japan Society for the Promotion of Science.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Supplemental Information
1. The video time table for the ATP synthesis experiment. 2. Standard curves of rotational velocity versus ATP concentration. (DOC 38 kb)
Rights and permissions
About this article
Cite this article
Rondelez, Y., Tresset, G., Nakashima, T. et al. Highly coupled ATP synthesis by F1-ATPase single molecules. Nature 433, 773–777 (2005). https://doi.org/10.1038/nature03277
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature03277
This article is cited by
-
Extreme-value analysis of intracellular cargo transport by motor proteins
Communications Physics (2024)
-
Micro- and nanochamber array system for single enzyme assays
Scientific Reports (2023)
-
Reconstitution of FoF1-ATPase-based biomimetic systems
Nature Reviews Chemistry (2019)
-
Control of rotation of the F1FO-ATP synthase nanomotor by an inhibitory α-helix from unfolded ε or intrinsically disordered ζ and IF1 proteins
Journal of Bioenergetics and Biomembranes (2018)
-
Oscillating Electric Field Measures the Rotation Rate in a Native Rotary Enzyme
Scientific Reports (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.