Abstract
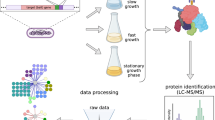
Proteins often function as components of multi-subunit complexes. Despite its long history as a model organism1, no large-scale analysis of protein complexes in Escherichia coli has yet been reported. To this end, we have targeted DNA cassettes into the E. coli chromosome to create carboxy-terminal, affinity-tagged alleles of 1,000 open reading frames (∼ 23% of the genome). A total of 857 proteins, including 198 of the most highly conserved, soluble non-ribosomal proteins essential in at least one bacterial species, were tagged successfully, whereas 648 could be purified to homogeneity and their interacting protein partners identified by mass spectrometry. An interaction network of protein complexes involved in diverse biological processes was uncovered and validated by sequential rounds of tagging and purification. This network includes many new interactions as well as interactions predicted based solely on genomic inference or limited phenotypic data2. This study provides insight into the function of previously uncharacterized bacterial proteins and the overall topology of a microbial interaction network, the core components of which are broadly conserved across Prokaryota.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Neidhardt, F. (ed.) Escherichia coli and Salmonella: Cellular and Molecular Biology (ASM Press, Washington DC, 1996)
Serres, M. H. et al. A functional update of the Escherichia coli K-12 genome. Genome Biol. 2, research0035.1–0035.7 (2001)
Gavin, A. C. et al. Functional organization of the yeast proteome by systematic analysis of protein complexes. Nature 415, 141–147 (2002)
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl Acad. Sci. USA 97, 6640–6645 (2000)
Yu, D. et al. An efficient recombination system for chromosome engineering in Escherichia coli . Proc. Natl Acad. Sci. USA 97, 5978–5983 (2000)
Zeghouf, M. et al. Sequential Peptide Affinity (SPA) system for the identification of mammalian and bacterial protein complexes. J. Proteome Res. 3, 463–468 (2004)
Stukenberg, P. T. & O'Donnell, M. Assembly of a chromosomal replication machine: two DNA polymerases, a clamp loader, and sliding clamps in one holoenzyme particle. V. Four different polymerase-clamp complexes on DNA. J. Biol. Chem. 270, 13384–13391 (1995)
Harmon, F. G., Brockman, J. P. & Kowalczykowski, S. C. RecQ helicase stimulates both DNA catenation and changes in DNA topology by topoisomerase III. J. Biol. Chem. 278, 42668–42678 (2003)
Witte, G., Urbanke, C. & Curth, U. DNA polymerase III chi subunit ties single-stranded DNA binding protein to the bacterial replication machinery. Nucleic Acids Res. 31, 4434–4440 (2003)
Nakayama, H. RecQ family helicases: roles as tumor suppressor proteins. Oncogene 21, 9008–9021 (2002)
von Mering, C. et al. Comparative assessment of large-scale data sets of protein–protein interactions. Nature 417, 399–403 (2002)
Salwinski, L. et al. The Database of Interacting Proteins: 2004 update. Nucleic Acids Res. 32 (Database issue), D449–D451 (2004)
Bader, G. D., Betel, D. & Hogue, C. W. BIND: the Biomolecular Interaction Network Database. Nucleic Acids Res. 31, 248–250 (2003)
von Mering, C. et al. STRING: a database of predicted functional associations between proteins. Nucleic Acids Res. 31, 258–261 (2003)
Bowers, P. et al. Prolinks: a database of protein functional linkages derived from coevolution. Genome Biol. 5, R35 (2004)
Rain, J. C. et al. The protein–protein interaction map of Helicobacter pylori . Nature 409, 211–215 (2001)
Gully, D., Moinier, D., Loiseau, L. & Bouveret, E. New partners of acyl carrier protein detected in Escherichia coli by tandem affinity purification. FEBS Lett. 548, 90–96 (2003)
Wuchty, S., Oltvai, Z. N. & Barabasi, A. L. Evolutionary conservation of motif constituents in the yeast protein interaction network. Nature Genet. 35, 176–179 (2003)
Jordan, I. K., Wolf, Y. I. & Koonin, E. V. No simple dependence between protein evolution rate and the number of protein-protein interactions: only the most prolific interactors tend to evolve slowly. BMC Evol. Biol. 3, 1 (2003)
Fraser, H. B., Wall, D. P. & Hirsh, A. E. A simple dependence between protein evolution rate and the number of protein-protein interactions. BMC Evol. Biol. 3, 11 (2003)
Peregrin-Alvarez, J. M., Tsoka, S. & Ouzounis, C. A. The phylogenetic extent of metabolic enzymes and pathways. Genome Res. 13, 422–427 (2003)
Tatusov, R. L., Koonin, E. V. & Lipman, D. J. A genomic perspective on protein families. Science 278, 631–637 (1997)
Wu, J., Kasif, S. & DeLisi, C. Identification of functional links between genes using phylogenetic profiles. Bioinformatics 19, 1524–1530 (2003)
Pellegrini, M., Marcotte, E. M., Thompson, M. J., Eisenberg, D. & Yeates, T. O. Assigning protein functions by comparative genome analysis: protein phylogenetic profiles. Proc. Natl Acad. Sci. USA 96, 4285–4288 (1999)
Date, S. V. & Marcotte, E. M. Discovery of uncharacterized cellular systems by genome-wide analysis of functional linkages. Nature Biotechnol. 21, 1055–1062 (2003)
Yu, H. et al. Annotation transfer between genomes: protein-protein interologs and protein-DNA regulogs. Genome Res. 14, 1107–1118 (2004)
Overbeek, R., Fonstein, M., D'Souza, M., Pusch, G. D. & Maltsev, N. The use of gene clusters to infer functional coupling. Proc. Natl Acad. Sci. USA 96, 2896–2901 (1999)
Haselbeck, R. et al. Comprehensive essential gene identification as a platform for novel anti-infective drug discovery. Curr. Pharm. Des. 8, 1155–1172 (2002)
Krogan, N. J. et al. High-definition macromolecular composition of yeast RNA-processing complexes. Mol. Cell 13, 225–239 (2004)
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997)
Acknowledgements
The authors wish to thank C. J. Ingles and M. Shales for comments on the manuscript. This work was supported by funds from the Ontario Research and Development Challenge Fund and Genome Canada to A.E. and J.G. G.B. was a recipient of a Charles H. Best Post-Doctoral Fellowship. J.M.P.-A. acknowledges support from the Hospital for Sick Children (Toronto, Ontario, Canada) Research Training Centre. Computer analyses were undertaken at the Centre for Computational Biology, Hospital for Sick Children.Authors’ contributions Informatics studies were performed and analysed by J.M.P.-A. and J.P. Experimental design and data analysis were coordinated by G.B. Tagging and purification experiments were performed by W.Y., X.Y., J.L. and G.B. V.C., A.S., D.R., B.B., N.J.K. and M.D. performed and assisted with mass spectrometry analysis. The manuscript was jointly drafted by G.B., A.E., J.G., J.M.P.-A. and J.P. The project was conceived and designed by J.G. and was directed by A.E.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Supplementary Notes 1
This contains the Supplementary Discussion, Supplementary Methods and legends to accompany the Supplementary Figures (S1-S6) and Supplementary Tables (S1-7). (DOC 69 kb)
Supplementary Notes 2
This contains a brief description for each of the Supplementary Figures and Supplementary Tables. (DOC 23 kb)
Supplementary Figures S1-6
This file contains Supplementary Figures S1-S6. (PDF 1878 kb)
Supplementary Tables S1-S7
This file contains Supplementary Tables S1-S7. (PDF 772 kb)
Rights and permissions
About this article
Cite this article
Butland, G., Peregrín-Alvarez, J., Li, J. et al. Interaction network containing conserved and essential protein complexes in Escherichia coli. Nature 433, 531–537 (2005). https://doi.org/10.1038/nature03239
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature03239
This article is cited by
-
Interaction networks of Escherichia coli replication proteins under different bacterial growth conditions
Scientific Data (2023)
-
Interaction between transcribing RNA polymerase and topoisomerase I prevents R-loop formation in E. coli
Nature Communications (2022)
-
Compartmentalization of the replication fork by single-stranded DNA-binding protein regulates translesion synthesis
Nature Structural & Molecular Biology (2022)
-
Network-based analysis of virulence factors for uncovering Aeromonas veronii pathogenesis
BMC Microbiology (2021)
-
Acyl carrier protein promotes MukBEF action in Escherichia coli chromosome organization-segregation
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.