Abstract
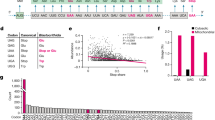
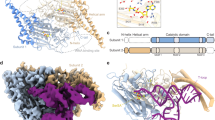
Analysis of the genome sequence of the small hyperthermophilic archaeal parasite Nanoarchaeum equitans1,2 has not revealed genes encoding the glutamate, histidine, tryptophan and initiator methionine transfer RNA species. Here we develop a computational approach to genome analysis that searches for widely separated genes encoding tRNA halves that, on the basis of structural prediction, could form intact tRNA molecules. A search of the N. equitans genome reveals nine genes that encode tRNA halves; together they account for the missing tRNA genes. The tRNA sequences are split after the anticodon-adjacent position 37, the normal location of tRNA introns. The terminal sequences can be accommodated in an intervening sequence that includes a 12–14-nucleotide GC-rich RNA duplex between the end of the 5′ tRNA half and the beginning of the 3′ tRNA half. Reverse transcriptase polymerase chain reaction and aminoacylation experiments of N. equitans tRNA demonstrated maturation to full-size tRNA and acceptor activity of the tRNAHis and tRNAGlu species predicted in silico. As the joining mechanism possibly involves tRNA trans-splicing, the presence of an intron might have been required for early tRNA synthesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Huber, H. et al. A new phylum of Archaea represented by a nanosized hyperthermophilic symbiont. Nature 417, 63–67 (2002)
Waters, E. et al. The genome of Nanoarchaeum equitans: insights into early archaeal evolution and derived parasitism. Proc. Natl Acad. Sci. USA 100, 12984–12988 (2003)
Di Giulio, M. The non-monophyletic origin of the tRNA molecule. J. Theor. Biol. 197, 403–414 (1999)
Tanaka, T. & Kikuchi, Y. Origin of the cloverleaf shape of transfer RNA—the double-hairpin model: Implication for the role of tRNA intron and the long extra loop. Viva Origino 29, 134–142 (2001)
Boucher, Y. & Doolittle, W. F. Something new under the sea. Nature 417, 27–28 (2002)
Marck, C. & Grosjean, H. tRNomics: analysis of tRNA genes from 50 genomes of Eukarya, Archaea, and Bacteria reveals anticodon-sparing strategies and domain-specific features. RNA 8, 1189–1232 (2002)
Eddy, S. R. & Durbin, R. RNA sequence analysis using covariance models. Nucleic Acids Res. 22, 2079–2088 (1994)
Lowe, T. M. & Eddy, S. R. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 25, 955–964 (1997)
Marck, C. & Grosjean, H. Identification of BHB splicing motifs in intron-containing tRNAs from 18 archaea: evolutionary implications. RNA 9, 1516–1531 (2003)
Hain, J., Reiter, W. D., Hudepohl, U. & Zillig, W. Elements of an archaeal promoter defined by mutational analysis. Nucleic Acids Res. 20, 5423–5428 (1992)
Connolly, S. A., Rosen, A. E., Musier-Forsyth, K. & Francklyn, C. S. G-1:C73 recognition by an arginine cluster in the active site of Escherichia coli histidyl-tRNA synthetase. Biochemistry 43, 962–969 (2004)
Sekine, S. et al. Major identity determinants in the ‘augmented D helix’ of tRNA(Glu) from Escherichia coli . J. Mol. Biol. 256, 685–700 (1996)
Schurer, H., Schiffer, S., Marchfelder, A. & Mörl, M. This is the end: processing, editing and repair at the tRNA 3′-terminus. Biol. Chem. 382, 1147–1156 (2001)
Xiong, Y., Li, F., Wang, J., Weiner, A. M. & Steitz, T. A. Crystal structures of an archaeal class I CCA-adding enzyme and its nucleotide complexes. Mol. Cell 12, 1165–1172 (2003)
Lohan, A. J. & Gray, M. W. Methods for analysis of mitochondrial tRNA editing in Acanthamoeba castellanii . Methods Mol. Biol. 265, 315–332 (2004)
Varshney, U., Lee, C. P. & RajBhandary, U. L. Direct analysis of aminoacylation levels of tRNAs in vivo. Application to studying recognition of Escherichia coli initiator tRNA mutants by glutaminyl-tRNA synthetase. J. Biol. Chem. 266, 24712–24718 (1991)
Konarska, M. M., Padgett, R. A. & Sharp, P. A. Trans splicing of mRNA precursors in vitro . Cell 42, 165–171 (1985)
Solnick, D. Does trans splicing in vitro require base pairing between RNAs? Cell 44, 211 (1986)
Wissinger, B., Schuster, W. & Brennicke, A. Trans splicing in Oenothera mitochondria: nad1 mRNAs are edited in exon and trans-splicing group II intron sequences. Cell 65, 473–482 (1991)
Abelson, J., Trotta, C. R. & Li, H. tRNA splicing. J. Biol. Chem. 273, 12685–12688 (1998)
Fabbri, S. et al. Conservation of substrate recognition mechanisms by tRNA splicing endonucleases. Science 280, 284–286 (1998)
Li, H. & Abelson, J. Crystal structure of a dimeric archaeal splicing endonuclease. J. Mol. Biol. 302, 639–648 (2000)
Kleman-Leyer, K., Armbruster, D. W. & Daniels, C. J. Properties of H. volcanii tRNA intron endonuclease reveal a relationship between the archaeal and eucaryal tRNA intron processing systems. Cell 89, 839–847 (1997)
Salgia, S. R., Singh, S. K., Gurha, P. & Gupta, R. Two reactions of Haloferax volcanii RNA splicing enzymes: joining of exons and circularization of introns. RNA 9, 319–330 (2003)
Maizels, N. & Weiner, A. M. Phylogeny from function: evidence from the molecular fossil record that tRNA originated in replication, not translation. Proc. Natl Acad. Sci. USA 91, 6729–6734 (1994)
Nagaswamy, U. & Fox, G. E. RNA ligation and the origin of tRNA. Orig. Life Evol. Biosph. 33, 199–209 (2003)
Gopalan, V., Vioque, A. & Altman, S. RNase P: variations and uses. J. Biol. Chem. 277, 6759–6762 (2002)
Schneider, T. D., Stormo, G. D., Gold, L. & Ehrenfeucht, A. Information content of binding sites on nucleotide sequences. J. Mol. Biol. 188, 415–431 (1986)
Curnow, A. W., Tumbula, D. L., Pelaschier, J. T., Min, B. & Söll, D. Glutamyl-tRNAGln amidotransferase in Deinococcus radiodurans may be confined to asparagine biosynthesis. Proc. Natl Acad. Sci. USA 95, 12838–12843 (1998)
Williams, M. A., Johzuka, Y. & Mulligan, R. M. Addition of non-genomically encoded nucleotides to the 3′-terminus of maize mitochondrial mRNAs: truncated rps12 mRNAs frequently terminate with CCA. Nucleic Acids Res. 28, 4444–4451 (2000)
Acknowledgements
We thank H. Huber and K. O. Stetter for advice and spirited discussions, M. Thomm for the use of laboratory facilities, and J. Yuan and J. Sabina for critically reading the manuscript. This work was supported by grants from the National Institute of General Medical Sciences and the Department of Energy (to D.S.) and by the German Federal Ministry of Education and Research (BMBF) for the Bioinformatics Competence Center ‘Intergenomics’ (to D.J.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Rights and permissions
About this article
Cite this article
Randau, L., Münch, R., Hohn, M. et al. Nanoarchaeum equitans creates functional tRNAs from separate genes for their 5′- and 3′-halves. Nature 433, 537–541 (2005). https://doi.org/10.1038/nature03233
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature03233
This article is cited by
-
G:U-Independent RNA Minihelix Aminoacylation by Nanoarchaeum equitans Alanyl-tRNA Synthetase: An Insight into the Evolution of Aminoacyl-tRNA Synthetases
Journal of Molecular Evolution (2020)
-
An RNA Ring was Not the Progenitor of the tRNA Molecule
Journal of Molecular Evolution (2020)
-
The Uroboros Theory of Life’s Origin: 22-Nucleotide Theoretical Minimal RNA Rings Reflect Evolution of Genetic Code and tRNA-rRNA Translation Machineries
Acta Biotheoretica (2019)
-
Binding Properties of Split tRNA to the C-terminal Domain of Methionyl-tRNA Synthetase of Nanoarchaeum equitans
Journal of Molecular Evolution (2017)
-
Genomics-informed isolation and characterization of a symbiotic Nanoarchaeota system from a terrestrial geothermal environment
Nature Communications (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.