Abstract
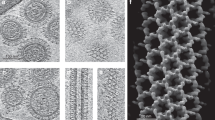
Membranes are essential for selectively controlling the passage of molecules in and out of cells and mediating the response of cells to their environment. Biological membranes and their associated proteins present considerable difficulties for structural analysis. Although enveloped viruses have been imaged at about 9 Å resolution by cryo-electron microscopy and image reconstruction1,2, no detailed crystallographic structure of a membrane system has been described. The structure of the bacteriophage PRD1 particle, determined by X-ray crystallography at about 4 Å resolution, allows the first detailed analysis of a membrane-containing virus3. The architecture of the viral capsid and its implications for virus assembly are presented in the accompanying paper3. Here we show that the electron density also reveals the icosahedral lipid bilayer, beneath the protein capsid, enveloping the viral DNA. The viral membrane contains about 26,000 lipid molecules asymmetrically distributed between the membrane leaflets. The inner leaflet is composed predominantly of zwitterionic phosphatidylethanolamine molecules, facilitating a very close interaction with the viral DNA, which we estimate to be packaged to a pressure of about 45 atm, factors that are likely to be important during membrane-mediated DNA translocation into the host cell. In contrast, the outer leaflet is enriched in phosphatidylglycerol and cardiolipin, which show a marked lateral segregation within the icosahedral asymmetric unit. In addition, the lipid headgroups show a surprising degree of order.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Zhang, W. et al. Visualization of membrane protein domains by cryo-electron microscopy of dengue virus. Nature Struct. Biol. 10, 907–912 (2003)
Mancini, E. J., Clarke, M., Gowen, B. E., Rutten, T. & Fuller, S. D. Cryo-electron microscopy reveals the functional organization of an enveloped virus, Semliki Forest virus. Mol. Cell 5, 255–266 (2000)
Abrescia, N. G. et al. Insights into assembly from structural analysis of bacteriophage PRD1. Nature doi:10.1038/nature03056 (this issue)
Bamford, D. H., Caldentey, J. & Bamford, J. K. Bacteriophage PRD1: a broad host range dsDNA tectivirus with an internal membrane. Adv. Virus Res. 45, 281–319 (1995)
Butcher, S. J., Bamford, D. H. & Fuller, S. D. DNA packaging orders the membrane of bacteriophage PRD1. EMBO J. 14, 6078–6086 (1995)
San Martín, C. et al. Combined EM/X-ray imaging yields a quasi-atomic model of the adenovirus-related bacteriophage PRD1 and shows key capsid and membrane interactions. Structure 9, 917–930 (2001)
Rydman, P. S., Bamford, J. K. & Bamford, D. H. A minor capsid protein P30 is essential for bacteriophage PRD1 capsid assembly. J. Mol. Biol. 313, 785–795 (2001)
Rydman, P. S. et al. Bacteriophage PRD1 contains a labile receptor-binding structure at each vertex. J. Mol. Biol. 291, 575–587 (1999)
Strömsten, N. J., Bamford, D. H. & Bamford, J. K. The unique vertex of bacterial virus PRD1 is connected to the viral internal membrane. J. Virol. 77, 6314–6321 (2003)
Gowen, B., Bamford, J. K., Bamford, D. H. & Fuller, S. D. The tailless icosahedral membrane virus PRD1 localizes the proteins involved in genome packaging and injection at a unique vertex. J. Virol. 77, 7863–7871 (2003)
Bamford, D. & Mindich, L. Structure of the lipid-containing bacteriophage PRD1: disruption of wild-type and nonsense mutant phage particles with guanidine hydrochloride. J. Virol. 44, 1031–1038 (1982)
Rydman, P. S. & Bamford, D. H. Bacteriophage PRD1 DNA entry uses a viral membrane-associated transglycosylase activity. Mol. Microbiol. 37, 356–363 (2000)
Rydman, P. S. & Bamford, D. H. The lytic enzyme of bacteriophage PRD1 is associated with the viral membrane. J. Bacteriol. 184, 104–110 (2002)
Bamford, J. K. et al. Diffraction quality crystals of PRD1, a 66-MDa dsDNA virus with an internal membrane. J. Struct. Biol. 139, 103–112 (2002)
Grahn, A. M., Daugelavicius, R. & Bamford, D. H. Sequential model of phage PRD1 DNA delivery: active involvement of the viral membrane. Mol. Microbiol. 46, 1199–1209 (2002)
Davis, T. N., Muller, E. D. & Cronan, J. E. Jr The virion of the lipid-containing bacteriophage PR4. Virology 120, 287–306 (1982)
Grahn, A. M., Caldentey, J., Bamford, J. K. & Bamford, D. H. Stable packaging of phage PRD1 DNA requires adsorption protein P2, which binds to the IncP plasmid-encoded conjugative transfer complex. J. Bacteriol. 181, 6689–6696 (1999)
Tuma, R., Bamford, J. K., Bamford, D. H. & Thomas, G. J. Jr Structure, interactions and dynamics of PRD1 virus. II. Organization of the viral membrane and DNA. J. Mol. Biol. 257, 102–115 (1996)
Laurinavicius, S., Käkelä, R., Somerharju, P. & Bamford, D. H. Phospholipid molecular species profiles of tectiviruses infecting Gram-negative and Gram-positive hosts. Virology 322, 328–336 (2004)
Nagle, J. F. & Tristram-Nagle, S. Structure of lipid bilayers. Biochim. Biophys. Acta 1469, 159–195 (2000)
Struth, B. et al. Organization of two-dimensional phospholipid monolayers on a gel-forming substrate. Phys. Rev. Lett. 88, 025502 (2002)
Cerritelli, M. E. et al. Encapsidated conformation of bacteriophage T7 DNA. Cell 91, 271–280 (1997)
Livolant, F. & Leforestier, A. Condensed phases of DNA: Structures and phase transitions. Prog. Polym. Sci. 21, 1115–1164 (1996)
Smith, D. E. et al. The bacteriophage φ29 portal motor can package DNA against a large internal force. Nature 413, 748–752 (2001)
Evans, E., Heinrich, V., Ludwig, F. & Rawicz, W. Dynamic tension spectroscopy and strength of biomembranes. Biophys. J. 85, 2342–2350 (2003)
Bevers, E. M., Comfurius, P., Dekkers, D. W. C. & Zwaal, R. F. A. Lipid translocation across the plasma membrane of mammalian cells. Biochim. Biophys. Acta 1439, 317–330 (1999)
Casper, D. L. & Kirschner, D. A. Myelin membrane structure at 10 Å resolution. Nature New Biol. 231, 46–52 (1971)
Brunger, A. T. X-PLOR, Version 3.1: A System for X-ray Crystallography and NMR (Yale University Press, New Haven, Connecticut, 1992)
Tao, Y. et al. Assembly of a tailed bacterial virus and its genome release studied in three dimensions. Cell 95, 431–437 (1998)
Acknowledgements
We thank M. Sansom and P. Bond (Laboratory of Molecular Biophysics, University of Oxford) for providing coordinates of lipid bilayers, the staff at the European Synchrotron Radiation Facility for beamline support, and E. Mancini for assistance with data collection. This investigation was supported by a research grant from the Academy of Finland to J.K.H.B., research grants and the Finnish Centre of Excellence Program (2000–2005) from the Academy of Finland to D.H.B., the Biotechnology and Biological Sciences Research Council and the Medical Research Council, UK, and a grant from the Human Frontiers Science Program to D.H.B. and D.I.S. J.M.G. is supported by the Royal Society, and D.I.S. by the UK Medical Research Council.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Supplementary Methods
1. The acquisition of Raman spectra from crystals of PRD1; 2. Initial estimation of the bulk solvent electron density in cell 2 crystals; 3. Calculation of the mean number of electrons per lipid for the different headgroup species from the mass spectrometry data; 4. Calculation of the number of lipids in the viral membrane based on published data in lipid molecular volumes. (DOC 26 kb)
Supplementary Figure 1
The Raman spectrum from PRD1 crystals showing that the viral lipids are in the liquid crystalline phase. (JPG 18 kb)
Supplementary Figure 2
Analysis of the lateral order in the electron density for the membrane. (JPG 35 kb)
Supplementary Figure 3
Autocorrelation function for features in the membrane outer leaflet headgroups. (JPG 18 kb)
Supplementary Figure 4
Summary of the mass spectrometry data on the lipid composition of the viral membrane (JPG 35 kb)
Rights and permissions
About this article
Cite this article
Cockburn, J., Abrescia, N., Grimes, J. et al. Membrane structure and interactions with protein and DNA in bacteriophage PRD1. Nature 432, 122–125 (2004). https://doi.org/10.1038/nature03053
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature03053
This article is cited by
-
Membranes as the third genetic code
Molecular Biology Reports (2020)
-
Structural basis for assembly of vertical single β-barrel viruses
Nature Communications (2019)
-
Structural conservation in a membrane-enveloped filamentous virus infecting a hyperthermophilic acidophile
Nature Communications (2018)
-
Cryo-EM structure of the mature dengue virus at 3.5-Å resolution
Nature Structural & Molecular Biology (2013)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.