Abstract
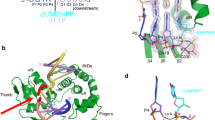
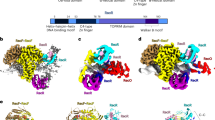
RecBCD is a multi-functional enzyme complex that processes DNA ends resulting from a double-strand break. RecBCD is a bipolar helicase that splits the duplex into its component strands and digests them until encountering a recombinational hotspot (Chi site). The nuclease activity is then attenuated and RecBCD loads RecA onto the 3′ tail of the DNA. Here we present the crystal structure of RecBCD bound to a DNA substrate. In this initiation complex, the DNA duplex has been split across the RecC subunit to create a fork with the separated strands each heading towards different helicase motor subunits. The strands pass along tunnels within the complex, both emerging adjacent to the nuclease domain of RecB. Passage of the 3′ tail through one of these tunnels provides a mechanism for the recognition of a Chi sequence by RecC within the context of double-stranded DNA. Gating of this tunnel suggests how nuclease activity might be regulated.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Kowalczykowski, S. C. Initiation of genetic recombination and recombination-dependent replication. Trends. Biochem. Sci. 25, 156–165 (2000)
Ponticelli, A. S., Schultz, D. W., Taylor, A. F. & Smith, G. R. Chi-dependent DNA strand cleavage by the RecBC enzyme. Cell 41, 145–151 (1985)
Taylor, A. F., Schultz, D. W., Ponticelli, A. S. & Smith, G. R. RecBC enzyme nicking at Chi sites during DNA unwinding: location and orientation-dependence of the cutting. Cell 41, 153–163 (1985)
Bianco, P. R. & Kowalczykowski, S. C. The recombination hotspot Chi is recognized by the translocating RecBCD enzyme as the single strand of DNA containing the sequence 5′-GCTGGTGG-3′. Proc. Natl Acad. Sci. USA 94, 6706–6711 (1997)
Spies, M. et al. A molecular throttle: the recombination hotspot Chi controls DNA translocation by the RecBCD helicase. Cell 114, 647–654 (2003)
Dixon, D. A. & Kowalczykowski, S. C. The recombination hotspot Chi is a regulatory sequence that acts by attenuating the nuclease activity of the E. coli RecBCD enzyme. Cell 73, 87–96 (1993)
Anderson, D. G. & Kowalczykowski, S. C. The recombination hot spot Chi is a regulatory element that switches the polarity of DNA degradation by the RecBCD enzyme. Genes Dev. 11, 571–581 (1997a)
Taylor, A. F. & Smith, G. R. Strand specificity of nicking of DNA at Chi sites by RecBCD enzyme. J. Biol. Chem. 270, 24459–24467 (1995b)
Anderson, D. G. & Kowalczykowski, S. C. The translocating RecBCD enzyme stimulates recombination by directing RecA protein onto ssDNA in a Chi-regulated manner. Cell 90, 77–86 (1997b)
Taylor, A. F. & Smith, G. R. Monomeric RecBCD enzyme binds and unwinds DNA. J. Biol. Chem. 270, 24451–24458 (1995a)
Boehmer, P. E. & Emmerson, P. T. Escherichia coli RecBCD enzyme: inducible overproduction and reconstitution of the ATP-dependent deoxyribonuclease from purified subunits. Gene 102, 1–6 (1991)
Yu, M., Souaya, J. & Julin, D. A. The 30 kDa C-terminal domain of the RecB protein is critical for the nuclease activity, but not the helicase activity, of the RecBCD enzyme from Escherichia coli. Proc. Natl Acad. Sci. USA 95, 981–986 (1998)
Handa, N., Ohashi, S., Kusano, K. & Kobayashi, I. Chi*, a chi-related 11-mer sequence partially active in an E. coli recC1004 strain. Genes Cells 2, 525–536 (1997)
Dillingham, M. S., Spies, M. & Kowalczykowski, S. C. RecBCD enzyme is a bipolar DNA helicase. Nature 423, 893–897 (2003)
Taylor, A. F. & Smith, G. R. RecBCD enzyme is a DNA helicase with fast and slow motors of opposite polarity. Nature 423, 889–893 (2003)
Roman, L. J. & Kowalczykowski, S. C. Characterisation of the adenosinetriphosphatase activity of the Escherichia coli RecBCD enzyme: relationship of ATP hydrolysis to the unwinding of duplex DNA. Biochemistry 28, 2873–2881 (1989)
Ganesan, S. & Smith, G. R. Strand-specific binding to duplex DNA ends by the subunits of the Escherichia coli RecBCD enzyme. J. Mol. Biol. 229, 67–78 (1993)
Farah, J. A. & Smith, G. R. The RecBCD enzyme initiation complex for DNA unwinding: enzyme positioning and DNA opening. J. Mol. Biol. 272, 699–715 (1997)
Boehmer, P. E. & Emmerson, P. T. The RecB subunit of the Escherichia coli RecBCD enzyme couples ATP hydrolysis to DNA unwinding. J. Biol. Chem. 267, 4981–4987 (1992)
Gorbalenya, A. E. & Koonin, E. V. Helicases: amino acid sequence comparisons and structure-function relationships. Curr. Opin. Struct. Biol. 3, 419–429 (1993)
Subramanya, H. S., Bird, L. E., Brannigan, J. A. & Wigley, D. B. Crystal structure of a DExx box helicase. Nature 384, 379–383 (1996)
Korolev, S., Hsieh, J., Gauss, G. H., Lohman, T. M. & Waksman, G. Major domain swiveling revealed by the crystal structures of complexes of E. coli Rep helicase bound to single-stranded DNA and ADP. Cell 90, 635–647 (1997)
Velankar, S. S., Soultanas, P., Dillingham, M. S., Subramanya, H. S. & Wigley, D. B. Crystal structures of complexes of PcrA helicase with a DNA substrate indicate an inchworm mechanism. Cell 97, 75–84 (1999)
Dillingham, M. S., Wigley, D. B. & Webb, M. R. Unidirectional single-stranded DNA translocation by PcrA helicase: measurement of step size and translocation speed. Biochemistry 39, 205–212 (2000)
Soultanas, P., Dillingham, M. S., Wiley, P., Webb, M. R. & Wigley, D. B. Uncoupling DNA translocation and helicase activity in PcrA: direct evidence for an active mechanism. EMBO J. 19, 3799–3810 (2000)
Singleton, M. R. & Wigley, D. B. Modularity and specialisation in Superfamily 1 and 2 helicases. J. Bacteriol. 184, 1819–1826 (2002)
Aravind, L., Makarova, K. S. & Koonin, E. V. Holliday junction resolvases and related nucleases: identification of new families, phyletic distribution and evolutionary trajectories. Nucleic Acids Res. 28, 3417–3432 (2000)
Kovall, R. & Matthews, B. W. Toroidal structure of λ-exonuclease. Science 277, 1824–1827 (1997)
Yu, M., Souaya, J. & Julin, D. A. Identification of the nuclease active site in the multifunctional RecBCD enzyme by creation of a chimeric enzyme. J. Mol. Biol. 283, 797–808 (1998)
Rosamond, J., Telander, K. M. & Linn, S. Modulation of the action of the RecBC enzyme of Escherichia coli K-12 by Ca2+. J. Biol. Chem. 254, 8648–8652 (1979)
Holm, L. & Sander, C. Protein structure comparison by alignment of distance matrices. J. Mol. Biol. 233, 123–138 (1993)
Amundsen, S. K., Taylor, A. F. & Smith, G. R. Domain of RecC required for assembly of the regulatory RecD subunit into the Escherichia coli RecBCD holoenzyme. Genetics 161, 483–492 (2002)
Chen, H. W., Ruan, B., Yu, M., Wang, J. & Julin, D. A. The RecD subunit of the RecBCD enzyme from Escherichia coli is a single-stranded DNA dependent ATPase. J. Biol. Chem. 272, 10072–10079 (1997)
Kuhn, B., Abdel-Monem, M., Krell, H. & Hoffmann-Berling, H. Evidence for two mechanisms for DNA unwinding catalyzed by DNA helicases. J. Biol. Chem. 254, 11343–11350 (1979)
Dillingham, M. S., Soultanas, P., Wiley, P., Webb, M. R. & Wigley, D. B. Defining the roles of individual residues in the single-stranded DNA binding site of PcrA helicase. Proc. Natl Acad. Sci. USA 98, 8381–8387 (2001)
Bianco, P. R. & Kowalczykowski, S. C. Translocation step size and mechanism of the RecBC DNA helicase. Nature 405, 368–372 (2000)
Korangy, F. & Julin, D. A. Efficiency of ATP hydrolysis and DNA unwinding by the RecBC enzyme from Escherichia coli. Biochemistry 33, 9552–9560 (1994)
Schultz, D. W., Taylor, A. F. & Smith, G. R. Escherichia coli RecBC pseudorevertants lacking Chi recombinational hotspot activity. J. Bacteriol. 155, 664–680 (1983)
Arnold, D. A., Bianco, P. R. & Kowalczykowski, S. C. The reduced levels of Chi recognition exhibited by the RecBC1004D enzyme reflect its recombination defect in vivo. J. Biol. Chem. 273, 16476–16486 (1998)
Arnold, D. A., Handa, N., Kobayashi, I. & Kowalczykowski, S. C. A novel, 11 nucleotide variant of Chi, Chi*: One of a class of sequences defining the Escherichia coli recombination hotspot Chi. J. Mol. Biol. 300, 469–479 (2000)
Kulkarni, A. & Julin, D. A. Specific inhibition of the E. coli RecBCD enzyme by Chi sequences in single-stranded oligodeoxynucleotides. Nucleic Acids Res. 32, 3672–3682 (2004)
Palas, K. M. & Kushner, S. R. Biochemical and physical properties of exonuclease V from Escherichia coli. J. Biol. Chem. 265, 3447–3454 (1990)
Churchill, J. J. & Kowalczykowski, S. C. Identification of the RecA protein-loading domain of RecBCD enzyme. J. Mol. Biol. 297, 537–542 (2000)
Kabsch, W. Automatic processing of rotation diffraction data from crystals of initially unknown symmetry and cell constants. J. Appl. Crystallogr. 26, 795–800 (1993)
Weeks, C. M. & Miller, R. Optimising Shake-and-bake for proteins. Acta Crystallogr. D 55, 492–500 (1999)
de La Fortelle, E. & Bricogne, G. Maximum-likelihood heavy-atom parameter refinement for the multiple isomorphous replacement and multiwavelength anomalous diffraction methods. Methods Enzymol. 276, 472–494 (1997)
Collaborative Computing Project No. 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Brunger, A. T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 904–925 (1998)
Sanner, M. F., Spehner, J. C. & Olson, A. J. Reduced surface: an efficient way to compute molecular surfaces. Biopolymers 38, 305–320 (1996)
Merritt, E. A. & Bacon, D. J. Raster3D: Photorealistic molecular graphics. Methods Enzymol. 277, 505–524 (1997)
Acknowledgements
We thank V. Ramakrishnan and J. Lowe for advice on the use and supply of tantalum bromide clusters, S. Halford for discussions about nucleases, C. Flensburg for advice on using SHARP and a pre-release version of the program, V. Dearing for oligonucleotide synthesis and purification, and D. Hall and E. Gordon for time and assistance on ESRF beamlines. This work was supported by Cancer Research UK (D.B.W.), NIH (S.C.K.) and a Wellcome Trust Travelling Research Fellowship (M.S.D.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Rights and permissions
About this article
Cite this article
Singleton, M., Dillingham, M., Gaudier, M. et al. Crystal structure of RecBCD enzyme reveals a machine for processing DNA breaks. Nature 432, 187–193 (2004). https://doi.org/10.1038/nature02988
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature02988
This article is cited by
-
A point mutation in recC associated with subclonal replacement of carbapenem-resistant Klebsiella pneumoniae ST11 in China
Nature Communications (2023)
-
Auxiliary ATP binding sites support DNA unwinding by RecBCD
Nature Communications (2022)
-
Proximate and ultimate causes of the bactericidal action of antibiotics
Nature Reviews Microbiology (2021)
-
Chi hotspot control of RecBCD helicase-nuclease by long-range intramolecular signaling
Scientific Reports (2020)
-
Quantification of very low-abundant proteins in bacteria using the HaloTag and epi-fluorescence microscopy
Scientific Reports (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.