Abstract
If a common ancestor of all living humans is defined as an individual who is a genealogical ancestor of all present-day people, the most recent common ancestor (MRCA) for a randomly mating population would have lived in the very recent past1,2,3. However, the random mating model ignores essential aspects of population substructure, such as the tendency of individuals to choose mates from the same social group, and the relative isolation of geographically separated groups. Here we show that recent common ancestors also emerge from two models incorporating substantial population substructure. One model, designed for simplicity and theoretical insight, yields explicit mathematical results through a probabilistic analysis. A more elaborate second model, designed to capture historical population dynamics in a more realistic way, is analysed computationally through Monte Carlo simulations. These analyses suggest that the genealogies of all living humans overlap in remarkable ways in the recent past. In particular, the MRCA of all present-day humans lived just a few thousand years ago in these models. Moreover, among all individuals living more than just a few thousand years earlier than the MRCA, each present-day human has exactly the same set of genealogical ancestors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Wachter, K. W. in Genealogical Demography (eds Dyke, B. & Morrill, W. T.) 85–93 (Academic, New York, 1980)
Chang, J. T. Recent common ancestors of all present-day individuals. Adv. Appl. Probab. 31, 1002–1026, 1027–1038 (1999)
Derrida, B., Manrubia, S. C. & Zanette, D. H. On the genealogy of a population of biparental individuals. J. Theor. Biol. 203, 303–315 (2000)
Ingman, M., Kaessmann, H., Pääbo, S. & Gyllensten, U. Mitochondrial genome variation and the origin of modern humans. Nature 408, 708–713 (2000)
Thomson, R., Pritchard, J. K., Shen, P., Oefner, P. J. & Feldman, M. W. Recent common ancestry of human Y chromosomes: Evidence from DNA sequence data. Proc. Natl Acad. Sci. USA 97, 7360–7365 (2000)
Hudson, R. R. in Oxford Surveys of Evolutionary Biology (eds Harvey, P. H. & Partridge, L.) 1–44 (Oxford Univ. Press, New York, 1990)
Stannard, D. E. American Holocaust: Columbus and the Conquest of the New World (Oxford Univ. Press, New York, 1992)
US National Office of Vital Statistics, Death Rates by Age, Race, and Sex, United States, 1900–1953, Vital Statistics—Special Reports Vol. 43 (US Government Printing Office, Washington DC, 1956)
Pletcher, S. D. Model fitting and hypothesis testing for age-specific mortality data. J. Evol. Biol. 12, 430–439 (1999)
Ohno, S. The Malthusian parameter of ascents: What prevents the exponential increase of one's ancestors? Proc. Natl Acad. Sci. USA 93, 15276–15278 (1996)
Derrida, B., Manrubia, S. C. & Zanette, D. H. Distribution of repetitions of ancestors in genealogical trees. Physica A 281, 1–16 (2000)
Wiuf, C. & Hein, J. On the number of ancestors to a DNA sequence. Genetics 147, 1459–1468 (1997)
Jones, R. Tasmanian archaeology: Establishing the sequences. Ann. Rev. Anthropol. 24, 423–446 (1995)
Fitzhugh, W. W. & Chausonnet, V. (eds) Crossroads of Continents: Cultures of Siberia and Alaska (Smithsonian Institution Press, Washington DC, 1988)
Bonné-Tamir, B. et al. Maternal and paternal lineages of the Samaritan isolate: Mutation rates and time to most recent common male ancestor. Ann. Hum. Genet. 67, 153–164 (2003)
Morton, N. E., Harris, D. E., Yee, S. & Lew, R. Pingelap and Mokil atolls: Migration. Am. J. Hum. Genet. 23, 339–349 (1971)
Hoerder, D. Cultures in Contact: World Migrations in the Second Millennium (Duke Univ. Press, Durham, North Carolina, 2002)
Zerjal, T. et al. The genetic legacy of the Mongols. Am. J. Hum. Genet. 72, 717–721 (2003)
Weiss, K. M. & Maruyama, T. Archeology, population genetics and studies of human racial ancestry. Am. J. Phys. Anthropol. 44, 31–50 (1976)
Ward, R. H. & Neel, J. V. Gene frequencies and microdifferentiation among the Makiritare indians. IV. A comparison of a genetic network with ethnohistory and migration matrices; a new index of genetic isolation. Am. J. Hum. Genet. 22, 538–561 (1970)
Jorde, L. B. in Current Developments in Anthropological Genetics (eds Mielke, J. H. & Crawford, M. H.) 135–208 (Plenum, New York, 1980)
Notohara, M. The coalescent and the genealogical process in geographically structured populations. J. Math. Biol. 29, 59–75 (1990)
Wilkinson-Herbots, H. M. Genealogy and subpopulation differentiation under various models of population structure. J. Math. Biol. 37, 535–585 (1998)
Hey, J. & Machado, C. A. The study of structured populations—new hope for a difficult and divided science. Nature Rev. Genet. 4, 535–543 (2003)
Acknowledgements
The research of D.L.T.R. was supported by the National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Supplementary Methods A
This file contains additional Methods (Further explanation and derivations of mathematical results) and an extra reference. (PDF 115 kb)
Supplementary Methods B
This file contains additional Methods (further details of the computational model), Supplementary Figure 1, Supplementary Table 1 and extra references. (PDF 196 kb)
Rights and permissions
About this article
Cite this article
Rohde, D., Olson, S. & Chang, J. Modelling the recent common ancestry of all living humans. Nature 431, 562–566 (2004). https://doi.org/10.1038/nature02842
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature02842
This article is cited by
-
The persistent homology of genealogical networks
Applied Network Science (2023)
-
Overcoming bioethical, legal, and hereditary barriers to mitochondrial replacement therapy in the USA
Journal of Assisted Reproduction and Genetics (2019)
-
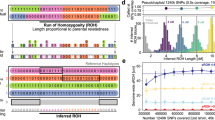
Runs of homozygosity: windows into population history and trait architecture
Nature Reviews Genetics (2018)
-
The Y chromosome as the most popular marker in genetic genealogy benefits interdisciplinary research
Human Genetics (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.