Abstract
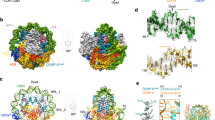
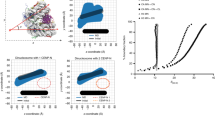
Mammalian centromeres are not defined by a consensus DNA sequence. In all eukaryotes a hallmark of functional centromeres—both normal ones and those formed aberrantly at atypical loci—is the accumulation of centromere protein A (CENP-A), a histone variant that replaces H3 in centromeric nucleosomes1,2,3,4,5,6,7. Here we show using deuterium exchange/mass spectrometry coupled with hydrodynamic measures that CENP-A and histone H4 form sub-nucleosomal tetramers that are more compact and conformationally more rigid than the corresponding tetramers of histones H3 and H4. Substitution into histone H3 of the domain of CENP-A responsible for compaction is sufficient to direct it to centromeres. Thus, the centromere-targeting domain of CENP-A confers a unique structural rigidity to the nucleosomes into which it assembles, and is likely to have a role in maintaining centromere identity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Cleveland, D. W., Mao, Y. & Sullivan, K. F. Centromeres and kinetochores: from epigenetics to mitotic checkpoint signaling. Cell 112, 407–421 (2003)
Schueler, M. G., Higgins, A. W., Rudd, M. K., Gustashaw, K. & Willard, H. F. Genomic and genetic definition of a functional human centromere. Science 294, 109–115 (2001)
Ohzeki, J., Nakano, M., Okada, T. & Masumoto, H. CENP-B box is required for de novo centromere chromatin assembly on human alphoid DNA. J. Cell Biol. 159, 765–775 (2002)
Rudd, M. K., Mays, R. W., Schwartz, S. & Willard, H. F. Human artificial chromosomes with alpha satellite-based de novo centromeres show increased frequency of nondisjunction and anaphase lag. Mol. Cell. Biol. 23, 7689–7697 (2003)
Sullivan, B. A., Blower, M. D. & Karpen, G. H. Determining centromere identity: cyclical stories and forking paths. Nature Rev. Genet. 2, 584–596 (2001)
Depinet, T. W. et al. Characterization of neo-centromeres in marker chromosomes lacking detectable alpha-satellite DNA. Hum. Mol. Genet. 6, 1195–1204 (1997)
Du Sart, D. et al. A functional neo-centromere formed through activation of a latent human centromere and consisting of non-alpha-satellite DNA. Nature Genet. 16, 144–153 (1997)
Stoler, S., Keith, K. C., Curnick, K. E. & Fitzgerald-Hayes, M. A mutation in CSE4, an essential gene encoding a novel chromatin-associated protein in yeast, causes chromosome nondisjunction and cell cycle arrest at mitosis. Genes Dev. 9, 573–586 (1995)
Buchwitz, B. J., Ahmad, K., Moore, L. L., Roth, M. B. & Henikoff, S. A histone-H3-like protein in C. elegans. Nature 401, 547–548 (1999)
Howman, E. V. et al. Early disruption of centromeric chromatin organization in centromere protein A (Cenpa) null mice. Proc. Natl Acad. Sci. USA 97, 1148–1153 (2000)
Takahashi, K., Chen, E. S. & Yanagida, M. Requirement of Mis6 centromere connector for localizing a CENP-A-like protein in fission yeast. Science 288, 2215–2219 (2000)
Blower, M. D. & Karpen, G. H. The role of Drosophila CID in kinetochore formation, cell-cycle progression and heterochromatin interactions. Nature Cell Biol. 3, 730–739 (2001)
Goshima, G., Kiyomitsu, T., Yoda, K. & Yanagida, M. Human centromere chromatin protein hMis12, essential for equal segregation, is independent of CENP-A loading pathway. J. Cell Biol. 160, 25–39 (2003)
Blower, M. D., Sullivan, B. A. & Karpen, G. H. Conserved organization of centromeric chromatin in flies and humans. Dev. Cell 2, 319–330 (2002)
Choo, K. H. Domain organization at the centromere and neocentromere. Dev. Cell 1, 165–177 (2001)
Earnshaw, W. C. & Rothfield, N. Identification of a family of human centromere proteins using autoimmune sera from patients with scleroderma. Chromosoma 91, 313–321 (1985)
Palmer, D. K. & Margolis, R. L. Kinetochore components recognized by human autoantibodies are present on mononucleosomes. Mol. Cell. Biol. 5, 173–186 (1985)
Hoofnagle, A. N., Resing, K. A. & Ahn, N. G. Protein analysis by hydrogen exchange mass spectrometry. Annu. Rev. Biophys. Biomol. Struct. 32, 1–25 (2003)
Hamuro, Y. et al. Dynamics of cAPK type IIβ activation revealed by enhanced amide H/2H exchange mass spectrometry (DXMS). J. Mol. Biol. 327, 1065–1076 (2003)
Richmond, T. J. & Davey, C. A. The structure of DNA in the nucleosome core. Nature 423, 145–150 (2003)
Glowczewski, L., Yang, P., Kalashnikova, T., Santisteban, M. S. & Smith, M. M. Histone-histone interactions and centromere function. Mol. Cell. Biol. 20, 5700–5711 (2000)
Shelby, R. D., Vafa, O. & Sullivan, K. F. Assembly of CENP-A into centromeric chromatin requires a cooperative array of nucleosomal DNA contact sites. J. Cell Biol. 136, 501–513 (1997)
Vermaak, D., Hayden, H. S. & Henikoff, S. Centromere targeting element within the histone fold domain of Cid. Mol. Cell. Biol. 22, 7553–7561 (2002)
Ahmad, K. & Henikoff, S. The histone variant H3.3 marks active chromatin by replication-independent nucleosome assembly. Mol. Cell 9, 1191–1200 (2002)
Shelby, R. D., Monier, K. & Sullivan, K. F. Chromatin assembly at kinetochores is uncoupled from DNA replication. J. Cell Biol. 151, 1113–1118 (2000)
Haushalter, K. A. & Kadonaga, J. T. Chromatin assembly by DNA-translocating motors. Nature Rev. Mol. Cell Biol. 4, 613–620 (2003)
Luger, K., Mader, A. W., Richmond, R. K., Sargent, D. F. & Richmond, T. J. Crystal structure of the nucleosome core particle at 2.8 Å resolution. Nature 389, 251–260 (1997)
Luger, K., Rechsteiner, T. J. & Richmond, T. J. Preparation of nucleosome core particle from recombinant histones. Methods Enzymol. 304, 3–19 (1999)
Eickbush, T. H. & Moudrianakis, E. N. The histone core complex: an octamer assembled by two sets of protein-protein interactions. Biochemistry 17, 4955–4964 (1978)
Earnshaw, W. C. et al. Molecular cloning of cDNA for CENP-B, the major human centromere autoantigen. J. Cell Biol. 104, 817–829 (1987)
Acknowledgements
We thank J. Kahana, J. Shah and K. Sullivan for reagents, and I. Cheeseman, B. Cottrell, P. Dyer, C. Gessner, F. Gordon, J. Kim, S. McBryant, D. Pantazatos and S. W. Englander for advice and technical assistance over the course of this study. We also thank J. Shah for comments on the manuscript. This research was supported by grants from the NIH to D.W.C and V.L.W., BioStar and Life Sciences Informatics grants from the University of California and ExSar Corporation to V.L.W., and by postdoctoral fellowships from the American Cancer Society (B.E.B.) and the NIH (D.R.F.). Salary support for D.W.C. was provided by the Ludwig Institute for Cancer Research.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
V.L.W. has an equity interest in ExSar Corporation.
Supplementary information
Supplementary Figure S1
Replacing particular CENP-A residues with the corresponding residues from histone H3 disrupts centromere targeting. (PDF 525 kb)
Supplementary Figure S2
A model for the epigenetic maintenance of centromere identity via specialized, CENP-A-containing chromatin. (PDF 159 kb)
Supplementary Figure Legends
Legends for Supplementary Figs S1 and S2. (PDF 47 kb)
Supplementary Methods
A detailed description of protein fragmentation mapping, deuterium exchange, and analysis of exchange profiles. (PDF 66 kb)
Rights and permissions
About this article
Cite this article
Black, B., Foltz, D., Chakravarthy, S. et al. Structural determinants for generating centromeric chromatin. Nature 430, 578–582 (2004). https://doi.org/10.1038/nature02766
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature02766
This article is cited by
-
Establishment of an Agrobacterium-mediated genetic transformation and CRISPR/Cas9-mediated mutagenesis of haploid inducer genes in Pak-choi plants (Brassica rapa ssp. chinensis)
Plant Biotechnology Reports (2024)
-
Uncovering natural allelic and structural variants of OsCENH3 gene by targeted resequencing and in silico mining in genus Oryza
Scientific Reports (2023)
-
Characterization of centromeric DNA of Gossypium anomalum reveals sequence-independent enrichment dynamics of centromeric repeats
Chromosome Research (2023)
-
Decoding allelic diversity, transcript variants and transcriptional complexity of CENH3 gene in Brassica oleracea var. botrytis
Protoplasma (2023)
-
Epigenetic regulation of centromere function
Cellular and Molecular Life Sciences (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.