Abstract
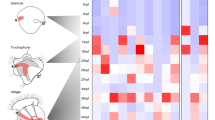
Tunicate embryos and larvae have small cell numbers and simple anatomical features in comparison with other chordates, including vertebrates. Although they branch near the base of chordate phylogenetic trees1, their degree of divergence from the common chordate ancestor remains difficult to evaluate. Here we show that the tunicate Oikopleura dioica has a complement of nine Hox genes in which all central genes are lacking but a full vertebrate-like set of posterior genes is present. In contrast to all bilaterians studied so far, Hox genes are not clustered in the Oikopleura genome. Their expression occurs mostly in the tail, with some tissue preference, and a strong partition of expression domains in the nerve cord, in the notochord and in the muscle. In each tissue of the tail, the anteroposterior order of Hox gene expression evokes spatial collinearity, with several alterations. We propose a relationship between the Hox cluster breakdown, the separation of Hox expression domains, and a transition to a determinative mode of development.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Wada, H. Evolutionary history of free-swimming and sessile lifestyles in urochordates as deduced from 18S rDNA molecular phylogeny. Mol. Biol. Evol. 15, 1189–1194 (1998)
Finnerty, J. R. The origins of axial patterning in the metazoa: how old is bilateral symmetry? Int. J. Dev. Biol. 47, 523–529 (2003)
Balavoine, G., de Rosa, R. & Adoutte, A. Hox clusters and bilaterian phylogeny. Mol. Phylogenet. Evol. 24, 366–373 (2002)
Von Allmen, G. et al. Splits in fruitfly Hox gene complexes. Nature 380, 116 (1996)
Burglin, T. R. & Ruvkun, G. The Caenorhabditis elegans homeobox gene cluster. Curr. Opin. Genet. Dev. 3, 615–620 (1993)
Akam, M. Hox and HOM: homologous gene clusters in insects and vertebrates. Cell 57, 347–349 (1989)
Ferrier, D. E., Minguillon, C., Holland, P. W. & Garcia-Fernandez, J. The amphioxus Hox cluster: deuterostome posterior flexibility and Hox14. Evol. Dev. 2, 284–293 (2000)
Powers, T. P. & Amemiya, C. T. Evidence for a Hox14 paralog group in vertebrates. Curr. Biol. 14, R183–R184 (2004)
Dehal, P. et al. The draft genome of Ciona intestinalis: insights into chordate and vertebrate origins. Science 298, 2157–2167 (2002)
Spagnuolo, A. et al. Unusual number and genomic organization of Hox genes in the tunicate Ciona intestinalis. Gene 309, 71–79 (2003)
Seo, H. C. et al. Miniature genome in the marine chordate Oikopleura dioica. Science 294, 2506 (2001)
Boutanaev, A. M., Kalmykova, A. I., Shevelyov, Y. Y. & Nurminsky, D. I. Large clusters of co-expressed genes in the Drosophila genome. Nature 420, 666–669 (2002)
Roy, P. J., Stuart, J. M., Lund, J. & Kim, S. K. Chromosomal clustering of muscle-expressed genes in Caenorhabditis elegans. Nature 418, 975–979 (2002)
Gionti, M. et al. Cihox5, a new Ciona intestinalis Hox-related gene, is involved in regionalization of the spinal cord. Dev. Genes Evol. 207, 515–523 (1998)
Locascio, A. et al. Patterning the ascidian nervous system: structure, expression and transgenic analysis of the CiHox3 gene. Development 126, 4737–4748 (1999)
Nagatomo, K. & Fujiwara, S. Expression of Raldh2, Cyp26 and Hox-1 in normal and retinoic acid-treated Ciona intestinalis embryos. Gene Expr. Patterns 3, 273–277 (2003)
Welsch, U. & Storch, V. Zur Feinstruktur der Chorda dorsalis niederer Chordaten Dendrodoa grossularia (v. Beneden) und Oikopleura dioica (Fol.). Z. Zellforsch. Mikrosk. Anat. 93, 547–559 (1969)
Lacalli, T. C. Tunicate tails, stolons, and the origin of the vertebrate trunk. Biol. Rev. Camb. Phil. Soc. 74, 177–198 (1999)
Ferrier, D. E. & Holland, P. W. Ciona intestinalis ParaHox genes: evolution of Hox/ParaHox cluster integrity, developmental mode, and temporal colinearity. Mol. Phylogenet. Evol. 24, 412–417 (2002)
van der Hoeven, F., Zakany, J. & Duboule, D. Gene transpositions in the HoxD complex reveal a hierarchy of regulatory controls. Cell 85, 1025–1035 (1996)
Averof, M. & Akam, M. Hox genes and the diversification of insect and crustacean body plans. Nature 376, 420–423 (1995)
Kmita, M. & Duboule, D. Organizing axes in time and space; 25 years of colinear tinkering. Science 301, 331–333 (2003)
Cowing, D. & Kenyon, C. Correct Hox gene expression established independently of position in Caenorhabditis elegans. Nature 382, 353–356 (1996)
Aboobaker, A. A. & Blaxter, M. L. Hox gene loss during dynamic evolution of the nematode cluster. Curr. Biol. 13, 37–40 (2003)
Spada, F. et al. Molecular patterning of the oikoplastic epithelium of the larvacean tunicate Oikopleura dioica. J. Biol. Chem. 276, 20624–20632 (2001)
Swofford, D. L. PAUP* 4.0.: Phylogenetic Analysis Using Parsimony (*and other methods), Version 4.0b10 (Sinauer, Sunderland, Massachusetts, 2002)
Schmidt, H. A., Strimmer, K., Vingron, M. & von Haeseler, A. TREE-PUZZLE: maximum likelihood phylogenetic analysis using quartets and parallel computing. Bioinformatics 18, 502–504 (2002)
Bassham, S. & Postlethwait, J. Brachyury (T) expression in embryos of a larvacean urochordate, Oikopleura dioica, and the ancestral role of T. Dev. Biol. 220, 322–332 (2000)
Acknowledgements
We thank A. Adoutte for advice throughout the course of this work; R. Aasland, M. Schartl, T. Stach and E. Thompson for their comments at several stages of manuscript preparation; and the personnel of the Oikopleura culture facility for technical support. Funding was provided by the Research Council of Norway and the University of Bergen in the frame of the Sars Centre – EMBL partnership contract.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Supplementary Figure 1
Coverage of Oikopleura genes by whole-genome shotgun data, measured by the coverage of a non-redundant collection of expressed sequence tags. (DOC 35 kb)
Supplementary Figure 2
Tandem duplication of the Hox4 gene. (JPG 58 kb)
Rights and permissions
About this article
Cite this article
Seo, HC., Edvardsen, R., Maeland, A. et al. Hox cluster disintegration with persistent anteroposterior order of expression in Oikopleura dioica. Nature 431, 67–71 (2004). https://doi.org/10.1038/nature02709
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature02709
This article is cited by
-
Non-collinear Hox gene expression in bivalves and the evolution of morphological novelties in mollusks
Scientific Reports (2021)
-
Transcriptomic analysis provides insights into candidate genes and molecular pathways involved in growth of Manila clam Ruditapes philippinarum
Functional & Integrative Genomics (2021)
-
The Chinese mitten crab genome provides insights into adaptive plasticity and developmental regulation
Nature Communications (2021)
-
Hox gene expression during development of the phoronid Phoronopsis harmeri
EvoDevo (2020)
-
Conservative route to genome compaction in a miniature annelid
Nature Ecology & Evolution (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.