Abstract
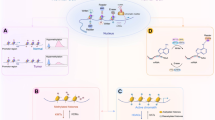
Epigenetic mechanisms, which involve DNA and histone modifications, result in the heritable silencing of genes without a change in their coding sequence. The study of human disease has focused on genetic mechanisms, but disruption of the balance of epigenetic networks can cause several major pathologies, including cancer, syndromes involving chromosomal instabilities, and mental retardation. The development of new diagnostic tools might reveal other diseases that are caused by epigenetic alterations. Great potential lies in the development of ‘epigenetic therapies’ — several inhibitors of enzymes controlling epigenetic modifications, specifically DNA methyltransferases and histone deacetylases, have shown promising anti-tumorigenic effects for some malignancies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Holliday, R. & Pugh, J. E. DNA modification mechanisms and gene activity during development. Science 187, 226–232 (1975).
Riggs, A. D. X inactivation, differentiation, and DNA methylation. Cytogenet. Cell Genet. 14, 9–25 (1975).
Takai, D. & Jones, P. A. Comprehensive analysis of CpG islands in human chromosomes 21 and 22. Proc. Natl Acad. Sci. USA 99, 3740–3745 (2002).
Jones, P. A. & Baylin, S. B. The fundamental role of epigenetic events in cancer. Nature Rev. Genet. 3, 415–428 (2002).
Lachner, M. & Jenuwein, T. The many faces of histone lysine methylation. Curr. Opin. Cell Biol. 14, 286–298 (2002).
Fischle, W., Wang, Y. & Allis, C. D. Binary switches and modification cassettes in histone biology and beyond. Nature 425, 475–479 (2003).
Strahl, B. D. & Allis, C. D. The language of covalent histone modifications. Nature 403, 41–45 (2000).
Tamaru, H. & Selker, E. U. A histone H3 methyltransferase controls DNA methylation in Neurospora crassa. Nature 414, 277–283 (2001).
Jackson, J. P., Lindroth, A. M., Cao, X. & Jacobsen, S. E. Control of CpNpG DNA methylation by the KRYPTONITE histone H3 methyltransferase. Nature 416, 556–560 (2002).
Malagnac, F., Bartee, L. & Bender, J. An Arabidopsis SET domain protein required for maintenance but not establishment of DNA methylation. EMBO J. 21, 6842–6852 (2002).
Johnson, L., Cao, X. & Jacobsen, S. Interplay between two epigenetic marks. DNA methylation and histone H3 lysine 9 methylation. Curr. Biol. 12, 1360–1367 (2002).
Soppe, W. J. et al. DNA methylation controls histone H3 lysine 9 methylation and heterochromatin assembly in Arabidopsis. EMBO J. 21, 6549–6559 (2002).
Tariq, M. et al. Erasure of CpG methylation in Arabidopsis alters patterns of histone H3 methylation in heterochromatin. Proc. Natl Acad. Sci. USA 100, 8823–8827 (2003).
Lehnertz, B. et al. Suv39h-mediated histone H3 lysine 9 methylation directs DNA methylation to major satellite repeats at pericentric heterochromatin. Curr. Biol. 13, 1192–1200 (2003).
Nan, X. et al. Transcriptional repression by the methyl-CpG-binding protein MeCP2 involves a histone deacetylase complex. Nature 393, 386–389 (1998).
Fuks, F. et al. The methyl-CpG-binding protein MeCP2 links DNA methylation to histone methylation. J. Biol. Chem. 278, 4035–4040 (2003).
Fuks, F., Burgers, W. A., Brehm, A., Hughes-Davies, L. & Kouzarides, T. DNA methyltransferase Dnmt1 associates with histone deacetylase activity. Nature Genet. 24, 88–91 (2000).
Hall, I. M. et al. Establishment and maintenance of a heterochromatin domain. Science 297, 2232–2237 (2002).
Volpe, T. A. et al. Regulation of heterochromatic silencing and histone H3 lysine-9 methylation by RNAi. Science 297, 1833–1837 (2002).
Zilberman, D., Cao, X. & Jacobsen, S. E. ARGONAUTE4 control of locus-specific siRNA accumulation and DNA and histone methylation. Science 299, 716–719 (2003).
Rougeulle, C. & Heard, E. Antisense RNA in imprinting: spreading silence through Air. Trends Genet. 18, 434–437 (2002).
Panning, B. & Jaenisch, R. RNA and the epigenetic regulation of X chromosome inactivation. Cell 93, 305–308 (1998).
Tufarelli, C. et al. Transcription of antisense RNA leading to gene silencing and methylation as a novel cause of human genetic disease. Nature Genet. 203, 157–165 (2003).
Okano, M., Bell, D. W., Haber, D. A. & Li, E. DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell 99, 247–257 (1999).
Amir, R. E. et al. Rett syndrome is caused by mutations in X-linked MECP2, encoding methyl-CpG-binding protein 2. Nature Genet. 23, 185–188 (1999).
Klose, R. & Bird, A. Molecular biology. MeCP2 repression goes nonglobal. Science 302, 793–795 (2003).
Chen, W. G. et al. Derepression of BDNF transcription involves calcium-dependent phosphorylation of MeCP2. Science 302, 885–889 (2003).
Martinowich, K. et al. DNA methylation-related chromatin remodeling in activity-dependent BDNF gene regulation. Science 302, 890–893 (2003).
Kane, M. F. et al. Methylation of the hMLH1 promoter correlates with lack of expression of hMLH1 in sporadic colon tumors and mismatch repair-defective human tumor cell lines. Cancer Res. 57, 808–811 (1997).
Gazzoli, I., Loda, M., Garber, J., Syngal, S. & Kolodner, R. D. A hereditary nonpolyposis colorectal carcinoma case associated with hypermethylation of the MLH1 gene in normal tissue and loss of heterozygosity of the unmethylated allele in the resulting microsatellite instability-high tumor. Cancer Res. 62, 3925–3928 (2002).
Suter, C. M., Martin, D. I. & Ward, R. L. Germline epimutation of MLH1 in individuals with multiple cancers. Nature Genet. advance online publication 4 April 2004 (doi:10.1038/ng1342).
Jones, P. A. & Laird, P. W. Cancer epigenetics comes of age. Nature Genet. 21, 163–167 (1999).
Hake, S. B., Xiao, A. & Allis, C. D. Linking the epigenetic ‘language’ of covalent histone modifications to cancer. Br. J. Cancer 90, 761–769 (2004).
Grignani, F. et al. Fusion proteins of the retinoic acid receptor-α recruit histone deacetylase in promyelocytic leukaemia. Nature 391, 815–818 (1998).
Jones, L. K. & Saha, V. Chromatin modification, leukaemia and implications for therapy. Br. J. Haematol. 118, 714–727 (2002).
Roberts, C. W. & Orkin, S. H. The SWI/SNF complex — chromatin and cancer. Nature Rev. Cancer 4, 133–142 (2004).
Wilson, V. L. & Jones, P. A. DNA methylation decreases in aging but not in immortal cells. Science 220, 1055–1057 (1983).
Richardson, B. C. Role of DNA methylation in the regulation of cell function: autoimmunity, aging and cancer. J. Nutr. 132, 2401S–2405S (2002).
Issa, J. P. CpG-island methylation in aging and cancer. Curr. Top. Microbiol. Immunol. 249, 101–118 (2000).
Beaudet, A. L. & Jiang, Y. H. A rheostat model for a rapid and reversible form of imprinting-dependent evolution. Am. J. Hum. Genet. 70, 1389–1397 (2002).
Laird, P. W. et al. Suppression of intestinal neoplasia by DNA hypomethylation. Cell 81, 197–205 (1995).
Sorm, F., Piskala, A., Cihak, A. & Vesely, J. 5-Azacytidine, a new, highly effective cancerostatic. Experientia 20, 202–203 (1964).
Constantinides, P. G., Jones, P. A. & Gevers, W. Functional striated muscle cells from non-myoblast precursors following 5-azacytidine treatment. Nature 267, 364–366 (1977).
Jones, P. A. & Taylor, S. M. Cellular differentiation, cytidine analogs and DNA methylation. Cell 20, 85–93 (1980).
Zhou, L. et al. Zebularine: a novel DNA methylation inhibitor that forms a covalent complex with DNA methyltransferases. J. Mol. Biol. 321, 591–599 (2002).
Michalowsky, L. A. & Jones, P. A. Differential nuclear protein binding to 5-azacytosine-containing DNA as a potential mechanism for 5-aza-2′-deoxycytidine resistance. Mol. Cell. Biol. 7, 3076–3083 (1987).
Issa, J. P. et al. Phase 1 study of low-dose prolonged exposure schedules of the hypomethylating agent 5-aza-2′-deoxycytidine (decitabine) in hematopoietic malignancies. Blood 103, 1635–1640 (2004).
Saunthararajah, Y. et al. Effects of 5-aza-2′-deoxycytidine on fetal hemoglobin levels, red cell adhesion, and hematopoietic differentiation in patients with sickle cell disease. Blood 102, 3865–3870 (2003).
Cheng, J. C. et al. Inhibition of DNA methylation and reactivation of silenced genes by zebularine. J. Natl Cancer Inst. 95, 399–409 (2003).
Lin, X. et al. Reversal of GSTP1 CpG island hypermethylation and reactivation of pi-class glutathione S-transferase (GSTP1) expression in human prostate cancer cells by treatment with procainamide. Cancer Res. 61, 8611–8616 (2001).
Fang, M. Z. et al. Tea polyphenol (–)-epigallocatechin-3-gallate inhibits DNA methyltransferase and reactivates methylation-silenced genes in cancer cell lines. Cancer Res. 63, 7563–7570 (2003).
Pina, I. C. et al. Psammaplins from the sponge Pseudoceratina purpurea: inhibition of both histone deacetylase and DNA methyltransferase. J. Org. Chem. 68, 3866–3873 (2003).
Yan, L. et al. Specific inhibition of DNMT1 by antisense oligonucleotides induces re-expression of estrogen receptor-α (ER) in ER-negative human breast cancer cell lines. Cancer Biol. Ther. 2, 552–556 (2003).
Xiao, H., Hasegawa, T. & Isobe, K. Both Sp1 and Sp3 are responsible for p21waf1 promoter activity induced by histone deacetylase inhibitor in NIH3T3 cells. J. Cell. Biochem. 73, 291–302 (1999).
Marks, P. A., Miller, T. & Richon, V. M. Histone deacetylases. Curr. Opin. Pharmacol. 3, 344–351 (2003).
Jahangeer, S., Elliott, R. M. & Henneberry, R. C. β-Adrenergic receptor induction in HeLa cells: synergistic effect of 5-azacytidine and butyrate. Biochem. Biophys. Res. Commun. 108, 1434–1440 (1982).
Ginder, G. D., Whitters, M. J. & Pohlman, J. K. Activation of a chicken embryonic globin gene in adult erythroid cells by 5-azacytidine and sodium butyrate. Proc. Natl Acad. Sci. USA 81, 3954–3958 (1984).
Cameron, E. E., Bachman, K. E., Myohanen, S., Herman, J. G. & Baylin, S. B. Synergy of demethylation and histone deacetylase inhibition in the re-expression of genes silenced in cancer. Nature Genet. 21, 103–107 (1999).
Suzuki, H. et al. A genomic screen for genes upregulated by demethylation and histone deacetylase inhibition in human colorectal cancer. Nature Genet. 31, 141–149 (2002).
Yamashita, K. et al. Pharmacologic unmasking of epigenetically silenced tumor suppressor genes in esophageal squamous cell carcinoma. Cancer Cell 2, 485–495 (2002).
Belinsky, S. A. et al. Inhibition of DNA methylation and histone deacetylation prevents murine lung cancer. Cancer Res. 63, 7089–7093 (2003).
Claus, R. & Lubbert, M. Epigenetic targets in hematopoietic malignancies. Oncogene 22, 6489–6496 (2003).
Plumb, J. A., Strathdee, G., Sludden, J., Kaye, S. B. & Brown, R. Reversal of drug resistance in human tumor xenografts by 2′-deoxy-5-azacytidine-induced demethylation of the hMLH1 gene promoter. Cancer Res. 60, 6039–6044 (2000).
Karpf, A. R. & Jones, D. A. Reactivating the expression of methylation silenced genes in human cancer. Oncogene 21, 5496–5503 (2002).
Weber, J. et al. Expression of the MAGE-1 tumor antigen is up-regulated by the demethylating agent 5-aza-2′-deoxycytidine. Cancer Res. 54, 1766–1771 (1994).
Laird, P. W. The power and the promise of DNA methylation markers. Nature Rev. Cancer 3, 253–266 (2003).
Liang, G., Gonzales, F. A., Jones, P. A., Orntoft, T. F. & Thykjaer, T. Analysis of gene induction in human fibroblasts and bladder cancer cells exposed to the methylation inhibitor 5-aza-2′-deoxycytidine. Cancer Res. 62, 961–966 (2002).
Mohandas, T., Sparkes, R. S. & Shapiro, L. J. Reactivation of an inactive X human chromosome: evidence for X inactivation by DNA methylation. Science 211, 393–396 (1981).
Wolf, S. F. & Migeon, B. R. Studies of X chromosome DNA methylation in normal human cells. Nature 295, 667–671 (1982).
Eversole-Cire, P. et al. Activation of an imprinted Igf 2 gene in mouse somatic cell cultures. Mol. Cell. Biol. 13, 4928–4938 (1993).
Jackson-Grusby, L., Laird, P. W., Magge, S. N., Moeller, B. J. & Jaenisch, R. Mutagenicity of 5-aza-2′-deoxycytidine is mediated by the mammalian DNA methyltransferase. Proc. Natl Acad. Sci. USA 94, 4681–4685 (1997).
Carr, B. I., Rahbar, S., Asmeron, Y., Riggs, A. & Winberg, C. D. Carcinogenicity and haemoglobin synthesis induction by cytidine analogues. Br. J. Cancer 57, 395–402 (1988).
Sato, N. et al. Frequent hypomethylation of multiple genes overexpressed in pancreatic ductal adenocarcinoma. Cancer Res. 63, 4158–4166 (2003).
Yang, A. S., Estecio, M. R., Garcia-Manero, G., Kantarjian, H. M. & Issa, J. P. Comment on ‘Chromosomal instability and tumors promoted by DNA hypomethylation’ and ‘Induction of tumors in nice by genomic hypomethylation’. Science 302, 1153 (2003).
Lubbert, M. et al. Cytogenetic responses in high-risk myelodysplastic syndrome following low-dose treatment with the DNA methylation inhibitor 5-aza-2′-deoxycytidine. Br. J. Haematol. 114, 349–357 (2001).
Karpf, A. R., Moore, B. C., Ririe, T. O. & Jones, D. A. Activation of the p53 DNA damage response pathway after inhibition of DNA methyltransferase by 5-aza-2′-deoxycytidine. Mol. Pharmacol. 59, 751–757 (2001).
Jackson-Grusby, L. et al. Loss of genomic methylation causes p53-dependent apoptosis and epigenetic deregulation. Nature Genet. 27, 31–39 (2001).
Peterson, E. J., Bogler, O. & Taylor, S. M. p53-mediated repression of DNA methyltransferase 1 expression by specific DNA binding. Cancer Res. 63, 6579–6582 (2003).
Juttermann, R., Li, E. & Jaenisch, R. Toxicity of 5-aza-2′-deoxycytidine to mammalian cells is mediated primarily by covalent trapping of DNA methyltransferase rather than DNA demethylation. Proc. Natl Acad. Sci. USA 91, 11797–11801 (1994).
Esteller, M., Corn, P. G., Baylin, S. B. & Herman, J. G. A gene hypermethylation profile of human cancer. Cancer Res. 61, 3225–3229 (2001).
Cui, H. et al. Loss of IGF2 imprinting: a potential marker of colorectal cancer risk. Science 299, 1753–1755 (2003).
Gibbons, R. J. & Higgs, D. R. Molecular-clinical spectrum of the ATR-X syndrome. Am. J. Med. Genet. 97, 204–212 (2000).
Oostra, B. A. & Willemsen, R. The X chromosome and fragile X mental retardation. Cytogenet. Genome Res. 99, 257–264 (2002).
Ehrlich, M. The ICF syndrome, a DNA methyltransferase 3B deficiency and immunodeficiency disease. Clin. Immunol. 109, 17–28 (2003).
Nicholls, R. D., Saitoh, S. & Horsthemke, B. Imprinting in Prader–Willi and Angelman syndromes. Trends Genet. 14, 194–200 (1998).
Goldstone, A. P. Prader–Willi syndrome: advances in genetics, pathophysiology and treatment. Trends Endocrinol. Metab. 15, 12–20 (2004).
Maher, E. R. & Reik, W. Beckwith–Wiedemann syndrome: imprinting in clusters revisited. J. Clin. Invest. 105, 247–252 (2000).
Feinberg, A. P. & Tycko, B. The history of cancer epigenetics. Nature Rev. Cancer 4, 143–153 (2004).
Soejima, H. et al. Silencing of imprinted CDKN1C gene expression is associated with loss of CpG and histone H3 lysine 9 methylation at DMR-LIT1 in esophageal cancer. Oncogene published online 8 March 2004 (doi:10.1038/sj.onc.1207576).
Ausio, J., Levin, D. B., De Amorim, G. V., Bakker, S. & Macleod, P. M. Syndromes of disordered chromatin remodeling. Clin. Genet. 64, 83–95 (2003).
Yoder, J. A., Walsh, C. P. & Bestor, T. H. Cytosine methylation and the ecology of intragenomic parasites. Trends Genet. 13, 335–340 (1997).
Nguyen, C. T., Gonzales, F. A. & Jones, P. A. Altered chromatin structure associated with methylation-induced gene silencing in cancer cells: correlation of accessibility, methylation, MeCP2 binding and acetylation. Nucleic Acids Res. 29, 4598–4606 (2001).
Bachman, K. E. et al. Histone modifications and silencing prior to DNA methylation of a tumor suppressor gene. Cancer Cell 3, 89–95 (2003).
El-Osta, A. & Wolffe, A. P. DNA methylation and histone deacetylation in the control of gene expression: basic biochemistry to human development and disease. Gene Expr. 9, 63–75 (2000).
Nguyen, C. T. et al. Histone H3-lysine 9 methylation is associated with aberrant gene silencing in cancer cells and is rapidly reversed by 5-aza-2′-deoxycytidine. Cancer Res. 62, 6456–6461 (2002).
Acknowledgements
We thank C. Nguyen for his help with the figures, and A. Yang and J. Rice for critical reading of the manuscript. This work was supported by the National Cancer Institute and the Max Kade Foundation.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
P.A.J. is a shareholder and consultant for Epigenomics AG and a consultant for SuperGen Inc., which is developing decitabine as a cancer treatment.
Rights and permissions
About this article
Cite this article
Egger, G., Liang, G., Aparicio, A. et al. Epigenetics in human disease and prospects for epigenetic therapy. Nature 429, 457–463 (2004). https://doi.org/10.1038/nature02625
Issue Date:
DOI: https://doi.org/10.1038/nature02625
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.