Abstract
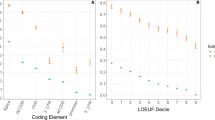
New alleles become fixed owing to random drift of nearly neutral mutations or to positive selection of substantially advantageous mutations1,2,3. After decades of debate, the fraction of fixations driven by selection remains uncertain4,5,6,7,8,9. Within 9,390 genes, we analysed 28,196 codons at which rat and mouse differ from each other at two nucleotide sites and 1,982 codons with three differences. At codons where rat–mouse divergence involved two non-synonymous substitutions, both of them occurred in the same lineage, either rat or mouse, in 64% of cases; however, independent substitutions would occur in the same lineage with a probability of only 50%. All three non-synonymous substitutions occurred in the same lineage for 46% of codons, instead of the 25% expected. Furthermore, comparison of 12 pairs of prokaryotic genomes also shows clumping of multiple non-synonymous substitutions in the same lineage. This pattern cannot be explained by correlated mutation or episodes of relaxed negative selection, but instead indicates that positive selection acts at many sites of rapid, successive amino acid replacement.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Kimura, M. The Neutral Theory of Molecular Evolution (Cambridge Univ. Press, Cambridge, 1983)
Gillespie, J. H. The Causes of Molecular Evolution (Oxford Univ. Press, Oxford, 1991)
Ohta, T. Near-neutrality in evolution of genes and gene regulation. Proc. Natl Acad. Sci. USA 99, 16134–16137 (2002)
Smith, N. G. C. & Eyre-Walker, A. Adaptive protein evolution in Drosophila. Nature 415, 1022–1024 (2002)
Fay, J. C., Wyckoff, G. J. & Wu, C.-I. Testing the neutral theory of molecular evolution with genomic data from Drosophila. Nature 415, 1024–1026 (2002)
Bustamante, C. D. et al. The cost of inbreeding in Arabidopsis. Nature 416, 531–534 (2002)
Eyre-Walker, A. Changing effective population size and the McDonald-Kreitman test. Genetics 162, 2017–2024 (2002)
Yang, Z. H. Inference of selection from multiple species alignments. Curr. Opin. Genet. Dev. 12, 688–694 (2002)
Anisimova, M., Nielsen, R. & Yang, Z. H. Effect of recombination on the accuracy of the likelihood method for detecting positive selection at amino acid sites. Genetics 164, 1229–1236 (2003)
Makalowski, W. & Boguski, M. S. Evolutionary parameters of the transcribed mammalian genome: an analysis of 2,820 orthologous rodent and human sequences. Proc. Natl Acad. Sci. USA 95, 9407–9412 (1998)
Kondrashov, A. S. Direct estimates of human per nucleotide mutation rates at 20 loci causing Mendelian diseases. Hum. Mutat. 21, 12–27 (2003)
Nachman, M. W. & Crowell, S. L. Estimate of the mutation rate per nucleotide in humans. Genetics 156, 297–304 (2000)
Fitch, W. M. & Markowitz, E. An improved method for determining codon variability in a gene and its application to the rate of fixation of mutations in evolution. Biochem. Genet. 4, 579–593 (1970)
Huelsenbeck, J. P. Testing a covariotide model of DNA substitution. Mol. Biol. Evol. 19, 698–707 (2002)
Miyata, T., Miyazawa, S. & Yasunaga, T. Two types of amino acid substitutions in protein evolution. J. Mol. Evol. 12, 219–236 (1979)
Bustamante, C. D., Nielsen, R. & Hartl, D. L. A maximum likelihood method for analyzing pseudogene evolution: implications for silent site evolution in humans and rodents. Mol. Biol. Evol. 19, 110–127 (2002)
Hellman, I. et al. Selection on human genes as revealed by comparisons to chimpanzee cDNA. Genome Res. 13, 831–837 (2003)
Wilbur, W. J. On the PAM matrix model of protein evolution. Mol. Biol. Evol. 2, 434–447 (1985)
International Human Genome Sequencing Consortium. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001)
Mouse Genome Sequencing Consortium. Initial sequencing and comparative analysis of the mouse genome. Nature 420, 520–562 (2002)
Rat Genome Sequencing Project Consortium. Genome sequence of the Brown Norway rat yields insights into mammalian evolution. Nature 428, 493–521 (2004)
Tatusov, R. L., Koonin, E. V. & Lipman, D. J. A genomic perspective on protein families. Science 278, 631–637 (1997)
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997)
Thompson, J. D., Higgins, D. G. & Gibson, T. J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994)
Yang, Z. H. PAML: a program package for phylogenetic analysis by maximum likelihood. Comput. Appl. Biosci. 13, 555–556 (1997)
Acknowledgements
We thank N. Bierne for a number of suggestions. G.A.B. was supported by a BWF graduate fellowship. S.S. was supported by Genome Canada Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Supplementary Figures and Discussion
Analysis of substitutions at adjacent codons, of possible biased misidentification of the rat-mouse common ancestor, and of fluctuating negative selection.
Rights and permissions
About this article
Cite this article
Bazykin, G., Kondrashov, F., Ogurtsov, A. et al. Positive selection at sites of multiple amino acid replacements since rat–mouse divergence. Nature 429, 558–562 (2004). https://doi.org/10.1038/nature02601
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature02601
This article is cited by
-
Improved inference of site-specific positive selection under a generalized parametric codon model when there are multinucleotide mutations and multiple nonsynonymous rates
BMC Evolutionary Biology (2019)
-
Crossing fitness valleys via double substitutions within codons
BMC Biology (2019)
-
mtProtEvol: the resource presenting molecular evolution analysis of proteins involved in the function of Vertebrate mitochondria
BMC Evolutionary Biology (2019)
-
Multinucleotide mutations cause false inferences of lineage-specific positive selection
Nature Ecology & Evolution (2018)
-
Compensatory evolution in mitochondrial tRNAs navigates valleys of low fitness
Nature (2010)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.