Abstract
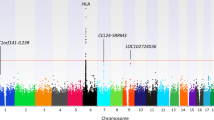
Leprosy is caused by Mycobacterium leprae and affects about 700,000 individuals each year1. It has long been thought that leprosy has a strong genetic component2, and recently we mapped a leprosy susceptibility locus to chromosome 6 region q25–q26 (ref. 3). Here we investigate this region further by using a systematic association scan of the chromosomal interval most likely to harbour this leprosy susceptibility locus. In 197 Vietnamese families we found a significant association between leprosy and 17 markers located in a block of approx. 80 kilobases overlapping the 5′ regulatory region shared by the Parkinson's disease gene PARK2 and the co-regulated gene PACRG. Possession of as few as two of the 17 risk alleles was highly predictive of leprosy. This was confirmed in a sample of 975 unrelated leprosy cases and controls from Brazil in whom the same alleles were strongly associated with leprosy. Variants in the regulatory region shared by PARK2 and PACRG therefore act as common risk factors for leprosy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
WHO, Leprosy. Global situation. Wkly Epidemiol. Rec. 77, 1–8 (2002)
Casanova, J. L. & Abel, L. Genetic dissection of immunity to mycobacteria: The human model. Annu. Rev. Immunol. 20, 581–620 (2002)
Mira, M. T. et al. Chromosome 6q25 is linked to susceptibility to leprosy in a Vietnamese population. Nature Genet. 33, 412–415 (2003)
Dupuis, J. & Siegmund, D. Statistical methods for mapping quantitative trait loci from a dense set of markers. Genetics 151, 373–386 (1999)
West, A. B., Lockhart, P. J., O'Farell, C. & Farrer, M. J. Identification of a novel gene linked to parkin via a bi-directional promoter. J. Mol. Biol. 326, 11–19 (2003)
Jacobson, R. R. & Krahenbuhl, J. L. Leprosy. Lancet 353, 655–660 (1999)
Kitada, T. et al. Mutations in the parkin gene cause autosomal recessive juvenile parkinsonism. Nature 392, 605–608 (1998)
Giasson, B. I. & Lee, V. M. Parkin and the molecular pathways of Parkinson's disease. Neuron 31, 885–888 (2001)
Imai, Y. et al. A product of the human gene adjacent to parkin is a component of Lewy bodies and suppresses Pael receptor-induced cell death. J. Biol. Chem. 278, 51901–51910 (2003)
Ridley, D. S. & Jopling, W. H. Classification of leprosy according to immunity. A five-group system. Int. J. Lepr. Other Mycobact. Dis. 34, 255–273 (1966)
Excoffier, L. & Slatkin, M. Maximum-likelihood estimation of molecular haplotype frequencies in a diploid population. Mol. Biol. Evol. 12, 921–927 (1995)
Abecasis, G. R. & Cookson, W. O. GOLD—graphical overview of linkage disequilibrium. Bioinformatics 16, 182–183 (2000)
Niu, T., Qin, Z. S., Xu, X. & Liu, J. S. Bayesian haplotype inference for multiple linked single-nucleotide polymorphisms. Am. J. Hum. Genet. 70, 157–169 (2002)
Horvath, S., Xu, X. & Laird, N. M. The family based association test method: Strategies for studying general genotype–phenotype associations. Eur. J. Hum. Genet. 9, 301–306 (2001)
Lake, S. L., Blacker, D. & Laird, N. M. Family-based tests of association in the presence of linkage. Am. J. Hum. Genet. 67, 1515–1525 (2000)
Thomson, G. Mapping disease genes: Family-based association studies. Am. J. Hum. Genet. 57, 487–498 (1995)
Schaid, D. J. & Rowland, C. Use of parents, sibs, and unrelated controls for detection of associations between genetic markers and disease. Am. J. Hum. Genet. 63, 1492–1506 (1998)
Devlin, B. & Roeder, K. Genomic control for association studies. Biometrics 55, 997–1004 (1999)
Acknowledgements
We thank all patients, their families and population controls that participated in this study; J. L. Casanova for support, suggestions and critical reading of the manuscript; T. H. M. Ottenhoff for his help and support in the Schwann cell work; and A. Sammak, C. Darmond-Zwaig, Y. Renaud, M. Girard, N. Martin, A. Villeneuve, D. Frappier and A. Verville for technical assistance. This study was supported by a grant from the Canadian Institutes of Health Research to E.S. The genotyping core at the McGill University and Genome Québec Innovation Centre is supported by the Canadian Genetic Diseases Network and Genome Québec, and the Schwann cell work in Leiden is supported by the Netherlands Leprosy Foundation. M.T.M. is supported by a graduate fellowship of the Brazilian Coordenacao de Aperfeicoamento de Pessoal de Nivel Superior (CAPES). A.A. and L.A. are supported in part by grants from Fondation Banque Nationale de Paris–Paribas, Fondation pour la Recherche Médicale and Fondation Schlumberger. T.J.H. and E.S. are Canadian Institutes of Health Research Investigators.Authors' contributions L.A. and E.S. share senior authorship.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
41586_2004_BFnature02326_MOESM1_ESM.xls
Supplementary Web Table 1: This file lists all oligonucleotides used for genotyping. Indicated are the method of genotyping, the lab name of the oligonucleotide, the nucleotide sequence, the type of nucleotide change, and the minor allele frequency observed among the Vietnamese families. ni: MAF < 5%, failed: genotyping assay did not produce reliable results. (XLS 106 kb)
41586_2004_BFnature02326_MOESM2_ESM.xls
Supplementary Web Table 2: This file lists all SNPs discovered by comparative sequencing of the 80 kb LD block in 12 unrelated individuals. Indicated are the SNP ID, the chromosome 6 position of a specific SNP on the July 2003 map, and the corresponding genotypes for each of the 12 individuals. (XLS 57 kb)
Rights and permissions
About this article
Cite this article
Mira, M., Alcaïs, A., Van Thuc, N. et al. Susceptibility to leprosy is associated with PARK2 and PACRG. Nature 427, 636–640 (2004). https://doi.org/10.1038/nature02326
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature02326
This article is cited by
-
Cell Biology of Parkin: Clues to the Development of New Therapeutics for Parkinson’s Disease
CNS Drugs (2022)
-
Genome-wide association study of Buruli ulcer in rural Benin highlights role of two LncRNAs and the autophagy pathway
Communications Biology (2020)
-
Interferon-γ signaling synergizes with LRRK2 in neurons and microglia derived from human induced pluripotent stem cells
Nature Communications (2020)
-
Human Genetic Susceptibility of Leprosy Recurrence
Scientific Reports (2020)
-
Human genetics of Buruli ulcer
Human Genetics (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.