Abstract
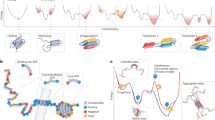
The manner in which a newly synthesized chain of amino acids transforms itself into a perfectly folded protein depends both on the intrinsic properties of the amino-acid sequence and on multiple contributing influences from the crowded cellular milieu. Folding and unfolding are crucial ways of regulating biological activity and targeting proteins to different cellular locations. Aggregation of misfolded proteins that escape the cellular quality-control mechanisms is a common feature of a wide range of highly debilitating and increasingly prevalent diseases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Vendruscolo, M., Zurdo, J., MacPhee, C. E. & Dobson, C. M. Protein folding and misfolding: a paradigm of self-assembly and regulation in complex biological systems. Phil. Trans. R. Soc. Lond. 361, 1205–1222 (2003).
Radford, S. E. & Dobson, C. M. From computer simulations to human disease: emerging themes in protein folding. Cell 97, 291–298 (1999).
Dobson, C. M., Sali, A. & Karplus, M. Protein folding: a perspective from theory and experiment. Angew. Chem. Int. Ed. Eng. 37, 868–893 (1998).
Wolynes, P. G., Onuchic, J. N. & Thirumalai, D. Navigating the folding routes. Science 267, 1619–1620 (1995).
Dill, K. A. & Chan, H. S. From Levinthal to pathways to funnels. Nature Struct. Biol. 4, 10–19 (1997).
Dinner, A. R., Sali, A., Smith, L. J., Dobson, C. M. & Karplus, M. Understanding protein folding via free energy surfaces from theory and experiment. Trends Biochem. Sci. 25, 331–339 (2000).
Baldwin, R. L. Protein folding: matching speed and stability. Nature 369, 183–184 (1994).
Eaton, W. A., Munoz, V., Thompson, P. A., Henry, E. R. & Hofrichter, J. Kinetics and dynamics of loops, α-helices, β-hairpins, and fast-folding proteins. Acc. Chem. Res. 31, 745–753 (1998).
Snow, C. D., Nguyen, H., Pande, V. S. & Gruebele, M. Absolute comparison of simulated and experimental protein-folding dynamics. Nature 420, 102–106 (2002).
Yang, W. Y. & Gruebele, M. Folding at the speed limit. Nature 423, 193–197 (2003).
Mayor, U. et al. The complete folding pathway of a protein from nanoseconds to microseconds. Nature 421, 863–867 (2003).
Plaxco, K. W., Simons, K. T. & Baker, D. Contact order, transition state placement and the refolding rates of single domain proteins. J. Mol. Biol. 277, 985–994 (1998).
Schuler, B., Lipman, E. A. & Eaton, W. A. Probing the free-energy surface for protein folding with single-molecule fluorescence spectroscopy. Nature 419, 743–747 (2002).
Fersht, A. R. Structure and Mechanism in Protein Science: A Guide to Enzyme Catalysis and Protein Folding (W.H. Freeman, New York, 1999).
Fersht, A. R. Transition-state structure as a unifying basis in protein-folding mechanisms: contact order, chain topology, stability, and the extended nucleus mechanism. Proc. Natl Acad. Sci. USA 97, 1525–1529 (2000).
Shea, J. E. & Brooks, C. L. From folding surfaces to folding proteins: a review and assessment of simulation studies of protein folding and unfolding. Annu. Rev. Phys. Chem. 52, 499–535 (2001).
Fersht, A. R. & Daggett, V. Protein folding and unfolding at atomic resolution. Cell 108, 573–582 (2002).
Vendruscolo, M., Paci, E., Dobson, C. M. & Karplus, M. Three key residues form a critical contact network in a transition state for protein folding. Nature 409, 641–646 (2001).
Makarov, D. E. & Plaxco, K. W. The topomer search model: a simple, quantitative theory of two-state protein folding kinetics. Protein Sci. 12, 17–26 (2003).
Baker, D. A surprising simplicity to protein folding. Nature 405, 39–42 (2000).
Roder, H. & Colon, W. Kinetic role of early intermediates in protein folding. Curr. Opin. Struct. Biol. 7, 15–28 (1997).
Sanchez, I. E. & Kiefhaber, T. Evidence for sequential barriers and obligatory intermediates in apparent two-state protein folding. J. Mol. Biol. 325, 367–376 (2003).
Khan, F., Chuang, J. I., Gianni, S. & Fersht, A. R. The kinetic pathway of folding of barnase. J. Mol. Biol. 333, 169–186 (2003).
Vendruscolo, M., Paci, E., Karplus, M. & Dobson, C. M. Structures and relative free energies of partially folded states of proteins. Proc. Natl Acad. Sci. USA 100, 14817–14821 (2003).
Cheung, M. S., Garcia, A. E. & Onuchic, J. N. Protein folding mediated by solvation: water expulsion and formation of the hydrophobic core occur after the structural collapse. Proc. Natl Acad. Sci. USA 99, 685–690 (2002).
Hardesty, B. & Kramer, G. Folding of a nascent peptide on the ribosome. Prog. Nucleic Acid Res. Mol. Biol. 66, 41–66 (2001).
Bukau, B. & Horwich, A. L. The Hsp70 and Hsp60 chaperone machines. Cell 92, 351–366 (1998).
Hartl, F. U. & Hayer-Hartl, M. Molecular chaperones in the cytosol: from nascent chain to folded protein. Science 295, 1852–1858 (2002).
Ellis, R. J. Macromolecular crowding: an important but neglected aspect of the intracellular environment. Curr. Opin. Struct. Biol. 11, 114–119 (2001).
Schiene, C. & Fischer, G. Enzymes that catalyse the restructuring of proteins. Curr. Opin. Struct. Biol. 10, 40–45 (2000).
Hammon, C. & Helenius, A. Quality control in the secretory pathway. Curr. Opin. Cell. Biol. 7, 523–529 (1995).
Kaufman, R. J. et al. The unfolded protein response in nutrient sensing and differentiation. Nature Rev. Mol. Cell Biol. 3, 411–421 (2002).
Wilson, M. R. & Easterbrook Smith, S. B. Clusterin is a secreted mammalian chaperone. Trends Biochem. Sci. 25, 95–98 (2000).
Schubert, U. et al. Rapid degradation of a large fraction of newly synthesized proteins by proteasomes. Nature 404, 770–774 (2000).
Bence, N. F., Sampat, R. M. & Kopito, R. R. Impairment of the ubiquitin–proteasome system by protein aggregation. Science 292, 1552–1555 (2001).
Thomas, P. J., Qu, B. H. & Pedersen, P. L. Defective protein folding as a basis of human disease. Trends Biochem. Sci. 20, 456–459 (1995).
Dobson, C. M. The structural basis of protein folding and its links with human disease. Phil. Trans. R. Soc. Lond. B 356, 133–145 (2001).
Horwich, A. Protein aggregation in disease: a role for folding intermediates forming specific multimeric interactions. J. Clin. Invest. 110, 1221–1232 (2002).
Bullock, A. N. & Fersht, A. R. Rescuing the functions of mutant p53. Nature Rev. Cancer 1, 68–76 (2001).
Tan, S. Y. & Pepys, M. B. Amyloidosis. Histopathology 25, 403–414 (1994).
Kelly, J. W. Alternative conformation of amyloidogenic proteins and their multi-step assembly pathways. Curr. Opin. Struct. Biol. 8, 101–106 (1998).
Sunde, M. & Blake, C. C. F. The structure of amyloid fibrils by electron microscopy and X-ray diffraction. Adv. Protein Chem. 50, 123–159 (1997).
Dobson, C. M. Protein misfolding, evolution and disease. Trends Biochem. Sci. 24, 329–332 (1999).
Fändrich, M. & Dobson, C. M. The behaviour of polyamino acids reveals an inverse side-chain effect in amyloid structure formation. EMBO J. 21, 5682–5690 (2002).
Jiménez, J. L. et al. Cryo-electron microscopy of an SH3 amyloid fibril and model of the molecular packing. EMBO J. 18, 815–821 (1999).
Petkova, A. T. et al. A structural model for Alzheimer's β-amyloid fibrils based on experimental constraints from solid state NMR. Proc. Natl Acad. Sci. USA 99, 16742–16747 (2002).
Chiti, F., Stefani, M., Taddei, N., Ramponi, G. & Dobson, C. M. Rationalization of mutational effects on protein aggregation rates. Nature 424, 805–808 (2003).
Caughey, B. & Lansbury, P. T. Jr. Protofibrils, pores, fibrils, and neurodegeneration: separating the responsible protein aggregates from the innocent bystanders. Annu. Rev. Neurosci. 26, 267–298 (2003).
Bitan, G. et al. Amyloid β-protein (Aβ) assembly: Aβ40 and Aβ42 oligomerize through distinct pathways. Proc. Natl Acad. Sci. USA 100, 330–335 (2003).
Nilsson, M. R., Driscoll, M. & Raleigh, D. P. Low levels of asparagine deamidation can have a dramatic effect on aggregation of amyloidogenic peptides: implications for the study of amyloid formation. Protein Sci. 11, 342–349 (2002).
Schlunegger, M. P., Bennett, M. J. & Eisenberg, D. Oligomer formation by 3D domain swapping: a model for protein assembly and misassembly. Adv. Protein Chem. 50, 61–122 (1997).
Bucciantini, M. et al. Inherent cytotoxicity of aggregates implies a common origin for protein misfolding diseases. Nature 416, 507–511 (2002).
Lashuel, H. A., Hartley, D., Petre, B. M., Walz, T. & Lansbury, P. T. Jr. Neurodegenerative disease: amyloid pores from pathogenic mutations. Nature 418, 291 (2002).
Dobson, C. M. Protein folding and disease: a view from the First Horizon Symposium. Nature Rev. Drug Discov. 2, 154–160 (2003).
True, H. L. & Lindquist, S. L. A yeast prion provides a mechanism for genetic variation and phenotypic diversity. Nature 407, 477–483 (2000).
Chapman, M. R. et al. Role of Escherichia coli curli operons in directing amyloid fiber formation. Science 295, 851–855 (2002).
Kelly, J. W. & Balch, W. E. Amyloid as a natural product. J. Cell Biol. 161, 461–462 (2003).
Broome, B. M. & Hecht, M. H. Nature disfavours sequences of alternating polar and non-polar amino acids: implications for amyloidogenesis. J. Mol. Biol. 296, 961–968 (2000).
Chiti, F. et al. Kinetic partitioning of protein folding and aggregation. Nature Struct. Biol. 9, 137–143 (2002).
Macario, A. J. L. & Macario, E. C. Sick chaperones and ageing: a perspective. Ageing Res. Rev. 1, 295–311 (2002).
Ramirez-Alvarado, M., Merkel, J. S. & Regan, L. A systematic exploration of the influence of the protein stability on amyloid fibril formation in vitro. Proc. Natl Acad. Sci. USA 97, 8979–8984 (2000).
Dumoulin, M. et al. A camelid antibody fragment inhibits amyloid fibril formation by human lysozyme. Nature 424, 783–788 (2003).
Prusiner, S. B. Prion diseases and the BSE crisis. Science 278, 245–251 (1997).
Taylor, J. P., Hardy, J. & Fischbeck, K. H. Toxic proteins in neurodegenerative disease. Science 296, 1991–1995 (2002).
Walsh, D. M. et al. Naturally secreted oligomers of amyloid β protein potently inhibit hippocampal long-term potentiation in vivo. Nature 416, 535–539 (2002).
Kayed, R. et al. Common structure of soluble amyloid oligomers implies common mechanisms of pathogenesis. Science 300, 486–489 (2003).
Stefani, M. & Dobson, C. M. Protein aggregation and aggregate toxicity: new insights into protein folding, misfolding diseases and biological evolution. J. Mol. Med. 81, 678–699 (2003).
Sherman, M. Y. & Goldberg, A. L. Cellular defenses against unfolded proteins: a cell biologist thinks about neurodegenerative diseases. Neuron 29, 15–32 (2001).
Muchowski, P. J. et al. Hsp70 and Hsp40 chaperones can inhibit self-assembly of polyglutamine proteins into amyloid-like fibrils. Proc. Natl Acad. Sci. USA 97, 7841–7846 (2000).
Csermely, P. Chaperone overload is a possible contributor to 'civilization diseases'. Trends Genet. 17, 701–704 (2001).
Dobson, C. M. Getting out of shape—protein misfolding diseases. Nature 418, 729–730 (2002).
Acknowledgements
I should like to thank in particular the Wellcome Trust, the Leverhulme Trust and the UK Research Councils for generous support over many years, without whom my own research activities in this area of science could not have been carried out.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Dobson, C. Protein folding and misfolding. Nature 426, 884–890 (2003). https://doi.org/10.1038/nature02261
Issue Date:
DOI: https://doi.org/10.1038/nature02261
This article is cited by
-
Terahertz Time-Domain Spectroscopic (THz-TDS) Insights into Protein Deformation
Brazilian Journal of Physics (2024)
-
Construction of synthetic anti-fouling consortia: fouling control effects and polysaccharide degradation mechanisms
Microbial Cell Factories (2023)
-
Constructing custom thermodynamics using deep learning
Nature Computational Science (2023)
-
Data-mining unveils structure–property–activity correlation of viral infectivity enhancing self-assembling peptides
Nature Communications (2023)
-
A fluorescent multi-domain protein reveals the unfolding mechanism of Hsp70
Nature Chemical Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.