Abstract
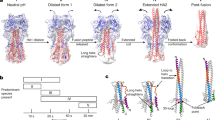
Fusion of biological membranes is mediated by specific lipid-interacting proteins that induce the formation and expansion of an initial fusion pore. Here we report the crystal structure of the ectodomain of the Semliki Forest virus fusion glycoprotein E1 in its low-pH-induced trimeric form. E1 adopts a folded-back conformation that, in the final post-fusion form of the full-length protein, would bring the fusion peptide loop and the transmembrane anchor to the same end of a stable protein rod. The observed conformation of the fusion peptide loop is compatible with interactions only with the outer leaflet of the lipid bilayer. Crystal contacts between fusion peptide loops of adjacent E1 trimers, together with electron microscopy observations, suggest that in an early step of membrane fusion, an intermediate assembly of five trimers creates two opposing nipple-like deformations in the viral and target membranes, leading to formation of the fusion pore.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Jahn, R., Lang, T. & Sudhof, T. C. Membrane fusion. Cell 112, 519–533 (2003)
Skehel, J. J. & Wiley, D. C. Receptor binding and membrane fusion in virus entry: the influenza hemagglutinin. Annu. Rev. Biochem. 69, 531–569 (2000)
Weissenhorn, W. et al. Structural basis for membrane fusion by enveloped viruses. Mol. Membr. Biol. 16, 3–9 (1999)
Danieli, T., Pelletier, S. L., Henis, Y. I. & White, J. M. Membrane fusion mediated by the influenza virus hemagglutinin requires the concerted action of at least three hemagglutinin trimers. J. Cell Biol. 133, 559–569 (1996)
Blumenthal, R., Sarkar, D. P., Durell, S., Howard, D. E. & Morris, S. J. Dilation of the influenza hemagglutinin fusion pore revealed by the kinetics of individual cell–cell fusion events. J. Cell Biol. 135, 63–71 (1996)
Markovic, I., Leikina, E., Zhukovsky, M., Zimmerberg, J. & Chernomordik, L. V. Synchronized activation and refolding of influenza hemagglutinin in multimeric fusion machines. J. Cell Biol. 155, 833–844 (2001)
Zimmerberg, J. & Chernomordik, L. V. Membrane fusion. Adv. Drug Deliv. Rev. 38, 197–205 (1999)
Blumenthal, R., Clague, M. J., Durell, S. R. & Epand, R. M. Membrane fusion. Chem. Rev. 103, 53–69 (2003)
Helenius, A., Kartenbeck, J., Simons, K. & Fries, E. On the entry of Semliki Forest virus into BHK-21 cells. J. Cell Biol. 84, 404–420 (1980)
Schlesinger, S. & Schlesinger, M. J. in Fields Virology (eds Knipe, D. M. & Howley, P. M.) 895–916 (Lippincott Williams and Wilkins, Philadelphia, 2001)
Lescar, J. et al. The fusion glycoprotein shell of Semliki Forest virus: an icosahedral assembly primed for fusogenic activation at endosomal pH. Cell 105, 137–148 (2001)
Zhang, W. et al. Placement of the structural proteins in Sindbis virus. J. Virol. 76, 11645–11658 (2002)
Rey, F. A., Heinz, F. X., Mandl, C., Kunz, C. & Harrison, S. C. The envelope glycoprotein from tick-borne encephalitis virus at 2Å resolution. Nature 375, 291–298 (1995)
Modis, Y., Ogata, S., Clements, D. & Harrison, S. C. A ligand-binding pocket in the dengue virus envelope glycoprotein. Proc. Natl Acad. Sci. USA 100, 6899–6901 (2003)
Wahlberg, J. M. & Garoff, H. Membrane fusion process of Semliki Forest virus I. Low pH-induced rearrangement in spike protein quaternary structure precedes virus penetration into cells. J. Cell Biol. 116, 339–348 (1992)
Kielian, M., Chatterjee, P. K., Gibbons, D. L. & Lu, Y. E. in Subcellular Biochemistry Vol. 34 Fusion of Biological Membranes and Related Problems (eds Hilderson, H. & Fuller, S.) 409–455 (Plenum, New York, 2000)
Klimjack, M. R., Jeffrey, S. & Kielian, M. Membrane and protein interactions of a soluble form of the Semliki Forest virus fusion protein. J. Virol. 68, 6940–6946 (1994)
Gibbons, D. L. & Kielian, M. Molecular dissection of the Semliki Forest virus homotrimer reveals two functionally distinct regions of the fusion protein. J. Virol. 76, 1194–1205 (2002)
Ahn, A., Gibbons, D. L. & Kielian, M. The fusion peptide of Semliki Forest virus associates with sterol-rich membrane domains. J. Virol. 76, 3267–3275 (2002)
Gibbons, D. L. et al. Visualization of the target-membrane-inserted fusion protein of Semliki Forest virus by combined electron microscopy and crystallography. Cell 114, 573–583 (2003)
Eckert, D. M. & Kim, P. S. Mechanisms of viral membrane fusion and its inhibition. Annu. Rev. Biochem. 70, 777–810 (2001)
Wahlberg, J. M., Bron, R., Wilschut, J. & Garoff, H. Membrane fusion of Semliki Forest virus involves homotrimers of the fusion protein. J. Virol. 66, 7309–7318 (1992)
Markosyan, R. M., Cohen, F. S. & Melikyan, G. B. HIV-1 envelope proteins complete their folding into six-helix bundles immediately after fusion pore formation. Mol. Biol. Cell 14, 926–938 (2003)
Kemble, G. W., Danieli, T. & White, J. M. Lipid-anchored influenza hemagglutinin promotes hemifusion, not complete fusion. Cell 76, 383–391 (1994)
Armstrong, R. T., Kushnir, A. S. & White, J. M. The transmembrane domain of influenza hemagglutinin exhibits a stringent length requirement to support the hemifusion to fusion transition. J. Cell Biol. 151, 425–437 (2000)
Bagai, S. & Lamb, R. A. Truncation of the COOH-terminal region of the paramyxovirus SV5 fusion protein leads to hemifusion but not complete fusion. J. Cell Biol. 135, 73–84 (1996)
Januszeski, M. M., Cannon, P. M., Chen, D., Rozenberg, Y. & Anderson, W. F. Functional analysis of the cytoplasmic tail of Moloney murine leukemia virus envelope protein. J. Virol. 71, 3613–3619 (1997)
Melikyan, G. B., Markosyan, R. M., Brener, S. A., Rozenberg, Y. & Cohen, F. S. Role of the cytoplasmic tail of ecotropic moloney murine leukemia virus Env protein in fusion pore formation. J. Virol. 74, 447–455 (2000)
Chernomordik, L., Frolov, V. A., Leikina, E., Bronk, P. & Zimmerberg, J. The pathway of membrane fusion catalyzed by influenza hemagglutinin: restriction of lipids, hemifusion, and lipidic fusion pore formation. J. Cell Biol. 140, 1369–1382 (1998)
Gaudin, Y. Rabies virus-induced membrane fusion pathway. J. Cell Biol. 150, 601–612 (2000)
Markosyan, R. M., Melikyan, G. B. & Cohen, F. S. Evolution of intermediates of influenza virus hemagglutinin-mediated fusion revealed by kinetic measurements of pore formation. Biophys. J. 80, 812–821 (2001)
Modis, Y., Ogata, S., Clements, D. & Harrison, S. C. Structure of the dengue virus envelope protein after membrane fusion. Nature 427, 313–319 (2004)
Bressanelli, S. et al. Structure of a flavivirus envelope glycoprotein in its low-pH-induced membrane fusion conformation. EMBO J. (in the press)
Gibbons, D. L. et al. Purification and crystallization reveal two types of interactions of the fusion protein homotrimer of Semliki Forest virus. J. Virol. (in the press)
Otwinowski, Z. & Minor, W. in Macromolecular Crystallography Part A (eds Carter, C. W. & Sweet, R. M.) 307–326 (Academic Press, London, 1997)
Terwilliger, T. C. & Berendzen, J. Automated MAD and MIR structure solution. Acta Crystallogr. D 55, 849–861 (1999)
de la Fortelle, E. & Bricogne, G. Maximum-likelihood heavy-atom parameter refinement for multiple isomorphous replacement and multiwavelength anomalous diffraction methods. Methods Enzymol. 276, 472–493 (1997)
Cowtan, K. dm: an automated procedure for phase improvement by density modification. Joint CCP4 ESF-EACBM Newslett. Protein Crystallogr. 31, 34–38 (1994)
Jones, T. A. & Kjeldgaard, M. in Macromolecular Crystallography Part B (eds Carter, C. W. & Sweet, R. M.) 173–208 (Academic Press, London, 1997)
Roussel, A. & Cambillaud, C. Silicon Graphics Geometry Partners Directory (Silicon Graphics, Mountain View, California, 1991)
Brünger, A. T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Carson, M. Ribbon models of macromolecules. J. Mol. Graph. 5, 103–106 (1987)
Acknowledgements
We thank S. Bressanelli, S. Duquerroy and P. Fernandez Varela for their help at different stages of this work; A. Ahn and A. Urian for help with virus and protein preparation; C. Schulze-Briese and T. Tomikazi for help during diffraction data collection; Y. Gaudin for critically reading the manuscript; and J. Navaza for helpful discussions. More than 80% of the data used to determine the crystal structure were collected at synchrotron beam line X06SA of the Swiss Light Source, Paul Scherrer Institut, Villigen, Switzerland. Other synchrotron sources used were beam lines ID14 and ID29 at the European Synchrotron Radiation Facility, Grenoble, France, and beam line BW7A at DESY, Hamburg, Germany. M.K. acknowledges support from the Public Health Service and from a Cancer Center Core Support Grant from the National Cancer Institute. F.A.R. acknowledges support from the CNRS and INRA, the SESAME Program of the Région Ile-de-France, the French Fondation pour la Recherche Médicale, the Association pour la Recherche contre le Cancer, the CNRS programs “Physique et Chimie du Vivant” and “Dynamique et réactivité des assemblages biologiques”, and the European Union ENhcV consortium. D.L.G. was supported by the Medical Scientist Training Program of the Albert Einstein College of Medicine, the Albert Cass Traveling Fellowship and the CNRS.Authors' contributions The crystallographic analyses reported in this paper were performed by F.A.R. and M.C.V.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Gibbons, D., Vaney, MC., Roussel, A. et al. Conformational change and protein–protein interactions of the fusion protein of Semliki Forest virus. Nature 427, 320–325 (2004). https://doi.org/10.1038/nature02239
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature02239
This article is cited by
-
A molecular understanding of alphavirus entry and antibody protection
Nature Reviews Microbiology (2023)
-
Identification of the flavivirus conserved residues in the envelope protein hinge region for the rational design of a candidate West Nile live-attenuated vaccine
npj Vaccines (2023)
-
Implication for alphavirus host-cell entry and assembly indicated by a 3.5Å resolution cryo-EM structure
Nature Communications (2018)
-
Entry of severe fever with thrombocytopenia syndrome virus
Virologica Sinica (2017)
-
Recent progress on chikungunya virus research
Virologica Sinica (2017)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.