Abstract
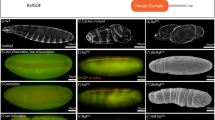
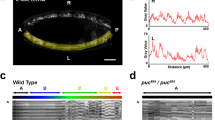
Drosophila thoracic mechanosensory bristles originate from cells that are singled out from ‘proneural’ groups of competent epithelial cells. Neural competence is restricted to individual sensory organ precursors (SOPs) by Delta/Notch-mediated ‘lateral inhibition’, whereas other cells in the proneural field adopt an epidermal fate. The precursors of the large macrochaetes differentiate separately from individual proneural clusters that comprise about 20–30 cells or as heterochronic pairs from groups of more than 100 cells1, whereas the precursors of the small regularly spaced microchaetes emerge from even larger proneural fields2. This indicates that lateral inhibition might act over several cell diameters; it was difficult to reconcile with the fact that the inhibitory ligand Delta is membrane-bound until the observation that SOPs frequently extend thin processes3,4 offered an attractive hypothesis. Here we show that the extension of these planar filopodia—a common attribute of wing imaginal disc cells—is promoted by Delta and that their experimental suppression reduces Notch signalling in distant cells and increases bristle density in large proneural groups, showing that these membrane specializations mediate long-range lateral inhibition.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Huang, F., Dambly-Chaudiere, C. & Ghysen, A. The emergence of sense organs in the wing disc of Drosophila. Development 111, 1087–1095 (1991)
Usui, K. & Kimura, K. Sequential emergence of the evenly spaced microchaetes on the notum of Drosophila. Roux's Arch. Dev. Biol. 203, 151–158 (1993)
Lai, E. C. & Rubin, G. M. neuralized functions cell-autonomously to regulate a subset of notch-dependent processes during adult Drosophila development. Dev. Biol. 231, 217–233 (2001)
Renaud, O. & Simpson, P. scabrous modifies epithelial cell adhesion and extends the range of lateral signalling during development of the spaced bristle pattern in Drosophila. Dev. Biol. 240, 361–376 (2001)
Wigglesworth, V. B. Local and general factors in the development of ‘pattern’ in Rhodnius prolixus Hemiptera. J. Exp. Biol. 17, 180–200 (1940)
Parks, A. L. & Muskavitch, M. A. Delta function is required for bristle organ determination and morphogenesis in Drosophila. Dev. Biol. 157, 484–496 (1993)
Jennings, B., Preiss, A., Delidakis, C. & Bray, S. The Notch signalling pathway is required for Enhancer of split bHLH protein expression during neurogenesis in the Drosophila embryo. Development 120, 3537–3548 (1994)
Lecourtois, M. & Schweisguth, F. The neurogenic suppressor of hairless DNA-binding protein mediates the transcriptional activation of the enhancer of split complex genes triggered by Notch signaling. Genes Dev. 9, 2598–2608 (1995)
Zeng, C., Younger-Shepherd, S., Jan, L. Y. & Jan, Y. N. Delta and Serrate are redundant Notch ligands required for asymmetric cell divisions within the Drosophila sensory organ lineage. Genes Dev. 12, 1086–1091 (1998)
Qi, H. et al. Processing of the notch ligand delta by the metalloprotease Kuzbanian. Science 283, 91–94 (1999)
Mishra-Gorur, K., Rand, M. D., Perez-Villamil, B. & Artavanis-Tsakonas, S. Down-regulation of Delta by proteolytic processing. J. Cell Biol. 159, 313–324 (2002)
Kooh, P. J., Fehon, R. G. & Muskavitch, M. A. Implications of dynamic patterns of Delta and Notch expression for cellular interactions during Drosophila development. Development 117, 493–507 (1993)
Sanson, B., Alexandre, C., Fascetti, N. & Vincent, J. P. Engrailed and hedgehog make the range of Wingless asymmetric in Drosophila embryos. Cell 98, 207–216 (1999)
Milan, M., Weihe, U., Perez, L. & Cohen, S. M. The LRR proteins capricious and Tartan mediate cell interactions during DV boundary formation in the Drosophila wing. Cell 106, 785–794 (2001)
Locke, M. & Huie, P. Epidermal feet in insect morphogenesis. Nature 293, 733–735 (1981)
Fehon, R. G. et al. Molecular interactions between the protein products of the neurogenic loci Notch and Delta, two EGF-homologous genes in Drosophila. Cell 61, 523–534 (1990)
Bretscher, A., Edwards, K. & Fehon, R. G. ERM proteins and merlin: integrators at the cell cortex. Nature Rev. Mol. Cell Biol. 3, 586–599 (2002)
Martin, M. et al. Ezrin NH2-terminal domain inhibits the cell extension activity of the COOH-terminal domain. J. Cell Biol. 128, 1081–1093 (1995)
Martin, M., Roy, C., Montcourrier, P., Sahuquet, A. & Mangeat, P. Three determinants in ezrin are responsible for cell extension activity. Mol. Biol. Cell 8, 1543–1557 (1997)
Meir, E., von Dassow, G., Munro, E. & Odell, G. M. Robustness, flexibility, and the role of lateral inhibition in the neurogenic network. Curr. Biol. 12, 778–786 (2002)
Turing, A. M. The chemical basis for morphogenesis. Phil. Trans. R. Soc. Lond. B 237, 37–72 (1952)
Bellaiche, Y., Gho, M., Kaltschmidt, J. A., Brand, A. H. & Schweisguth, F. Frizzled regulates localization of cell-fate determinants and mitotic spindle rotation during asymmetric cell division. Nature Cell Biol. 3, 50–57 (2001)
Lee, T. & Luo, L. Mosaic analysis with a repressible cell marker for studies of gene function in neuronal morphogenesis. Neuron 22, 451–461 (1999)
Hinz, U., Giebel, B. & Campos-Ortega, J. A. The basic-helix–loop–helix domain of Drosophila lethal of scute protein is sufficient for proneural function and activates neurogenic genes. Cell 76, 77–87 (1994)
Seugnet, L., Simpson, P. & Haenlin, M. Transcriptional regulation of Notch and Delta: requirement for neuroblast segregation in Drosophila. Development 124, 2015–2025 (1997)
Haenlin, M., Kunisch, M., Kramatschek, B. & Campos-Ortega, J. A. Genomic regions regulating early embryonic expression of the Drosophila neurogenic gene Delta. Mech. Dev. 47, 99–110 (1994)
Bender, L. B., Kooh, P. J. & Muskavitch, M. A. Complex function and expression of Delta during Drosophila oogenesis. Genetics 133, 967–978 (1993)
Lamb, R. F. et al. Essential functions of ezrin in maintenance of cell shape and lamellipodial extension in normal and transformed fibroblasts. Curr. Biol. 7, 682–688 (1997)
Andreoli, C., Martin, M., Le Borgne, R., Reggio, H. & Mangeat, P. Ezrin has properties to self-associate at the plasma membrane. J. Cell Sci. 107, 2509–2521 (1994)
Ramirez-Weber, F. A. & Kornberg, T. B. Cytonemes: cellular processes that project to the principal signaling center in Drosophila imaginal discs. Cell 97, 599–607 (1999)
Acknowledgements
This work was initiated in the unité INSERM 432, Université Montpellier II. We thank S. Baghdiguian, C. Dambly-Chaudière, A. Ghysen, S. Layalle, P.-H. Mangeat and A.-M. Martinez for discussions; C. Dambly-Chaudière, M. Haenlin, F. Schweisguth, J.-P. Vincent, the Bloomington Stock Center and the Developmental Studies Hybridoma Bank for fly stocks and reagents; N. Lautrédou-Audouy for help with confocal microscopy; F. Mérezègue for help with electron microscopy; E. Gazave and A. Sahuquet for help with morphometry and statistics; and C. Roy for critically reading the manuscript. C.d.J. and D.A. thank the researchers into Drosophila at the Institut de Génétique Humaine (Montpellier) for hospitality. This work was supported by grants to D.A. from the Association pour la Recherche contre le Cancer and the Université Montpellier II. C.d.J. was supported by the Fondation pour la Recherche Médicale.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Rights and permissions
About this article
Cite this article
de Joussineau, C., Soulé, J., Martin, M. et al. Delta-promoted filopodia mediate long-range lateral inhibition in Drosophila. Nature 426, 555–559 (2003). https://doi.org/10.1038/nature02157
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature02157
This article is cited by
-
Adjusting the range of cell–cell communication enables fine-tuning of cell fate patterns from checkerboard to engulfing
Journal of Mathematical Biology (2023)
-
Predictive model for cytoneme guidance in Hedgehog signaling based on Ihog- Glypicans interaction
Nature Communications (2022)
-
Regulatory mechanisms of cytoneme-based morphogen transport
Cellular and Molecular Life Sciences (2022)
-
Generation of extracellular morphogen gradients: the case for diffusion
Nature Reviews Genetics (2021)
-
A role for actomyosin contractility in Notch signaling
BMC Biology (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.