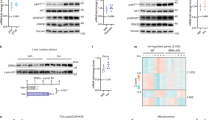
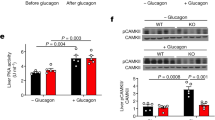
Abstract
Fasting triggers a series of hormonal cues that promote energy balance by inducing glucose output and lipid breakdown in the liver1. In response to pancreatic glucagon and adrenal cortisol, the cAMP-responsive transcription factor CREB activates gluconeogenic and fatty acid oxidation programmes by stimulating expression of the nuclear hormone receptor coactivator PGC-1 (refs 2–5). In parallel, fasting also suppresses lipid storage and synthesis (lipogenic) pathways1, but the underlying mechanism is unknown. Here we show that mice deficient in CREB activity have a fatty liver phenotype and display elevated expression of the nuclear hormone receptor PPAR-γ, a key regulator of lipogenic genes6,7. CREB inhibits hepatic PPAR-γ expression in the fasted state by stimulating the expression of the Hairy Enhancer of Split (HES-1) gene, a transcriptional repressor that is shown here to be a mediator of fasting lipid metabolism in vivo. The coordinate induction of PGC-1 and repression of PPAR-γ by CREB during fasting provides a molecular rationale for the antagonism between insulin and counter-regulatory hormones, and indicates a potential role for CREB antagonists as therapeutic agents in enhancing insulin sensitivity in the liver.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Saltiel, A. R. New perspectives into the molecular pathogenesis and treatment of type 2 diabetes. Cell 104, 517–529 (2001)
Herzig, S. et al. CREB regulates hepatic gluconeogenesis through the coactivator PGC-1. Nature 413, 179–183 (2001)
Puigserver, P. et al. Insulin-regulated hepatic gluconeogenesis through FOXO1–PGC-1α interaction. Nature 423, 550–555 (2003)
Vega, R. B., Huss, J. M. & Kelly, D. P. The coactivator PGC-1 cooperates with peroxisome proliferator-activated receptor α in transcriptional control of nuclear genes encoding mitochondrial fatty acid oxidation enzymes. Mol. Cell. Biol. 20, 1868–1876 (2000)
Yoon, J. C. et al. Control of hepatic gluconeogenesis through the transcriptional coactivator PGC-1. Nature 413, 131–138 (2001)
Yu, S. et al. Adipocyte-specific gene expression and adipogenic steatosis in the mouse liver due to peroxisome proliferator-activated receptor γ 1 (PPAR-γ1) overexpression. J. Biol. Chem. 278, 498–505 (2003)
Matsusue, K. et al. Liver-specific disruption of PPARγ in leptin-deficient mice improves fatty liver but aggravates diabetic phenotypes. J. Clin. Invest. 111, 737–747 (2003)
Rudolph, D. et al. Impaired fetal T cell development and perinatal lethality in mice lacking the cAMP response element binding protein. Proc. Natl Acad. Sci. USA 95, 4481–4486 (1998)
Horton, J. D. & Shimomura, I. Sterol regulatory element-binding proteins: activators of cholesterol and fatty acid biosynthesis. Curr. Opin. Lipidol. 10, 143–150 (1999)
Conkright, M. D. et al. Genome-wide analysis of CREB target genes reveals a core promoter requirement for cAMP responsiveness. Mol. Cell 11, 1101–1108 (2003)
Fajas, L. et al. The organization, promoter analysis, and expression of the human PPAR-γ gene. J. Biol. Chem. 272, 18779–18789 (1997)
Fajas, L. et al. Regulation of peroxisome proliferator-activated receptor γ expression by adipocyte differentiation and determination factor 1/sterol regulatory element binding protein 1: implications for adipocyte differentiation and metabolism. Mol. Cell. Biol. 19, 5495–5503 (1999)
Sasai, Y., Kageyama, R., Tagawa, Y., Shigemoto, R. & Nakanishi, S. Two mammalian helix–loop–helix factors structurally related to Drosophila hairy and Enhancer of split. Genes Dev. 6, 2620–2634 (1992)
Jarriault, S. et al. Signalling downstream of activated mammalian Notch. Nature 377, 355–358 (1995)
Gekakis, N. et al. Role of the CLOCK protein in the mammalian circadian mechanism. Science 280, 1564–1569 (1998)
Iso, T., Kedes, L. & Hamamori, Y. HES and HERP families: multiple effectors of the Notch signaling pathway. J. Cell. Physiol. 194, 237–255 (2003)
Sui, G. et al. A DNA vector-based RNAi technology to suppress gene expression in mammalian cells. Proc. Natl Acad. Sci. USA 99, 5515–5520 (2002)
Lee, Y. et al. Liporegulation in diet-induced obesity. The antisteatotic role of hyperleptinemia. J. Biol. Chem. 276, 5629–5635 (2001)
Koo, S. H. & Towle, H. C. Glucose regulation of mouse S(14) gene expression in hepatocytes. Involvement of a novel transcription factor complex. J. Biol. Chem. 275, 5200–5207 (2000)
Michael, L. F., Asahara, H., Shulman, A. I., Kraus, W. L. & Montminy, M. The phosphorylation status of a cyclic AMP-responsive activator is modulated via a chromatin-dependent mechanism. Mol. Cell. Biol. 20, 1596–1603 (2000)
Hirata, H. et al. Oscillatory expression of the bHLH factor Hes1 regulated by a negative feedback loop. Science 298, 840–843 (2002)
Canettieri, G. et al. Attenuation of a phosphorylation-dependent activator by an HDAC–PP1 complex. Nature Struct. Biol. 10, 175–181 (2003)
Daitoku, H., Yamagata, K., Matsuzaki, H., Hatta, M. & Fukamizu, A. Regulation of PGC-1 promoter activity by protein kinase B and the forkhead transcription factor FKHR. Diabetes 52, 642–649 (2003)
Takebayashi, K. et al. Structure, chromosomal locus, and promoter analysis of the gene encoding the mouse helix–loop–helix factor HES-1. Negative autoregulation through the multiple N box elements. J. Biol. Chem. 269, 5150–5156 (1994)
Orellana, S. A. & McKnight, G. S. Mutations in the catalytic subunit of cAMP-dependent protein kinase result in unregulated biological activity. Proc. Natl Acad. Sci. USA 89, 4726–4730 (1992)
McLarren, K. W. et al. The mammalian basic helix loop helix protein HES-1 binds to and modulates the transactivating function of the runt-related factor Cbfa1. J. Biol. Chem. 275, 530–538 (2000)
Nakajima, T., Uchida, C., Anderson, S. F., Parvin, J. D. & Montminy, M. Analysis of a cAMP-responsive activator reveals a two-component mechanism for transcriptional induction via signal-dependent factors. Genes Dev. 11, 738–747 (1997)
Acknowledgements
We thank K. Suter and L. Vera for performing injections; S. Stifani, R. Kageyama, J. Auwerx, Y. Shi and T. Sudo for providing reagents; and G. Schuetz for CREB knockout mice. We also thank R. Evans for reviewing the manuscript, and I. Verma for support. This work was supported by the NIH (M.M.), the American Diabetes Association, the Hillblom Foundation and the Deutsche Forschungsgemeinschaft (S.H.). F.G. is also supported by Dipartimento di Scienze Biomediche, Università di Sassari, Italy.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Rights and permissions
About this article
Cite this article
Herzig, S., Hedrick, S., Morantte, I. et al. CREB controls hepatic lipid metabolism through nuclear hormone receptor PPAR-γ. Nature 426, 190–193 (2003). https://doi.org/10.1038/nature02110
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature02110
This article is cited by
-
mir-98-5p regulates gluconeogenesis and lipogenesis by targeting PPP1R15B in hepatocytes
Journal of Cell Communication and Signaling (2023)
-
Chromogranin A and its derived peptides: potential regulators of cholesterol homeostasis
Cellular and Molecular Life Sciences (2023)
-
A propolis-derived small molecule ameliorates metabolic syndrome in obese mice by targeting the CREB/CRTC2 transcriptional complex
Nature Communications (2022)
-
Molecular signaling in pancreatic ductal metaplasia: emerging biomarkers for detection and intervention of early pancreatic cancer
Cellular Oncology (2022)
-
Unique adaptations in neonatal hepatic transcriptome, nutrient signaling, and one-carbon metabolism in response to feeding ethyl cellulose rumen-protected methionine during late-gestation in Holstein cows
BMC Genomics (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.