Abstract
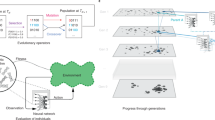
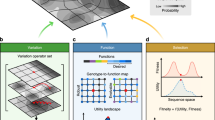
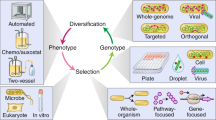
A long-standing challenge to evolutionary theory has been whether it can explain the origin of complex organismal features. We examined this issue using digital organisms—computer programs that self-replicate, mutate, compete and evolve. Populations of digital organisms often evolved the ability to perform complex logic functions requiring the coordinated execution of many genomic instructions. Complex functions evolved by building on simpler functions that had evolved earlier, provided that these were also selectively favoured. However, no particular intermediate stage was essential for evolving complex functions. The first genotypes able to perform complex functions differed from their non-performing parents by only one or two mutations, but differed from the ancestor by many mutations that were also crucial to the new functions. In some cases, mutations that were deleterious when they appeared served as stepping-stones in the evolution of complex features. These findings show how complex functions can originate by random mutation and natural selection.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Darwin, C. On the Origin of Species by Means of Natural Selection (Murray, London, 1859)
Jacob, F. Evolution and tinkering. Science 196, 1161–1166 (1977)
Salvini-Plawen, L. V. & Mayr, E. On the evolution of photoreceptors and eyes. Evol. Biol. 10, 207–263 (1977)
Mortlock, R. R. (ed.) Microorganisms as Model Systems for Studying Evolution (Plenum, New York, 1984)
Dawkins, R. The Blind Watchmaker (Norton, New York, 1986)
Goldsmith, T. H. Optimization, constraint, and history in the evolution of eyes. Q. Rev. Biol. 65, 281–322 (1990)
Piatigorsky, J. & Wistow, G. The recruitment of crystallins: new functions precede gene duplication. Science 252, 1078–1079 (1991)
Land, M. F. & Fernald, R. D. The evolution of eyes. Annu. Rev. Neurosci. 15, 1–29 (1992)
Nilsson, D.-E. & Pelger, S. A pessimistic estimate of the time required for an eye to evolve. Proc. R. Soc. Lond. B 256, 53–58 (1994)
Meléndez-Hevia, E., Waddell, T. G. & Cascante, M. The puzzle of the Krebs citric acid cycle: assembling the pieces of chemically feasible reactions, and opportunism in the design of metabolic pathways during evolution. J. Mol. Evol. 43, 293–303 (1996)
Chen, L., DeVries, A. L. & Cheng, C.-H. C. Evolution of antifreeze glycoprotein gene from a trypsinogen gene in Antarctic notothenioid fish. Proc. Natl Acad. Sci. USA 94, 3811–3816 (1997)
Dean, A. M. & Golding, G. B. Protein engineering reveals ancient adaptive replacements in isocitrate dehydrogenase. Proc. Natl Acad. Sci. USA 94, 3104–3109 (1997)
Newcomb, R. D. et al. A single amino acid substitution converts a carboxylesterase to an organophosphorus hydrolase and confers insecticide resistance on a blowfly. Proc. Natl Acad. Sci. USA 94, 7464–7468 (1997)
Miller, K. R. Finding Darwin's God (Cliff Street, New York, 1999)
Yokoyama, S. & Radlwimmer, F. B. The molecular genetics and evolution of red and green colour vision in vertebrates. Genetics 158, 1697–1710 (2001)
Wilkins, A. S. The Evolution of Developmental Pathways (Sinauer, Sunderland, Massachusetts, 2002)
Ray, T. S. in Artificial Life II (eds Langton, C. G., Taylor, C., Farmer, J. D. & Rasmussen, S.) 371–408 (Addison–Wesley, Redwood City, California, 1991)
Bedau, M. A., Snyder, E., Brown, C. T. & Packard, N. H. in Proc. 4th Eur. Conf. Artif. Life (eds Husband, P. & Harvey, I.) 125–134 (MIT Press, Cambridge, Massachusetts, 1997)
Adami, C. Introduction to Artificial Life (Springer, New York, 1998)
Lenski, R. E., Ofria, C., Collier, T. C. & Adami, C. Genome complexity, robustness and genetic interactions in digital organisms. Nature 400, 661–664 (1999)
Wagenaar, D. & Adami, C. in Artificial Life VII (eds Bedau, M. A., McCaskill, J. S., Packard, N. H. & Rasmussen, S.) 216–220 (MIT Press, Cambridge, Massachusetts, 2000)
Wilke, C. O., Wang, J. L., Ofria, C., Lenski, R. E. & Adami, C. Evolution of digital organisms at high mutation rate leads to survival of the flattest. Nature 412, 331–333 (2001)
Yedid, G. & Bell, G. Microevolution in an electronic microcosm. Am. Nat. 157, 465–487 (2001)
Ofria, C., Adami, C. & Collier, T. C. Design of evolvable computer languages. IEEE Trans. Evol. Comput. 6, 420–424 (2002)
Wilke, C. O. & Adami, C. The biology of digital organisms. Trends Ecol. Evol. 17, 528–532 (2002)
Yedid, G. & Bell, G. Macroevolution simulated with autonomously replicating computer programs. Nature 420, 810–812 (2002)
Dennett, D. in Encyclopedia of Evolution (ed. Pagel, M.) E83–E92 (Oxford Univ. Press, New York, 2002)
Holland, J. H. Adaptation in Natural and Artificial Systems (MIT Press, Cambridge, Massachusetts, 1992)
Koza, J. R. Genetic Programming (MIT Press, Cambridge, Massachusetts, 1992)
Lipson, H. & Pollack, J. B. Automatic design and manufacture of robotic life forms. Nature 406, 974–978 (2000)
Foster, J. A. Evolutionary computation. Nature Rev. Genet. 2, 428–436 (2001)
Dennett, D. C. Darwin's Dangerous Idea (Simon & Schuster, New York, 1995)
Drake, J. W., Charlesworth, B., Charlesworth, D. & Crow, J. F. Rates of spontaneous mutation. Genetics 148, 1667–1686 (1998)
Acknowledgements
We thank A. Bennett, J. Bull, J. Coyne, D. Lenski, M. Lenski and E. Zuckerkandl for comments. The authors' work is supported by the US National Science Foundation Biocomplexity Program and by the MSU Foundation. Part of this work was carried out at the Jet Propulsion Laboratory under contract with the US National Aeronautics and Space Administration.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
41586_2003_BFnature01568_MOESM1_ESM.pdf
Supplementary Information: Sections I, II and III provide more background information on Avida and the logic functions that digital organisms perform. Section IV provides additional data on phenotypic and genomic evolution along the line of descent in the case-study population, up to the time of origin of the EQU function. (PDF 42 kb)
Rights and permissions
About this article
Cite this article
Lenski, R., Ofria, C., Pennock, R. et al. The evolutionary origin of complex features. Nature 423, 139–144 (2003). https://doi.org/10.1038/nature01568
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature01568
This article is cited by
-
Ontology for the Avida digital evolution platform
Scientific Data (2023)
-
Ongoing shuffling of protein fragments diversifies core viral functions linked to interactions with bacterial hosts
Nature Communications (2023)
-
Self-replication of a quantum artificial organism driven by single-photon pulses
Scientific Reports (2021)
-
Strong Emergence in Biological Systems: Is It Open to Mathematical Reasoning?
Acta Biotheoretica (2021)
-
Inclusive groups can avoid the tragedy of the commons
Scientific Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.