Abstract
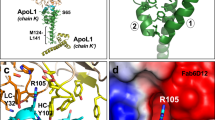
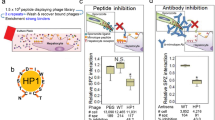
Human sleeping sickness in east Africa is caused by the parasite Trypanosoma brucei rhodesiense. The basis of this pathology is the resistance of these parasites to lysis by normal human serum (NHS)1,2. Resistance to NHS is conferred by a gene that encodes a truncated form of the variant surface glycoprotein termed serum resistance associated protein (SRA)3,4. We show that SRA is a lysosomal protein, and that the amino-terminal α-helix of SRA is responsible for resistance to NHS. This domain interacts strongly with a carboxy-terminal α-helix of the human-specific serum protein apolipoprotein L-I (apoL-I). Depleting NHS of apoL-I, by incubation with SRA or anti-apoL-I, led to the complete loss of trypanolytic activity. Addition of native or recombinant apoL-I either to apoL-I-depleted NHS or to fetal calf serum induced lysis of NHS-sensitive, but not NHS-resistant, trypanosomes. Confocal microscopy demonstrated that apoL-I is taken up through the endocytic pathway into the lysosome. We propose that apoL-I is the trypanosome lytic factor of NHS, and that SRA confers resistance to lysis by interaction with apoL-I in the lysosome.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Hager, K. M. et al. Endocytosis of a cytotoxic human high density lipoprotein results in disruption of acidic intracellular vesicles and subsequent killing of African trypanosomes. J. Cell Biol. 126, 155–167 (1994)
Raper, J., Portela, M. P., Lugli, E., Frevert, U. & Tomlinson, S. Trypanosome lytic factors: novel mediators of human innate immunity. Curr. Opin. Microbiol. 4, 402–408 (2001)
De Greef, C. & Hamers, R. The serum-resistance associated (SRA) gene of Trypanosoma brucei rhodesiense encodes a VSG-like protein. Mol. Biochem. Parasitol. 68, 277–284 (1994)
Xong, H. V. et al. A VSG expression site-associated gene confers resistance to human serum in Trypanosoma rhodesiense. Cell 95, 839–846 (1998)
Nolan, D. P., Geuskens, M. & Pays, E. N-linked glycans containing linear poly-N-acetyllactosamine as sorting signals in endocytosis in Trypanosoma brucei. Curr. Biol. 9, 1169–1172 (1999)
Kelley, R. J., Alexander, D. L., Cowan, C., Balber, A. E. & Bangs, J. D. Molecular cloning of p67, a lysosomal membrane glycoprotein from Trypanosoma brucei. Mol. Biochem. Parasitol. 98, 17–28 (1999)
Blum, M. L. et al. A structural motif in the variant surface glycoproteins of Trypanosoma brucei. Nature 362, 603–609 (1993)
Duchateau, P. N. et al. Apolipoprotein L, a new human high density lipoprotein apolipoprotein expressed by the pancreas. Identification, cloning, characterization, and plasma distribution of apolipoprotein L. J. Biol. Chem. 272, 25576–25582 (1997)
Page, N. M., Butlin, D. J., Lomthaisong, K. & Lowry, P. J. The human apolipoprotein L gene cluster: identification, classification, and sites of distribution. Genomics 74, 71–78 (2001)
Duchateau, P. N., Pullinger, C. R., Cho, M. H., Eng, C. & Kane, J. P. Apolipoprotein L gene family: tissue-specific expression, splicing, promoter regions; discovery of a new gene. J. Lipid Res. 42, 620–630 (2001)
Monajemi, H., Fontijn, R. D., Pannekoek, H. & Horrevoets, A. J. The apolipoprotein L gene cluster has emerged recently in evolution and is expressed in human vascular tissue. Genomics 79, 539–546 (2002)
Lins, L., Charloteaux, B., Thomas, A. & Brasseur, R. Computational study of lipid-destabilizing protein fragments: towards a comprehensive view of tilted peptides. Proteins 44, 435–447 (2001)
Smith, A. B., Esko, J. D. & Hajduk, S. L. Killing of trypanosomes by the human haptoglobin-related protein. Science 268, 284–286 (1995)
Hatada, S. et al. No trypanosome lytic activity in the sera of mice producing human haptoglobin-related protein. Mol. Biochem. Parasitol. 119, 291–294 (2002)
Drain, J., Bishop, J. R. & Hajduk, S. L. Haptoglobin-related protein mediates trypanosome lytic factor binding to trypanosomes. J. Biol. Chem. 276, 30254–30260 (2001)
Shimamura, M., Hager, K. M. & Hajduk, S. L. The lysosomal targeting and intracellular metabolism of trypanosome lytic factor by Trypanosoma brucei brucei. Mol. Biochem. Parasitol. 115, 227–237 (2001)
Do Thi, C. D., Aerts, D., Steinert, M. & Pays, E. High homology between variant surface glycoprotein gene expression sites of Trypanosoma brucei and Trypanosoma gambiense. Mol. Biochem. Parasitol. 48, 199–210 (1991)
Salmon, D. et al. A novel heterodimeric transferrin receptor encoded by a pair of VSG expression site-associated genes in T. brucei. Cell 78, 75–86 (1994)
Guex, N. & Peitsch, M. C. SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modeling. Electrophoresis 18, 14–23 (1997)
Laskowski, R. A., MacArthur, M. W., Moss, D. S. & Thornton, J. M. Procheck: a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993)
Brasseur, R. WinMGM: a fast CPK molecular graphics program for analyzing molecular structure. J. Mol. Graphics 12, 212–218 (1994)
Acknowledgements
We thank M. Chamekh for the pQ30-SRA plasmid, A. Michel and V. Stroobant for peptide synthesis, S. Kler and C. Flore for technical assistance, and D. Perez-Morza and S. Schurmans for help. This work was supported by the Belgian Fund for Scientific Research (FRSM, FWO and Crédit aux Chercheurs) and by the Interuniversity Attraction Poles Programme, Belgian State, Federal Office for Scientific, Technical and Cultural Affairs. R.B., L.V. and L.L. are, respectively, Research Director and Research Associates at the Belgian Fund for Scientific Research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Rights and permissions
About this article
Cite this article
Vanhamme, L., Paturiaux-Hanocq, F., Poelvoorde, P. et al. Apolipoprotein L-I is the trypanosome lytic factor of human serum. Nature 422, 83–87 (2003). https://doi.org/10.1038/nature01461
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature01461
This article is cited by
-
DNAzyme-based faithful probing and pulldown to identify candidate biomarkers of low abundance
Nature Chemistry (2024)
-
Apolipoprotein L1 (APOL1) renal risk variant-mediated podocyte cytotoxicity depends on African haplotype and surface expression
Scientific Reports (2024)
-
A SNARE protective pool antagonizes APOL1 renal toxicity in Drosophila nephrocytes
Cell & Bioscience (2023)
-
Macrophage-mediated trogocytosis contributes to destroying human schistosomes in a non-susceptible rodent host, Microtus fortis
Cell Discovery (2023)
-
Identification and validation of candidate clinical signatures of apolipoprotein L isoforms in hepatocellular carcinoma
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.