Abstract
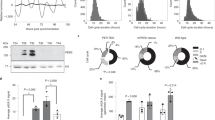
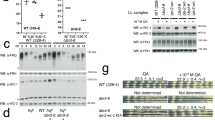
In the mouse circadian clock, a transcriptional feedback loop is at the centre of the clockwork mechanism. Clock and Bmal1 are essential transcription factors that drive the expression of three period genes (Per1–3) and two cryptochrome genes (Cry1 and Cry2)1,2,3,4,5. The Cry proteins feedback to inhibit Clock/Bmal1-mediated transcription by a mechanism that does not alter Clock/Bmal1 binding to DNA6. Here we show that transcriptional regulation of the core clock mechanism in mouse liver is accompanied by rhythms in H3 histone acetylation, and that H3 acetylation is a potential target of the inhibitory action of Cry. The promoter regions of the Per1, Per2 and Cry1 genes exhibit circadian rhythms in H3 acetylation and RNA polymerase II binding that are synchronous with the corresponding steady-state messenger RNA rhythms. The histone acetyltransferase p300 precipitates together with Clock in vivo in a time-dependent manner. Moreover, the Cry proteins inhibit a p300-induced increase in Clock/Bmal1-mediated transcription. The delayed timing of the Cry1 mRNA rhythm, relative to the Per rhythms, is due to the coordinated activities of Rev-Erbα and Clock/Bmal1, and defines a new mechanism for circadian phase control.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
King, D. P. et al. Positional cloning of the mouse circadian Clock gene. Cell 89, 641–653 (1997)
Bunger, M. K. et al. Mop3 is an essential component of the master circadian pacemaker in mammals. Cell 103, 1009–1017 (2000)
Gekakis, N. et al. Role of the CLOCK protein in the mammalian circadian mechanism. Science 280, 1564–1569 (1998)
Jin, X. et al. A molecular mechanism regulating rhythmic output from the suprachiasmatic circadian clock. Cell 96, 57–68 (1999)
Kume, K. et al. mCRY1 and mCRY2 are essential components of the negative limb of the circadian clock feedback loop. Cell 98, 193–205 (1999)
Lee, C., Etchegaray, J. P., Cagampang, F. R., Loudon, A. S. & Reppert, S. M. Posttranslational mechanisms regulate the mammalian circadian clock. Cell 107, 855–867 (2001)
Reppert, S. M. & Weaver, D. R. Coordination of circadian timing in mammals. Nature 418, 935–941 (2002)
Akhtar, R. A. et al. Circadian cycling of the mouse liver transcriptome, as revealed by cDNA microarray, is driven by the suprachiasmatic nucleus. Curr. Biol. 12, 540–550 (2001)
Panda, S. et al. Coordinated transcription of key pathways in the mouse by the circadian clock. Cell 109, 307–320 (2002)
Storch, K. F. et al. Extensive and divergent circadian gene expression in liver and heart. Nature 417, 78–83 (2002)
Yagita, K., Tamanini, F., van Der Horst, G. T. & Okamura, H. Molecular mechanisms of the biological clock in cultured fibroblasts. Science 292, 278–281 (2001)
Orphanides, G. & Reinberg, D. A unified theory of gene expression. Cell 108, 439–451 (2002)
Jenuwein, T. & Allis, C. D. Translating the histone code. Science 293, 1074–1080 (2001)
Fry, C. J. & Peterson, C. L. Chromatin remodeling enzymes: who's on first? Curr. Biol. 11, R185–R197 (2001)
Takahata, S. et al. Transactivation mechanisms of mouse clock transcription factors, mClock and mArnt. Genes Cells 5, 739–747 (2000)
Preitner, N. et al. The orphan nuclear receptor REV-ERBα controls circadian transcription within the positive limb of the mammalian circadian oscillator. Cell 110, 251–260 (2002)
Ueda, H. R. et al. A transcription factor response element for gene expression during circadian night. Nature 418, 534–539 (2002)
Crosio, C., Cermakian, N., Allis, C. D. & Sassone-Corsi, P. Light induced chromatin modification in cells of the mammalian circadian clock. Nature Neurosci. 3, 1241–1247 (2000)
Wade, P. A., Jones, P. L., Vermaak, D. & Wolffe, A. P. Purification of a histone deacetylase complex from Xenopus laevis: preparation of substrates and assay procedures. Methods Enzymol. 304, 715–725 (1999)
Rundlett, S. E. et al. HDA1 and RPD3 are members of distinct yeast histone deacetylase complexes that regulate silencing and transcription. Proc. Natl Acad. Sci. USA 93, 14503–14508 (1996)
Carmen, A. A., Rundlett, S. E. & Grunstein, M. HDA1 and HDA3 are components of a yeast histone deacetylase (HDA) complex. J. Biol. Chem. 271, 15837–15844 (1996)
Guschin, D., Wade, P. A., Kikyo, N. & Wolffe, A. P. ATP-dependent histone octamer mobilization and histone deacetylation mediated by the Mi-2 chromatin remodeling complex. Biochemistry 39, 5238–5245 (2000)
Pfeifer, G. P., Chen, H. H., Komura, J. & Riggs, A. D. Chromatin structure analysis by ligation-mediated and terminal transferase-mediated polymerase chain reaction. Methods Enzymol. 304, 548 (1999)
Acknowledgements
We thank D. Reinberg for the anti-RPB1 antibody; V. Sartorelli for the p300 expression construct; U. Schibler for Rev-Erbα reagents; H. R. Ueda for PATSER sequence analysis; and D. R. Weaver and C. L. Peterson for suggestions. This work was supported by grants from the NIH and the Defense Advanced Research Projects Agency (DARPA).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Etchegaray, JP., Lee, C., Wade, P. et al. Rhythmic histone acetylation underlies transcription in the mammalian circadian clock. Nature 421, 177–182 (2003). https://doi.org/10.1038/nature01314
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature01314
This article is cited by
-
LKRSDH-dependent histone modifications of insulin-like peptide sites contribute to age-related circadian rhythm changes
Nature Communications (2024)
-
Epigenetics and seasonal timing in animals: a concise review
Journal of Comparative Physiology A (2023)
-
Modulation of cellular processes by histone and non-histone protein acetylation
Nature Reviews Molecular Cell Biology (2022)
-
Origins of human disease: the chrono-epigenetic perspective
Nature Reviews Genetics (2021)
-
MRG15 orchestrates rhythmic epigenomic remodelling and controls hepatic lipid metabolism
Nature Metabolism (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.