Abstract
Only a small proportion of the mouse genome is transcribed into mature messenger RNA transcripts. There is an international collaborative effort to identify all full-length mRNA transcripts from the mouse, and to ensure that each is represented in a physical collection of clones. Here we report the manual annotation of 60,770 full-length mouse complementary DNA sequences. These are clustered into 33,409 ‘transcriptional units’, contributing 90.1% of a newly established mouse transcriptome database. Of these transcriptional units, 4,258 are new protein-coding and 11,665 are new non-coding messages, indicating that non-coding RNA is a major component of the transcriptome. 41% of all transcriptional units showed evidence of alternative splicing. In protein-coding transcripts, 79% of splice variations altered the protein product. Whole-transcriptome analyses resulted in the identification of 2,431 sense–antisense pairs. The present work, completely supported by physical clones, provides the most comprehensive survey of a mammalian transcriptome so far, and is a valuable resource for functional genomics.
Similar content being viewed by others
Main
With the availability of draft sequences of the human genome1,2, increasing attention has focused on identifying the complete set of mammalian genes, both protein-coding and non-protein-coding. As various analyses3,4 have noted, different annotation criteria lead to different sets of predicted genes, and even the true number of protein-coding genes remains uncertain. Biases in the training sets used for optimizing gene-finding algorithms lead to systematic biases in the types of genes that are predicted computationally—and so important genes and gene classes can be missed. A systematic analysis of transcripts from human chromosome 21 using dense oligonucleotide arrays revealed that the apparent transcriptional output of processed cytoplasmic messenger RNAs exceeds the predicted exons by an order of magnitude5.
One significant class of ‘genes’ missing from the existing genome annotation are those that give rise to non-protein-coding RNAs. Non-coding RNAs, although not highly transcribed6, constitute a major functional output of the genome. In addition to their role in protein synthesis (ribosomal and transfer RNAs), non-coding RNAs have been implicated in control processes such as genomic imprinting7 and perhaps more globally in control of genetic networks8.
Because of these and other limitations, it is clear that the utility of mammalian genome sequences will be more fully realized when there is a complete description of a mammalian transcriptome. The transcriptome includes all RNAs synthesized in an organism, including protein-coding, non-protein-coding, alternatively spliced, alternatively polyadenylated, alternatively initiated, sense, antisense, and RNA-edited transcripts. Attempts to catalogue the mammalian transcriptome have been based upon assembly of sequences from large-scale expressed-sequence-tag (EST) sequencing projects9. More recently, serial analysis of gene expression10 and large-scale sequencing of open reading frame (ORF) sequence tags11 have contributed to the definition of the transcriptome and have improved genome annotation. Each has the limitation that no physical cDNA clones are fully sequenced. The RIKEN Mouse Gene Encyclopedia Project is a large-scale effort to isolate and sequence novel full-length mouse cDNAs. The initial results and validation of our approach have been reported previously, and resulted in the functional annotation of 21,076 full-length cDNAs (FANTOM1)12. Here we describe the characterization and annotation of the FANTOM2 clone set (60,770), consisting of 39,694 new cDNA clones (FANTOM2 new set) and the 21,076 FANTOM1 clone set. We also present a global analysis of the mouse transcriptome based upon an integration of the FANTOM2 sequences with all available mouse mRNA data from the public sequence databases13. This data set provides the most comprehensive view so far of the transcriptional potential and diversity within the mouse genome, and serves as an important model for analysing the transcriptomes of other higher eukaryotes.
To enable a global description of the transcriptome, we use the term transcriptional unit (TU) to describe a segment of the genome from which transcripts are generated. A TU is defined by the identification of a cluster of transcripts that contains a common core of genetic information (in some cases, a protein-coding region). The main advantage of this approach is that the definition is purely computational and unequivocal. The existence of a TU is inferred from the identification of mRNAs via full-length cDNA isolation and sequencing. A single cDNA sequence may define a TU. When multiple cDNA transcripts that share DNA sequence are identified, we define these sequence-based clusters as a single TU. For each TU, the 5′ boundary is defined at the most distal transcription start site, and the 3′ boundary at the most extreme poly(A) sequence. TUs are DNA strand-specific, and are typically bounded by promoters at one end and termination sequences at the other. With this definition, alternative spliced transcripts, alternative 5′ start and 3′ ends, and variants arising from recombination (as in the immunoglobulin and T-cell receptor loci) are subsumed within a single TU, even though they may generate protein or RNA products with very different functions. TUs on opposite strands are counted separately, even if they overlap spatially in the genome. Thus, antisense transcripts are considered to be made from separate TUs.
The FANTOM2 clone set
The cDNA clones used in the project were selected from 246 full-length enriched cDNA libraries, most of which were also normalized and subtracted, with most from C57BL/6J mice, as described elsewhere12,14,15. Information about tissue source and other information regarding these libraries is available in Supplementary Information section 1.
1,442,236 sequences were grouped into 171,144 3′-end clusters based on sequence similarity (Supplementary Information section 2)14,15,16. Of these, 159,789 had no significant BLAST hit to known mouse genes, and were considered potentially novel. However, from the annotation of the FANTOM1 clone set12 it became clear that alternative polyadenylation is common in the mouse transcriptome, and that the set of 3′-end clusters is significantly redundant. To address this problem, 547,149 of these clones were sequenced from their 5′ ends to provide additional discrimination.
Representative cDNAs were selected to represent target clusters and fully sequenced as described in Supplementary Information section 2. In total, we generated 60,770 high-quality, full-insert cDNA sequences with an average length of 1.97 kilobases (kb) (Supplementary Information section 3). If the smaller FANTOM1 clones are removed, the average insert size is 2.35 kb. The sequences of all of the cDNAs are available from DDBJ/EMBL/GenBank. A summary of all the sequence quality information, including the distribution of Phred/Phrap scores, is provided in Supplementary Information section 4. This table also demonstrates the excellent agreement between the cDNA sequences and the completed mouse genome sequence. Overall, there was 99.8% identity of aligned bases, and more than half the differences were mismatches rather than insertions or deletions. Among the 14,276 clones having >98% similarity to SWISS-PROT + TrEMBL cDNAs, automatic non-annotated alignment suggested that at least 67% of the cDNA clones were completely full length (Supplementary Information section 5) including alternative amino or carboxy terminals, while about 4% showed complete alignment but for unknown reasons lacked start or stop codons.
Annotating and mapping the RIKEN mouse cDNAs
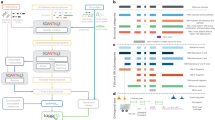
To expedite manual annotation, the 60,770 cDNAs were first subjected to automated annotation. The decision tree used for automated annotation is presented in Fig. 1 and Supplementary Information section 6. Briefly, clone sequences that had a high degree of similarity to known genes in the Mouse Genome Informatics (MGI, http://www.informatics.jax.org/mgihome/) and LocusLink/RefSeq (http://www.ncbi.nlm.nih.gov/LocusLink/)17 databases were assigned the official gene name and available Gene Ontology18 (GO) terms. Sequences that were considered possible paralogues to known mouse genes or possible orthologues of genes from another mammalian species based upon an imperfect match to previously characterized genes were annotated using an analogous process—except that the name assigned to the clone sequence was qualified with the suffix ‘homologue’, or the prefix ‘similar to’ or ‘weakly similar to’, depending on the degree of similarity of the protein match. Unassigned sequences with matches to mouse mRNA/EST clusters (UniGene or TIGR's tentative consensus sequences) were assigned the annotation of the EST cluster. Clones with good coding sequence predictions in which the only available information was a match to a known protein domain or predicted motifs were assigned a name based on the domain match. Clone sequences with no nucleotide or protein match but with a valid coding sequence prediction of at least 100 amino acids in length were called ‘hypothetical proteins’. Clones for which there was an independent match to at least one other mouse EST from a distinct library were classified as ‘unknown EST’. All other clones were labelled ‘unclassifiable’. Throughout the process, an effort was made to use only informative names.
Nomenclature and criteria used in the gene name are described on the right-hand side. Gene names were annotated (right-hand side) from the gene function descriptor of the reference (left-hand side) according to the nomenclature. Priority was given to reference descriptors from which functional information could be inferred, even if references with less informative descriptors were more similar to the clones. CDS, coding sequence; ID, identity; aa, amino acid; bp, base pair; EST, expressed sequence tag.
The results of the automated computational analyses were reviewed by an international consortium of sequence annotators and gene-family experts over 2 months during the Mouse Annotation Teleconference for RIKEN cDNA sequences (MATRICS). These annotators altered the computational annotation in 24.8% of cases. The best coding sequence (CDS) was chosen by MATRICS annotators using a standardized set of criteria. If no good CDS existed, annotators classified sequences as unknown, or alternatively as 5′ untranslated region (UTR), or 3′ UTR if there was evidence of alignment with a longer transcript from a known gene. Clones with CDS of less than 100 amino acids were annotated only if supported by known proteins, motifs, predicted signal peptide, transmembrane regions or splicing evidence.
The FANTOM2 cDNA annotation system used in MATRICS also provides a tool for continued annotation and a high-quality viewer for each transcript (Supplementary Information section 7). The FANTOM2 annotation viewer is accessible at http://fantom2.gsc.riken.go.jp/. The results of the annotation for the representatives of these clones are described below.
The cDNA sequences were provided to the Mouse Genome Sequencing Consortium, and have been used in their genome sequence annotation in the accompanying report19. We independently mapped the cDNA sequences to the mouse genome sequence (MGSCv3) assembly (ftp://wolfram.wi.mit.edu/pub/mouse_contigs/MGSC_V3) as described in Methods. This provided a basis for determining orthology between mouse and human genes in the annotation process, based on knowledge of conserved linkage between these two species20. Genome analysis also provided insight into the intron–exon structure of mouse genes. The unambiguously assigned chromosomal locations are summarized, and distribution of the clones along the mouse chromosomes is presented, in Supplementary Information section 8.
Analysis of the transcriptome
Creating a comprehensive transcript data set for the mouse
To enable an overview of the transcriptome, a single transcript sequence from each TU was chosen as a representative. Because the focus of the RIKEN Mouse Gene Encyclopedia Project has been to identify and sequence novel transcripts, the 60,770 FANTOM2 cDNA set does not contain a representative of all known mouse TUs. We therefore derived a comprehensive representative transcript and protein set (RTPS), using a layered strategy of redundancy detection to identify transcripts that correspond to the same TUs, first in the FANTOM2 collection and then among representative sequences of the FANTOM2 TUs and known mouse cDNAs (Supplementary Information section 9). The FANTOM2 cDNAs were first clustered by sequence similarity using ClusTrans (see Methods). Manual annotation corrected the number of clusters by ClusTrans from 35,957 to 35,637 and in some cases changed the representative clones. Of the 35,637 representative sequences, 11,108 were derived from multiple sequence clusters, while 24,529 were unclustered singletons.
The set of 35,637 cluster-resolved FANTOM2 representatives were then combined with a redundant set of 44,106 transcripts for mouse mRNA sequences compiled from LocusLink, MGI and GenBank (non-EST) databases (mouse public data set) (Supplementary Information section 9). The method for redundancy reduction and clustering criteria are outlined in Supplementary Information section 21. This step reduced the number of unique representative FANTOM2 sequences to 33,409 and the total number of representative transcripts in the RTPS to 37,086 (Fig. 2). 10,009 of the 33,409 TUs within the 60,770 FANTOM2 cDNA clone set correspond to known mouse genes of non-FANTOM2 sequences characterized in public databases (MGI, NCBI LocusLink, GenBank). The FANTOM2 data set represents an additional 15,923 novel TUs, a significant addition to the elucidation of the mouse transcriptome. Among these, 4,258 TUs contain protein-coding transcripts, and the remaining 11,665 TUs lack protein-coding potential. The proportion of novel non-protein-coding transcripts increased in the FANTOM2 new set compared to the FANTOM1 set. This could be a consequence of the intensive library normalization and subtraction strategy designed to collect less-abundant transcripts. The implication is that non-coding RNAs are relatively rare. The features of non-coding RNAs compared to the protein-coding RNAs are described later, and are shown in Supplementary Information section 14.
[Public] consists of 44,106 sequences derived from the mouse public data set (transcripts for mouse mRNA sequences compiled from LocusLink, Mouse Genome Informatics and GenBank (non-EST) databases). [FANTOM1] contains 21,076 sequences that have been reported previously12. [FANTOM2 New] contains 39,694 new sequences reported in the present work. In each box, the top row shows the count of TUs in each category, and the breakdown (protein-coding and non-protein-coding) is shown in the bottom row.
Functional annotation and CDS analysis by manual annotation
Table 1 represents the results of gene annotation for the FANTOM2 clone set. Of the 33,409 representatives, 13,736 (41.1%) were assigned some functional information. 6,929 (20.7%) TUs were assigned gene names from the categories of ‘known mouse DNA’, ‘known mouse protein’ or ‘inferred from EST/mRNA cluster’, which are the exact matches to a mouse gene with a known function. The categories ‘homologue’, ‘similar to’, ‘weakly similar to’ and ‘domain/motif’ represent functional information for 4,623 TUs (13.8%), inferred from sequence similarity. Others were assigned as ‘hypothetical protein’, ‘unknown EST’ or ‘unclassifiable’, hence no functional information was available. These TUs are potential candidates for novel genes.
Manually annotated CDS status was also evaluated for the representative clones of the FANTOM2 set (Supplementary Information section 10). Of the 33,409 FANTOM2 representative TUs, 17,594 were considered to have coding potential. Of these, 12,267 (69.7%) and 2,078 (11.8%) were translated into complete and partial protein sequences, respectively. Combining the protein sequences of FANTOM2 with those of public databases, the estimate of the protein coding TUs in the RTPS was 20,487.
Additional TU predicted by 5′ and 3′ EST mapping
In addition to mapping the cDNA sequences to the MGSCv3 genome to assist annotation, we also mapped 547,149 5′ ESTs and 1,442,236 3′ ESTs from the phase 1 sequencing and non-RIKEN contributions to a public EST database (NCBI dbEST) (Supplementary Information section 20). EST clusters were mapped to the mouse genome assembly based upon a minimal match of 95% over 100 base pairs (bp). The number of candidate TUs that were aligned outside the regions encompassed by the boundaries of TU within the RTPS is shown in Table 2. 5′ end, 3′ end and 5′ + 3′ pairs were counted separately. Of these, 3,425 candidate TUs that have both 5′ + 3′ pairs with additional EST support provide a minimal additional count to the 37,086 TUs in the RTPS. 3,422 5′ ESTs and 6,703 3′ ESTs are also supported by other ESTs. If we include singletons, the number of candidate TUs is estimated at 70,000.
Assignment of the transcription start sites of TUs
The correlation of expression measurements with identified promoter regions permits the association of specific cis-acting elements with particular patterns of expression21. In the mouse, this will only be possible when there is a complete enumeration of the transcription start sites for all TUs, so that the flanking sequences can be identified and analysed to identify promoters. To assess the reliability of 5′ end assignment, we performed an alignment of FANTOM2 and RIKEN 5′ ESTs to known genes. 131,335 sequences were aligned with previously known complete cDNAs. Of these, 116,660 5′ ESTs or complete cDNAs could be aligned on the 5′ UTR of the corresponding gene. 57,636 (4,431 genes) extended at least 10 nucleotides further than the recorded 5′ ends, and probably represent a more accurate assignment of the transcription start point; 29,359 (3,068 genes) were within 10 bp of the 5′ end of the mRNA and confirmed the starting point; and 29,665 (3,021 genes) were at least 10 nucleotides shorter than the published starting point although they still carry the first ATG. Some truncated transcripts will represent alternative promoter use, or the known diversity of start sites in TATA-less promoters22.
Alternative splicing in the mouse transcriptome
The evolution of higher eukaryotes is associated with an increase in the number of introns and the incidence of alternative exon splicing. Previous EST-based estimates of alternative splice variants1,23 suggested that at least 59% of human and 33% of mouse genes are alternatively spliced. To assess the incidence of alternative splicing in mouse, we aligned 60,770 RIKEN clone sequences and the 44,106 mouse mRNA transcripts from the LocusLink, MGI (non-EST) and GenBank (non-EST) databases to the MGSCv3 mouse genome assembly using BLAST followed by SIM4 (ref. 24). 77,640 sequences out of 104,876 sequences were mapped to the mouse genome and were grouped into 33,042 clusters (Table 3). The removal of unspliced and single-exon derived transcripts, which are also irrelevant for splice analysis, reduced the number of clusters to 18,970. Of these, 7,293 clusters were singletons and excluded from further analysis. 4,750 (41%) clusters out of the remaining 11,677 clusters showed evidence of cryptic exons (exons included in only a subset of transcripts from a cluster) or exon length variation (Table 3). Modrek et al.25 obtained similar numbers (42%) for human splice forms. The details of the status of each of the exon types from the 4,750 clusters are shown in Supplementary Information section 11. All variant and non-variant cluster transcripts can be explored at http://genomes.rockefeller.edu/MouSDB/.
Patterns in untranslated regions of mRNAs
We compared the entire clone set to the database of regulatory elements specific to mRNA UTRs (http://bighost.area.ba.cnr.it/BIG/UTRHome/) using PatSearch26. The incidence of UTR-specific functional motifs (Supplementary Information section 12) provided additional functional annotation. The relevant annotation for each pattern was added only to those clones where the nature of the predicted protein and the specific location of the pattern allowed a very conservative assessment of its biological activity. This allowed confirmation of gene annotation in instances where the CDS was only partially sequenced. Additionally for clones annotated for the SECIS element (selenocysteine insertion sequence), it was possible to correctly define the CDS as it is known that in this case an in-frame UGA stop codon codes for selenocysteine.
Repeats
Insertion events in CDS of transcripts may have direct impact on the protein coding function. The FANTOM2 clone set contains 105,861 repeats, including 14,929 low-complexity regions (Supplementary Information section 13A). 48.5% (51,372) of repeats were found in 64.8% (16,473) of FANTOM2 RTPS representatives covering 16.5% of the TU lengths (Supplementary Information section 13B). Among cDNAs with an annotated CDS, 14.2% (1,232) carry one or more repeats that overlap with the CDS. The majority contain repeats of the SINE (short interspersed nucleotide element), simple repeats, LINE (long interspersed nucleotide element) and LTR (long terminal repeat) classes. Most simple repeats did not cause frameshifts or premature stop codons, whereas the dispersed elements commonly changed the CDS. For example, the insertion of a SINE B2 element into the CDS of a transcript A630030N07 (AK041699) similar to the intron-less ubiquitin-conjugating enzyme Mhr6bn will disrupt translation or produce a non-functional protein. The inserted B2 may affect the transcription of the neighbouring genes by acting as Pol II promoter27. Sequencing of genomes and cDNA collections from strains other than C57BL/6J will distinguish inserted elements that are a source of functional polymorphism from those that represent species-specific mutagenic events.
Non-protein-coding RNAs
The largest class of new transcripts in the FANTOM2 set lack an apparent protein-coding region; 15,815 TUs in the FANTOM2 set may represent functional non-coding RNAs. Supplementary Information section 14 compares sequence information for protein-coding and non-coding transcripts in the FANTOM2 set. The sequence quality does not distinguish the two classes, and does not explain the failure to annotate a CDS. Annotators had access to sequence quality information and also to CDS predictions that take account of possible frame shifts. The clearest difference is that 71% of the non-coding transcripts are unspliced/single exon compared to genomic sequence, whereas 18% of protein-coding transcripts are unspliced/single exon. The finding of 4,148 non-coding RNAs showing splice evidence is also a very interesting new feature. The frequency of polyadenylation signals does not differ significantly between the two classes of transcripts, suggesting that most non-coding transcripts are indeed the product of RNA Pol II-mediated transcription, processing and nuclear export.
To identify the strongest candidates as functional and novel non-coding RNAs, we further eliminated from the 15,815 potential non-coding RNAs any with similarity to known protein sequences identified using BLASTX (E-value <10-5). We mapped the resulting 12,382 sequences to the MGSCv3, and selected clones that aligned at greater than 90% identity over more than 90% of their length. To further filter the data, we used exon predictions from GENSCAN analysis of the genomic sequence. Because GENSCAN may miss non-coding regions and UTRs of protein-coding transcripts, we considered sequences mapping on the same strand and within 10 kb of any predicted exon as protein-coding candidates and eliminated these from further consideration. The remaining 4,280 transcripts were considered the strongest non-coding RNA candidates.
Functional non-coding RNAs are likely to be distinguished by their reproducible expression and some measure of sequence conservation to mouse, human, rat EST or human genome. We assembled a line of evidence to support non-coding RNA annotation, including the number of mouse, rat and human EST hits, homology with human genome sequence and CpG islands ((C + G)-rich region) in genomic regions 5′ upstream of each candidate. The nucleotide sequences of 1,194 (27.9%) transcripts out of 4,280 matched previously sequenced mouse ESTs (Table 4). Of these 4,280 representative transcripts from the RTPS, 252 transcripts matched rat ESTs (5.8%), and 111 transcripts matched human ESTs. 454 (10.6%) sequences mapped to both the mouse and human genome sequences, and we were able to identify CpG islands upstream of the transcription start site for 919 (21.5%) transcripts. Additionally, 1,150 (26.9%) of the functional non-coding RNA candidates were spliced (which is not significantly different from the overall frequency in the class) and 323 (7.5%) were also identified as candidate antisense transcripts.
Even applying the most conservative criteria, this suggests that several hundred mouse transcripts are conserved, regulated, non-coding mRNAs. Although the functional role played by these transcripts remains to be studied in detail, the significant fraction of the transcriptional potential devoted to non-protein-coding genes suggests that they indeed have important biological and regulatory roles.
Antisense RNA
There is increasing recognition that the production of RNA transcripts from both strands in a genomic locus can produce coordinate regulation. For example, Air, the antisense of Igf2r, contributes to the paternal imprinting of three genes (Igf2r, Scl22a2 and Scl22a3)28. We focused on antisense transcripts using a slightly different strategy from that used to identify non-coding RNA. First, we aligned the sequences in the FANTOM2 set and public mRNA sequences with the mouse genome MGSCv3, and searched for pairs of sequences that mapped to the same genomic region in opposite orientations. We identified 2,431 pairs of sense–antisense transcripts overlapping in the exons of the sense gene by at least 20 bases. Of these, 1,573 pairs were not previously identified, and 316 of sense–antisense pairs contained the strongest non-coding RNA candidates described above.
Analysis of the predicted proteome
Our analysis of the proteome is based primarily on the previously described representative protein set (RPS), which contained protein sequences for each nucleotide where prediction was possible. We also created a variant-based proteome set (VPS) combining RPS and complete protein sequences representing splice variants not included in RPS. The VPS includes variant forms of known genes identified by sequencing of the FANTOM2 clones. The table of protein families for RPS and VPS is presented in Supplementary Information section 15. This table also includes the number of known members of a given family, including previously known human orthologues and those newly discovered. This analysis provides some indication of the extent to which the RPS represents the proteome. The overall appearance of the predicted mouse proteome is similar to that of the human (http://www.ensembl.org) deduced by genomic sequencing. Although there are fewer sequences in the current human set, the proportion covered by known domain families is the same, and the most abundant families have approximately the same number of members in both proteomes.
Protein domain/motif analysis and functional assignment
The constructed mouse proteome sequence set was analysed using a number of protein domain and motif databases including InterPro29, Superfamily30 and MDS31. An InterPro analysis gives a general functional overview of the sequence set and a functional overview of the transcriptome. InterPro domains were discovered in about 70% of the RPS, similar to other eukaryotic proteomes (http://www.ebi.ac.uk/proteome). Further analysis using the protein sets for human and mouse from the International Protein Index (IPI; Table 5) with the RPS suggests that the general domain composition is very similar for all three sets. The domain distribution in RPS and VPS is also very similar for top InterPro entries. The total number of protein sequences in the FANTOM2 set covered by all domain databases (InterPro, Superfamily) is 17,236 (92% of the RPS). As expected, the most abundant domains are the zinc fingers, which are often duplicated in the same sequence; protein kinases are also highly abundant.
Superfamily domain analysis
Superfamily30 analysis (http://supfam.org/SUPERFAMILY/cgi-bin/gen_list.cgi?genome=mr;password=rumplestiltskin) is used to detect and classify evolutionarily related groups of domains for which there is a known structural representative. An accurate superfamily definition is obtained by detailed manual analysis of structural, sequence and functional evidence for a common evolutionary ancestor32. The ancestral domain from each superfamily represents a genetic building block. These building blocks have been duplicated, recombined and mutated to create the proteins that are currently observed in the genome. A small number of domains have been duplicated a very large number of times, but most have been duplicated only a very few times. 98% of the domains identified by this analysis have been produced by duplication from 715 ancestral domains. The domain architecture for each sequence indicates the recombination of ancestral domains that has taken place during evolution.
Using strict criteria, a number of novel structural domain combinations were found in the FANTOM2 set. We identified 120 unknown, structural, pair-wise combinations of known structural domains; of these pairs, 30 have not been found previously in any sequenced proteome. As well as representing unique recombination events in evolution, these domain pairs provide targets for structural genomics projects that are assured to be novel. These newly discovered domain combinations can be found at http://supfam.org/FANTOM2/domcombs.html.
New domains
The availability of a very large set of new protein-coding transcripts, combined with existing information, permits computational detection of new motifs that are present in multiple independent gene products31. A number of new protein motifs, summarized in the MDS database (http://motif.ics.es.osaka-u.ac.jp/FANTOM2/), were predicted from the FANTOM2 sequences. One example is a structural GTPase submotif (MDS00154), specific for the immune associated nucleotide family (IAN), which also contain the canonical D-X-X-G pattern of the Walker B motif33. The IAN specificity of the MDS00154 motif suggests that it modulates GTPase activity and specificity for signalling in the T-cell differentiation and selection process. For example, mouse Ian1 was reported to be upregulated during the positive and β selection of thymocytes expressing TCR-β and TCR-α/β chains, a process that is essential to the development of a peripheral immune response34.
Membrane and secreted proteins
Membrane and secreted proteins are of particular interest because of their central role in intercellular communication and their accessibility as drug targets. We analysed the RTPS, using two independent methods to predict endoplasmic reticulum signal peptides35 and to predict transmembrane helices, TMHMM 2.0 (ref. 36) and SVMtm (http://genet.imb.uq.edu.au/predictors/). First, we removed from the RTPS 1,559 readily identifiable partial ORFs that did not contain an initial methionine. As the two transmembrane helix prediction methods are subject to the misprediction of signal peptides as transmembrane domains, we developed a filter for predicted N-terminal transmembrane segments. If the predicted transmembrane's starting point was within the first 15 residues of the ORF and a signal peptide was predicted, then the region was regarded as a signal peptide rather than a transmembrane domain.
This analysis identified six classes of proteins on the basis of their membrane organization: (A) non-secretory proteins, (B) soluble/secreted proteins, (C) type I membrane proteins, (D) type II membrane proteins, (E) multi-span membrane proteins, and (F) unclassified proteins (Table 6). For each class, we adopted a stringent consensus method to restrict false-positive predictions, keeping only those motifs that were predicted using multiple methods. On the basis of this consensus prediction protocol, 80.1% (15,174) of the protein ORFs within the RTPS fall into classes A–E. Within the RTPS CDSs, 10.9% (1,877) were annotated as putative secreted proteins. Included among this class are the majority of soluble proteins involved in intercellular communication and the maintenance of the extracellular matrix, and 521 novel putative secreted proteins originating from the FANTOM2 project.
Small proteins
Functional proteins less than 100 amino acids in length were annotated only if they showed significant homology to known proteins from other species or members of gene families. On the basis of these stringent criteria, only 376 proteins of less than 100 amino acids were annotated. 4,558 contained predicted CDS regions encoding proteins between 50 to 99 amino acids in length, and these were reanalysed to identify high-quality, putative short CDSs. First, clones were eliminated if the CDS was not initiated within the first 500 nucleotides, on the assumption that few genuine 5′ UTRs exceed this length. For each of the remaining 3,159 transcripts, we used tBLASTn to search the translated Ensembl human gene predictions. Among the 1,823 transcripts that had homology to a single entry in the human genome annotation, we searched for the potential splicing and identified 557 spliced transcripts. These are regarded as the most likely short protein candidates, although the possibility that some short CDSs are encoded by single exon TUs cannot be discounted. This analysis suggests that short proteins might increase the predicted proteome by as much as 10%, but each candidate will require further individual annotation and validation.
Functional analysis of predicted proteins using Gene Ontology
To summarize the functional capacity of the transcriptome, we used the structured vocabularies of the Gene Ontology (GO) project37 (http://www.geneontology.org/), as described in Methods. Of the 18,768 representative proteins, we were able to assign molecular function GO terms to 11,125 TUs, biological process GO terms to 10,443 TUs and cellular component GO terms to 10,488 TUs, representing 59.3%, 55.6% and 55.9%, respectively. GO annotations for individual genes are displayed in the FANTOM web interface, along with codes denoting the method used in the assignment (http://fantom2.gsc.riken.go.jp/). Supplementary Information section 16 summarizes the results of our analysis.
We also compared the results of our analysis with a similar binning of the Ensembl annotation of the human genome (Supplementary Information section 16). The distributions are very similar, re-emphasizing the importance of the mouse as a model system for human biology. Notable differences are seen in the areas where annotators focused on genes with special biological properties. For example, membrane proteins and extracellular proteins show significant increases in the mouse genome compared to the human genome: 2,892 versus 940 genes, and 4,907 versus 4,068 genes, respectively. Another area where the number of mouse genes was significantly greater than human genes, 3,152 versus 2,310, was in the molecular function ‘transporter’ and the biological process transport. The richness of the mouse annotations from this analysis provides an important foundation for the functional annotation and investigation of human orthologues.
Human disease genes
The catalogue of the mouse transcriptome can also be used to identify candidate genes associated with human diseases and mouse phenotypic variants. We obtained a list of 1,712 human loci associated with diseases by searching LocusLink (http://www.ncbi.nlm.nih.gov/LocusLink). Of these, 1,022 had associated sequence information, while 690 did not. We searched the 1,022 disease-associated protein sequences against the FANTOM2 data set using tBLASTn with a minimum E-value of less than 10-50 as the criterion for significance. Of 921 human diseases that had previously been identified with a homologous mouse gene, 740 (80%) were found in the FANTOM2 data set. For the 101 human disease genes without previously identified mouse homologues, 67 (66%) were mapped to the FANTOM2 data set (http://fantom2.gsc.riken.go.jp/supplement/disease_genes/).
Other gene discoveries
The FANTOM2 cDNA collection also revealed a substantial number of new protein-coding transcripts for which the likely function can be inferred from domain assignments or sequence homology (Supplementary Information section 17). For many of these proteins, we can also ascertain tissue-specific expression patterns from both the library of origin and the RIKEN Expression Array Database (http://read.gsc.riken.go.jp). Furthermore, using GO assignments and sequence similarity, we can assign these to gene families representing the full spectrum of functions of an archetypal mammalian cell. A number of highlights are described below.
• Cell movement. Miki et al.38 have reported the identification of kinesin superfamily proteins (KIFs) in the mouse and human genome. KIFs are motor proteins contributing to intracellular trafficking by transporting vesicles, protein complexes and chromosomes39. In FANTOM2, we found 33 KIFs equivalent to 73.3% of the 45 known loci. Of the 33 KIFs, 17 were full length. Previously, 2 alternative splice variants had been reported. In FANTOM2, we found 4 additional splice variants. Clones of 25 loci were derived from neural tissue or mixtures of neural and other tissue, consistent with previous reports.
• Protein metabolism and turnover. Ubiquitination and targeted protein degradation within the proteosome complex controls many processes in eukaryotic cells, including meiosis, cellular proliferation and development40. Unlike cell death and the cell cycle, which have been studied extensively, many ubiquitination regulatory proteins remain to be discovered in mammals. Our analysis identified a number of new gene products participating in this process, including 4 E1 ubiquitin activating enzymes, 13 E2 ubiquitin conjugating enzymes, 98 E3 ubiquitin ligases and 6 de-ubiquitinating enzymes. The extent of these gene families suggests that the targeting of cellular proteins for degradation is, indeed, a very highly regulated process.
• G-protein-coupled receptors (GPCRs). These comprise the largest family of receptor proteins in mammals, and may be promising drug targets. At present, there are approximately 600 intact GPCR genes that have been identified within the human genome, excluding nearly 350 odorant receptor genes1,2. 410 clones in the FANTOM2 collection encoded candidate GPCRs; clustering to the TUs reduced this number to 213, although the high degree of relatedness and frequency of alternative splicing among some GPCR families make such clustering challenging. Out of these, 165 genes were contained within the 308 GPCRs previously annotated in the MGI database. The remaining 48 were unique to the mouse, and 14 of these had no clear mammalian orthologue. Among these, we identified two new members C630030A14 (AK083234) and 5330439C02 (AK030625) of G-protein-coupled receptor family C, which contains metabotropic glutamate receptors, γ-aminobutyric acid type B receptor, and Ca2+-sensing receptors. These are highly homologous to previously identified GPCRs in the Caenorhabditis elegans and Drosophila genomes.
• The ‘metabolome’. The InterPro and GO functional assignments allowed us to assign corresponding Enzyme Commission (EC) numbers41 to many of the predicted proteins with inferred metabolic enzymatic activities. In total, 726 distinct EC numbers were assigned to 3,583 TUs in the RTPS (approximately 10% of the total), of which nearly 90% (3,182) contained FANTOM cDNA clones. In comparison, the human enzyme class in the KEGG metabolic pathway database42 contains 720 unique EC number assignments.
The RTPS contains representatives of the mouse orthologues of most genes involved in known metabolic pathways. For example, the entirety of the tricarboxylic acid (TCA) cycle (Fig. 3) appears in the RTPS. All the enzymes in the TCA cycle were covered by FANTOM2 clones.
• The impact of alternative splicing on the proteome. As alternative splicing can change protein function, we evaluated its effect on putative translations of 4,750 variant clusters consisting of 22,150 sequences (Table 3), which showed evidence of cryptic splicing or exon length variation. Of the 4,750 variant clusters, 4,263 clusters were potentially protein coding. The analyses of all the exons derived from these 4,750 variant clusters are shown in Supplementary Information section 11. 8,500 exons (2,530 constitutive exons with lengths variation + 3,338 cryptic internal exons + 2,582 cryptic terminal exons) exhibited length variation. Of these 8,500 variant exons, 6,247 (73.5%) are within the CDS and will affect the CDS of 3,378 (79.2%) variant clusters out of 4,263 potential protein-coding clusters. The high number of splice forms translated into potential variant proteins is in line with a previous estimate that 74% of splice variation in human transcripts alters the coding sequence25.
Alternative splicing, especially exon-skipping, can radically change the function of the protein product, and many examples can be seen in our data set. For example, the tyrosine phosphatase receptor type R (D130067J21 (AK051719)) of variant cluster scl1364 (1500019M22 (AK028054), D130067J21 (AK051719) and RefSeq NM_011217) lacks a transmembrane domain owing to alternative splicing. However, the sequence retained a cleavable signal peptide, implying that it could be secreted from cells. The variant also lacks the tyrosine phosphatase domain, and may therefore act as a dominant-negative form of the full-length receptor. Other splice variants among putative signalling proteins cause loss of protein interaction domains. Examples include PDZ (Rap guanine nucleotide exchange factor, scl1714), pleckstrin (Rho interacting protein 3, scl1734), phosphatase (putative phosphatase, scl1827) and leucine-rich repeat (S-phase kinase-associated protein 2, scl2706), whose splice variants may generate proteins that have very different functions in cellular regulation than do their canonical forms.
Discussion
The transcriptome discovery strategy
Our aggressive approach to cDNA library normalization and subtraction, and the use of cDNA libraries from a wide range of tissues, has succeeded in providing the largest set of independent and distinct full-length cDNAs available for any species. In the early stages of this project, smaller clones were prioritized for full-length sequencing12. As the project progressed, larger cDNAs were selected to facilitate completion of the transcriptome. This places greater demands on the full-length cDNA synthesis process, and may potentially result in truncated transcripts. The successful application of library construction strategies developed by the RIKEN team has allowed a high success rate in identification of full-length transcripts. This is evidenced by our identification of many new cDNAs well over 4 kb in length, including those encoding G-protein-coupled receptors and ion channels whose sequences have been validated by mapping to the draft mouse genome.
One consequence of the selection for both rare transcripts and for longer cDNAs is that the largest class of new and novel cDNAs are non-coding. Our analysis shows that the majority (∼80%) of non-coding transcripts are not spliced. Sequence alone cannot distinguish genuine functional unspliced non-coding RNAs (such as Air28) from unspliced heterogeneous nuclear RNAs (hnRNAs) that might be included in libraries through the presence of an internal poly(A) sequence. The rigorous isolation of cytoplasmic mRNA43 introduced in later stages in this project may reduce the incidence of hnRNA if it does make a significant contribution. However, a disadvantage of targeting cytoplasmic RNAs is the assumption that all functional RNAs enter the cytosol. Such a strategy might also exclude non-coding RNAs that are active in the nucleus and that may have a regulatory role by interfering with splicing, polyadenylation or nuclear export. A large proportion of the non-coding transcripts are supported by independent ESTs. By definition, the ‘unclassifiable’ class represents singletons, but around 60% of ‘unclassifiable’ transcripts analysed thus far are detected reproducibly using cDNA microarrays (http://read.gsc.riken.go.jp). This finding supports the analysis of polyadenylation, which indicates that the majority are genuine mature products of transcription mediated by pol II. Overall, it is clear that non-coding RNAs are a major component of the mammalian transcriptome.
The size of the transcriptome and the proteome
Much of the discussion arising from completion of mouse and human genome sequences has revolved around estimating the number of genes. To analyse the draft transcriptome, we developed a stringent definition of a TU as a unit of genetic information transcribed into mRNA. Our analysis of the available sequence data from this project and of that previously reported in the public databases suggest that the mouse genome contains 37,086 TUs. However, it should be noted that even this is an incomplete survey of the transcriptional potential in mouse. For example, of the 22,444 protein-coding genes predicted with a high level of confidence in mouse Ensembl19, 3,074 were not covered by RTPS or by any RIKEN 5′ or 3′ EST sequencing.
The number of clusters is, of course, a function of the clustering parameters. Many larger clusters, some containing up to 90 representatives, may contain multiple representatives of gene families. Examples include the Gapd, RpL21, RpL29, Hmgb1, Rps2 and RpL7a clusters. The sequences in each of these clusters are very similar over their entire length, and readily meet stringent clustering cut-offs, but they clearly derive from different loci and should therefore be considered separate TUs. For example, Gapd is known to have 70 loci in the mouse genome (http://www.ncbi.nih.gov/LocusLink/list.cgi?V=1&Q=NM_008084).
As noted previously, there are a large number of 3′ and 5′ EST clusters from the phase 1 sequencing project that remain to be sequenced. To make a better estimate of the number of TUs in the mouse genome, we analysed the 547,149 5′ ESTs and 1,442,236 3′ ESTs from the phase 1 sequencing and non-RIKEN entries in dbEST. These ESTs were mapped to the draft mouse genome sequence. To count the number of additional TU candidates, we counted the number of EST clusters that were mapped outside the region of TUs determined by RTPS (Table 2), thereby increasing the estimated number of TUs in the mouse genome to around 70,000.
In addition, we have a number of reasons to suspect that our present estimate represents a conservative lower bound to the number of TUs in the mouse genome:
(1) Members of the FANTOM consortium and the RIKEN Genome Exploration Research Group continue to collaborate in the production of full-length cDNA libraries from novel tissue sources. In these continuing efforts, which use a very aggressive approach to normalization and subtraction of cDNA libraries, novel TUs are being discovered at a significant rate. Furthermore, much of our effort is now focused on inducible genes expressed in cell types such as macrophages, lymphocytes, dendritic cells, melanocytes and specialized cells in the central and peripheral nervous systems.
(2) Long mRNAs on the scale of genes such as titin44 and dystrophin45 are under-represented in all available cDNA libraries owing to the limitations in reverse transcription, cloning and stability of long cDNA clones; such genes are also very difficult to identify and annotate in genomic sequence. Continuing efforts are directed at selected cloning of longer mRNAs using novel RIKEN full-length cloning strategies46.
(3) Our definition of TUs did not account for the fact that some overlapping transcripts may encode distinct functions and may be independently regulated. Complete functional annotation of the genome will require a development of criteria that will allow such clusters to be separated. Transcript 6030402A12 (AK031286), which contributes a single TU to our estimate, contains at least 12 distinct transcripts that differ at both the 5′ and 3′ ends and consequently include quite different protein functional domains. Some of this variation has been noted previously47. By alignment to the mouse genome sequence, we can demonstrate that proteins that contain both N-terminal activation (LER) and repression (KRAB) domains—or proteins that lack either of these—arise from alternative promoter usage. Alternative polyadenylation (as well as internal exon skipping) yields C-terminal C2H2 zinc-finger arrays containing zero, 5, 6, 7, 8, 9, 17, 18 or 21 repeats in individual variant forms. It is not self-evident that the SCAN-KRAB zinc-finger locus contains a single TU (as defined here), or a single gene.
We can begin to infer that estimates of the number of TUs in the human genome based on EST assembly9 were not unrealistic. At least 41% of TUs encode more than one form of mRNA, consistent with computational analysis in the Human Alternative Splicing Database25 (http://www.bioinformatics.ucla.edu/~splice/HASDB/). This will certainly be an underestimate, because our approach sought to minimize the number of replicate sequences from a single TU. Furthermore, tissue-specific or infrequently spliced variants are sampled less frequently by the cloning strategy that we used, and might be eliminated through subtraction or clone selection. Alternative splicing cDNA libraries48 and oligonucleotide-based microarrays49 could be used to expand our knowledge of the impact of alternative splicing in the mouse transcriptome and proteome. Depending on how alternative 5′ ends, 3′ ends and cryptic exons are combined, the total number of transcripts will be at least twice the number of TUs.
Final comments
With the simultaneous, and synergistic, availability of genome and transcriptome sequences, we have reached a milestone in the era of large-scale data acquisition. We expect that additional mammalian transcriptome projects will be guided by the experience of the Mouse Gene Encyclopedia project, and that their outputs will support and refine the functional annotation of the TUs described here. As we continue this project, we hope that we can provide the tools necessary for a complete understanding of the biological processes that underlie mammalian development, growth, adaptation and disease. On the basis of the analysis presented here, we believe that a genome sequence alone can provide us with, at best, an incomplete picture of the transcriptional capacity of a complex genome. Looking forward, we anticipate that transcriptional analysis, both through sequencing and large-scale expression analysis, will be essential to the building of a complete picture of how genes and their products interact to form the regulatory, signalling and metabolic pathways that are the basis for complex life on Earth. Finally, although the mouse may be an imperfect model for the understanding of human biology, we believe that the resources made available through our efforts and those of the Mouse Genome Sequence Consortium—in combination with the exquisite genetic resources that are already available—will make the mouse the most appropriate animal for human studies for years to come.
Methods
Creation of the cDNA resource
Library construction, clone selection and sequencing is described in detail in Supplementary Information section 2. The cDNA sequence was assembled with the Phred/Phrap/Consed system, taking advantage of the anchor sequences of the 5′ and 3′ ends of cDNAs. Sequence and quality data for all assembled clones is available through DDBJ/EMBL/GenBank (Supplementary Information section 18), as well as through the FANTOM website (http://fantom2.gsc.riken.go.jp).
Mapping sequences to mouse draft genome
Sequences to be mapped were first screened using RepeatMasker (http://ftp.genome.washington.edu/RM/RepeatMasker.html). We aligned the full-length sequences with the draft mouse genome sequence (ftp://wolfram.wi.mit.edu/pub/mouse_contigs/MGSC_V3), using Paracel BLAST, with an E-value of 10-5. We extracted the genomic region of the best locus assigned to each cDNA (>95% identity over >100 bases), and used SIM4 (ref. 24) to align the cDNA to the genomic region. We aligned the repeat-masked EST sequences with the mouse genome, using BLAT50.
Clustering of FANTOM2 clone set and RTPS using ClusTrans
The 60,770 FANTOM2 clone set was first clustered using our original cDNA clustering program ClusTrans. Pairwise comparisons and global alignment were performed for all cDNAs using the SSEARCH program distributed with FASTA51,52, using the following parameters: match score, +1; mismatch score, -2; penalty for the first residue in a gap, -8; and penalty for additional residues in a gap, 0. After the global alignment, cDNA clusters were defined on the basis of percentage sequence identity and match length. The detailed criteria for defining a cluster are summarized in Supplementary Information section 19.
Creating an RTPS
To derive a comprehensive yet unique representative transcript and protein set (RTPS), the 60,770 FANTOM2 cDNA sequences, clustered initially with ClusTrans as above, were further clustered with 44,106 mouse transcript sequences compiled from the LocusLink, MGI (non-EST) and GenBank (non-EST) databases. The detailed strategy for construction of the RTPS is described in Supplementary Information sections 19 and 21.
Each cluster in the combined FANTOM2/public domain compilation defines a TU, and a single transcript sequence was chosen to represent each TU. For protein-coding TUs, a single polypeptide sequence was chosen to represent each TU with the following selection rules, longest SWISS-PROT record > longest RefSeq protein (NP_) record > longest CDS sequence. The number of protein sequences successfully translated and included in the RPS was 18,768. A variant-based proteome set (VPS) was prepared by identifying alternative transcription products that have different CDS sequences compared to the CDS sequences of cluster representatives. Each FANTOM2 clone with a unique CDS due to alternative transcript production, contributed to the VPS. The VPS contains all 18,768 protein sequences in the RPS plus 15,518 coding sequences.
Assignment of Gene Ontology terms
For genes represented in the MGI databases (http://www.informatics.jax.org/), we adopted GO terms that were assigned by MGI annotators53. In some cases, MATRICS annotators assigned GO terms. Terms were also assigned to TUs using translation tables of SWISS-PROT keywords, InterPro domains, and EC assignments (ftp://ftp.geneontology.org/pub/go/external2go) to GO. A preliminary SCOP (Structural Classification of Proteins; http://scop.wehi.edu.au/scop/) structural domain to GO translation table, which is continuously being refined, was created for this project (http://www.supfam.org/SUPERFAMILY/GO/). GO binning strategies are described in Supplementary Information section 21.
Analysis of alternative splice variants
To confirm the splice candidates in the 4,750 variant clusters, variant exons were compared to mouse mRNA sequences in the dbEST division of GenBank. dbEST sequences were mapped to the mouse genome as for the mapping of full-length cDNAs: the locus was identified using BLAT50, and intron/exon boundaries were identified using SIM4. ESTs which mapped with at least 95% identity to the genome, with every exon mapped at 95% identity or less than 5 gaps or mismatches, were interrogated for the presence of the exon forms observed in the cDNA data.
The impact of variant splicing on the translated protein sequences was assessed by comparing all variant clusters to the protein sequences of the FANTOM2 RTPS. cDNA-to-genome alignments were used to determine the genomic coordinate of the CDS start and end.
Databases, tools and systems
The databases, tools and systems used for the analyses in this paper are described in Supplementary Information section 20.
Enzymes are represented in boxes with their corresponding Enzyme Commission (EC) numbers, and are connected with their metabolites. EC numbers in unshaded boxes were found for both the public data set and the FANTOM2 clone set; numbers in black boxes are newly found in the FANTOM2 clone set. CoA, coenzyme A; ThPP, thiamine pyrophosphate.
References
International Human Genome Sequencing Consortium. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001)
Venter, J. C. et al. The sequence of the human genome. Science 291, 1304–1351 (2001)
Hogenesch, J. B. et al. A comparison of the Celera and Ensembl predicted gene sets reveals little overlap in novel genes. Cell 106, 413–415 (2001)
Daly, M. J. Estimating the human gene count. Cell 109, 283–284 (2002)
Kapranov, P. et al. Large-scale transcriptional activity in chromosomes 21 and 22. Science 296, 916–919 (2002)
Harrison, P. M. et al. Molecular fossils in the human genome: identification and analysis of the pseudogenes in chromosomes 21 and 22. Genome Res. 12, 272–280 (2002)
Reik, W. & Walter, J. Genomic imprinting: parental influence on the genome. Nature Rev. Genet. 2, 21–32 (2001)
Mattick, J. S. & Gagen, M. J. The evolution of controlled multitasked gene networks: the role of introns and other noncoding RNAs in the development of complex organisms. Mol. Biol. Evol. 18, 1611–1630 (2001)
Liang, F. et al. Gene index analysis of the human genome estimates approximately 120,000 genes. Nature Genet. 25, 239–240 (2000)
Saha, S. et al. Using the transcriptome to annotate the genome. Nature Biotechnol. 20, 508–512 (2002)
Camargo, A. A. et al. The contribution of 700,000 ORF sequence tags to the definition of the human transcriptome. Proc. Natl Acad. Sci. USA 98, 12103–12108 (2001)
The RIKEN Genome Exploration Research Group Phase II Team and the FANTOM Consortium. Functional annotation of a full-length mouse cDNA collection. Nature 409, 685–690 (2001)
Strausberg, R. L., Feingold, E. A., Klausner, R. D. & Collins, F. S. The mammalian gene collection. Science 286, 455–457 (1999)
Carninci, P. et al. Normalization and subtraction of cap-trapper-selected cDNAs to prepare full-length cDNA libraries for rapid discovery of new genes. Genome Res. 10, 1617–1630 (2000)
Carninci, P. et al. Balanced-size and long-size cloning of full-length, cap-trapped cDNAs into vectors of the novel lambda-FLC family allows enhanced gene discovery rate and functional analysis. Genomics 77, 79–90 (2001)
Konno, H. et al. Computer-based methods for the mouse full-length cDNA encyclopedia: real-time sequence clustering for construction of a nonredundant cDNA library. Genome Res. 11, 281–289 (2001)
Pruitt, K. D. & Maglott, D. R. RefSeq and LocusLink: NCBI gene-centered resources. Nucleic Acids Res. 29, 137–140 (2001)
Ashburner, M. et al. Gene ontology: tool for the unification of biology. Nature Genet. 25, 25–29 (2000)
Mouse Genome Sequencing Consortium. Initial sequencing and comparative analysis of the mouse genome. Nature (this issue)
Mural, R. J. et al. A comparison of whole-genome shotgun-derived mouse chromosome 16 and the human genome. Science 296, 1661–1671 (2002)
Pilpel, Y., Sudarsanam, P. & Church, G. M. Identifying regulatory networks by combinatorial analysis of promoter elements. Nature Genet. 29, 153–159 (2001)
Smale, S. T. Transcription initiation from TATA-less promoters within eukaryotic protein-coding genes. Biochim. Biophys. Acta 1351, 73–88 (1997)
Brett, D., Pospisil, H., Valcarcel, J., Reich, J. & Bork, P. Alternative splicing and genome complexity. Nature Genet. 30, 29–30 (2002)
Florea, L., Hartzell, G., Zhang, Z., Rubin, G. M. & Miller, W. A computer program for aligning a cDNA sequence with a genomic DNA sequence. Genome Res. 8, 967–974 (1998)
Modrek, B., Resch, A., Grasso, C. & Lee, C. Genome-wide detection of alternative splicing in expressed sequences of human genes. Nucleic Acids Res. 29, 2850–2859 (2001)
Pesole, G., Liuni, S. & D'Souza, M. PatSearch: a pattern matcher software that finds functional elements in nucleotide and protein sequences and assesses their statistical significance. Bioinformatics 16, 439–450 (2000)
Ferrigno, O. et al. Transposable B2 SINE elements can provide mobile RNA polymerase II promoters. Nature Genet. 28, 77–81 (2001)
Sleutels, F., Zwart, R. & Barlow, D. P. The non-coding Air RNA is required for silencing autosomal imprinted genes. Nature 415, 810–813 (2002)
Apweiler, R. et al. InterPro—an integrated documentation resource for protein families, domains and functional sites. Bioinformatics 16, 1145–1150 (2000)
Gough, J., Karplus, K., Hughey, R. & Chothia, C. Assignment of homology to genome sequences using a library of hidden Markov models that represent all proteins of known structure. J. Mol. Biol. 313, 903–919 (2001)
Kawaji, H. et al. Exploration of novel motifs derived from mouse cDNA sequences. Genome Res. 12, 367–378 (2002)
Murzin, A. G. Structural classification of proteins: new superfamilies. Curr. Opin. Struct. Biol. 6, 386–394 (1996)
Leipe, D. D., Wolf, Y. I., Koonin, E. V. & Aravind, L. Classification and evolution of P-loop GTPases and related ATPases. J. Mol. Biol. 317, 41–72 (2002)
Poirier, G. M. et al. Immune-associated nucleotide-1 (IAN-1) is a thymic selection marker and defines a novel gene family conserved in plants. J. Immunol. 163, 4960–4969 (1999)
Nielsen, H. & Krogh, A. Prediction of signal peptides and signal anchors by a hidden Markov model. Proc. Int. Conf. Intell. Syst. Mol. Biol. 6, 122–130 (1998)
Krogh, A., Larsson, B., von Heijne, G. & Sonnhammer, E. L. Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J. Mol. Biol. 305, 567–580 (2001)
The Gene Ontology Consortium Creating the gene ontology resource: design and implementation. Genome Res. 11, 1425–1433 (2001)
Miki, H., Setou, M., Kaneshiro, K. & Hirokawa, N. All kinesin superfamily protein, KIF, genes in mouse and human. Proc. Natl Acad. Sci. USA 98, 7004–7011 (2001)
Hirokawa, N. Kinesin and dynein superfamily proteins and the mechanism of organelle transport. Science 279, 519–526 (1998)
Weissman, A. M. Themes and variations on ubiquitylation. Nature Rev. Mol. Cell Biol. 2, 169–178 (2001)
Bairoch, A. The ENZYME database in 2000. Nucleic Acids Res. 28, 304–305 (2000)
Kanehisa, M., Goto, S., Kawashima, S. & Nakaya, A. The KEGG databases at GenomeNet. Nucleic Acids Res. 30, 42–46 (2002)
Carninci, P., Nakamura, M., Sato, K., Hayashizaki, Y. & Brownstein, M. J. Cytoplasmic RNA extraction from fresh and frozen mammalian tissues. Biotechniques 33, 306–309 (2002)
Bang, M. L. et al. The complete gene sequence of titin, expression of an unusual approximately 700-kDa titin isoform, and its interaction with obscurin identify a novel Z-line to I-band linking system. Circ. Res. 89, 1065–1072 (2001)
Koenig, M. et al. Complete cloning of the Duchenne muscular dystrophy (DMD) cDNA and preliminary genomic organization of the DMD gene in normal and affected individuals. Cell 50, 509–517 (1987)
Carninci, P., Shiraki, T., Mizuno, Y., Muramatsu, M. & Hayashizaki, Y. Extra-long first-strand cDNA synthesis. Biotechniques 32, 984–985 (2002)
Dreyer, S. D., Zheng, Q., Zabel, B., Winterpacht, A. & Lee, B. Isolation, characterization, and mapping of a zinc finger gene, ZFP95, containing both a SCAN box and an alternatively spliced KRAB A domain. Genomics 62, 119–122 (1999)
Schweighoffer, F. et al. Qualitative gene profiling: a novel tool in genomics and in pharmacogenomics that deciphers messenger RNA isoforms diversity. Pharmacogenomics 1, 187–197 (2000)
Shoemaker, D. D. et al. Experimental annotation of the human genome using microarray technology. Nature 409, 922–927 (2001)
Kent, W. J. BLAT—the BLAST-like alignment tool. Genome Res. 12, 656–664 (2002)
Smith, T. F. & Waterman, M. S. Identification of common molecular subsequences. J. Mol. Biol. 147, 195–197 (1981)
Pearson, W. R. Searching protein sequence libraries: comparison of the sensitivity and selectivity of the Smith-Waterman and FASTA algorithms. Genomics 11, 635–650 (1991)
Hill, D. P. et al. Program description: Strategies for biological annotation of mammalian systems: implementing gene ontologies in mouse genome informatics. Genomics 74, 121–128 (2001)
Acknowledgements
We thank the following (listed in alphabetical order) for discussion, encouragement and technical assistance: K. Abe, T. Akimura, T. Hanagaki, K. Hayashida, K. Hiramoto, T. Hiraoka, M. Honda, F. Hori, T. Huber, M. Izawa, H. Kato, M. Kohda, Y. Kojima, S. Koya, K. Koyama, C. Kurihara, J. Mashima, T. Matsuyama, M. Murata, K. Nishi, K. Nomura, R. Numazaki, M. Ohno, H. Saitou, C. Sakai, H. Sano, Y. Shibata, A. J. G. Simpson, Y. Sogabe, M. Tagami, A. Tagawa, F. Takahashi, S. Takaku, Y. Takeda, T. Tanaka, A. Tomaru, W. W. Wasserman, A. Watahiki, C. Wicking, H. Yamanishi, T. Yao, K. Yoshida. We especially thank A. Wada, M. Muramatsu, T. Ogawa, A. Kira, and all the members of RIKEN Yokohama Research Promotion Division for supporting and encouraging the project. We also thank the Laboratory for Genome Exploration Research Group for secretarial and technical assistance. This work was mainly supported by a research grant for the RIKEN Genome Exploration Research Project from the Ministry of Education, Culture, Sports, Science and Technology (MEXT) of the Japanese Government to Y.H., and by a Research and Development for Applying Advanced Computational Science and Technology (ACT) grant from the Japan Science and Technology Corporation to Y.H. and J.K.
Author information
Authors and Affiliations
Consortia
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 117 kb)
Rights and permissions
About this article
Cite this article
The FANTOM Consortium and the RIKEN Genome Exploration Research Group Phase I & II Team*. Analysis of the mouse transcriptome based on functional annotation of 60,770 full-length cDNAs. Nature 420, 563–573 (2002). https://doi.org/10.1038/nature01266
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature01266
This article is cited by
-
LncRNA MAFG-AS1 is involved in human cancer progression
European Journal of Medical Research (2023)
-
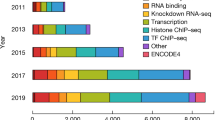
The status of the human gene catalogue
Nature (2023)
-
Germline genetic biomarkers to stratify patients for personalized radiation treatment
Journal of Translational Medicine (2022)
-
Notch-associated lncRNAs profiling circuiting epigenetic modification in colorectal cancer
Cancer Cell International (2022)
-
Scoring model based on the signature of non-m6A-related neoantigen-coding lncRNAs assists in immune microenvironment analysis and TCR-neoantigen pair selection in gliomas
Journal of Translational Medicine (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.