Abstract
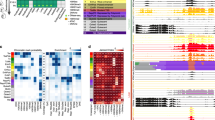
Chromosomes are divided into domains of open chromatin, where genes have the potential to be expressed, and domains of closed chromatin, where genes are not expressed1. Classic examples of open chromatin domains include ‘puffs’ on polytene chromosomes in Drosophila and extended loops from lampbrush chromosomes2,3. If multiple genes were typically expressed together from a single open chromatin domain, the position of co-expressed genes along the chromosomes would appear clustered. To investigate whether co-expressed genes are clustered, we examined the chromosomal positions of the genes expressed in muscle of Caenorhabditis elegans at the first larval stage. Here we show that co-expressed genes in C. elegans are clustered in groups of 2–5 along the chromosomes, suggesting that expression from a chromatin domain can extend over several genes. These observations reveal a higher-order organization of the structure of the genome, in which the order of genes along the chromosome is correlated with their expression in specific tissues.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Weintraub, H. Tissue-specific gene expression and chromatin structure. Harvey Lect. 79, 217–244 (1984)
Beermann, W. Control of differentiation at the chromosomal level. J. Exp. Zool. 157, 49–62 (1964)
Gall, J. G. & Callan, H. G. H3 Uridine incorporation in lampbrush chromosomes. Proc. Natl Acad. Sci. USA 48, 562–570 (1962)
Gorlach, M., Burd, C. G. & Dreyfuss, G. The mRNA poly(A)-binding protein: localization, abundance, and RNA-binding specificity. Exp. Cell Res. 211, 400–407 (1994)
Okkema, P. G., Harrison, S. W., Plunger, V., Aryana, A. & Fire, A. Sequence requirements for myosin gene expression and regulation in Caenorhabditis elegans. Genetics 135, 385–404 (1993)
Jones, A. R., Francis, R. & Schedl, T. GLD-1, a cytoplasmic protein essential for oocyte differentiation, shows stage- and sex-specific expression during Caenorhabditis elegans germline development. Dev. Biol. 180, 165–183 (1996)
Jiang, M. et al. Genome-wide analysis of developmental and sex-regulated gene expression profiles in Caenorhabditis elegans. Proc. Natl Acad. Sci. USA 98, 218–223 (2001)
Stein, L., Sternberg, P., Durbin, R., Thierry-Mieg, J. & Spieth, J. WormBase: network access to the genome and biology of Caenorhabditis elegans. Nucleic Acids Res. 29, 82–86 (2001)
Fukami-Kobayashi, K., Tomoda, S. & Go, M. Evolutionary clustering and functional similarity of RNA-binding proteins. FEBS Lett. 335, 289–293 (1993)
Blumenthal, T. Trans-splicing and polycistronic transcription in Caenorhabditis elegans. Trends Genet. 11, 132–136 (1995)
Blumenthal, T. Gene clusters and polycistronic transcription in eukaryotes. BioEssays 20, 480–487 (1998)
Reinke, V. et al. A global profile of germline gene expression in C. elegans. Mol. Cell 6, 605–616 (2000)
Kim, S. K. et al. A gene expression map for Caenorhabditis elegans. Science 293, 2087–2092 (2001)
Cohen, B. A., Mitra, R. D., Hughes, J. D. & Church, G. M. A computational analysis of whole-genome expression data reveals chromosomal domains of gene expression. Nature Genet. 26, 183–186 (2000)
Caron, H. et al. The human transcriptome map: clustering of highly expressed genes in chromosomal domains. Science 291, 1289–1292 (2001)
Hebbes, T. R., Clayton, A. L., Thorne, A. W. & Crane-Robinson, C. Core histone hyperacetylation co-maps with generalized DNase I sensitivity in the chicken β-globin chromosomal domain. EMBO J. 13, 1823–1830 (1994)
Stalder, J. et al. Tissue-specific DNA cleavages in the globin chromatin domain introduced by DNAase I. Cell 20, 451–460 (1980)
Goodwin, E. B., Okkema, P. G., Evans, T. C. & Kimble, J. Translational regulation of tra-2 by its 3′ untranslated region controls sexual identity in C. elegans. Cell 75, 329–339 (1993)
Wang, E., Miller, L. D., Ohnmacht, G. A., Liu, E. T. & Marincola, F. M. High-fidelity mRNA amplification for gene profiling. Nature Biotechnol. 18, 457–459 (2000)
Tenenbaum, S. A., Carson, C. C., Lager, P. J. & Keene, J. D. Identifying mRNA subsets in messenger ribonucleoprotein complexes by using cDNA arrays. Proc. Natl Acad. Sci. USA 97, 14085–14090 (2000)
Acknowledgements
We thank J. Shaw, K. Mach, M. Laub, J. Lieb, M. Hiller, E. Harmon and J. Wang for discussions; T. Blumenthal and J. Wang for sharing unpublished results; J. Wang for wild-type poly(A) mRNA; and M. Kiraly for assistance with the Supplementary Information. We also thank the programmers at the Stanford Microarray Database for microarray analysis and database management, and Proteome and Wormbase for annotation of C. elegans genes. P.J.R. is a Stanford University Beckman Fellow. This work was supported by a Human Frontiers Fellowship (P.J.R.) and grants from the National Institutes of Health (S.K.K.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Roy, P., Stuart, J., Lund, J. et al. Chromosomal clustering of muscle-expressed genes in Caenorhabditis elegans. Nature 418, 975–979 (2002). https://doi.org/10.1038/nature01012
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature01012
This article is cited by
-
Impact of poly(A)-tail G-content on Arabidopsis PAB binding and their role in enhancing translational efficiency
Genome Biology (2019)
-
QTL analysis of cocoon shell weight identifies BmRPL18 associated with silk protein synthesis in silkworm by pooling sequencing
Scientific Reports (2017)
-
A complete annotation of the chromosomes of the cellulase producer Trichoderma reesei provides insights in gene clusters, their expression and reveals genes required for fitness
Biotechnology for Biofuels (2016)
-
Histone modifications facilitate the coexpression of bidirectional promoters in rice
BMC Genomics (2016)
-
Overlapping cell population expression profiling and regulatory inference in C. elegans
BMC Genomics (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.