Abstract
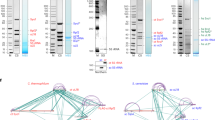
Although the U3 small nucleolar RNA (snoRNA), a member of the box C/D class of snoRNAs, was identified with the spliceosomal small nuclear RNAs (snRNAs) over 30 years ago1,2, its function and its associated protein components have remained more elusive. The U3 snoRNA is ubiquitous in eukaryotes and is required for nucleolar processing of pre-18S ribosomal RNA in all organisms where it has been tested3,4. Biochemical and genetic analyses suggest that U3–pre-rRNA base-pairing interactions mediate endonucleolytic pre-rRNA cleavages3. Here we have purified a large ribonucleoprotein (RNP) complex from Saccharomyces cerevisiae that contains the U3 snoRNA and 28 proteins. Seventeen new proteins (Utp1–17) and Rrp5 were present, as were ten known components. The Utp proteins are nucleolar and specifically associated with the U3 snoRNA. Depletion of the Utp proteins impedes production of the 18S rRNA, indicating that they are part of the active pre-rRNA processing complex. On the basis of its large size (80S; calculated relative molecular mass of at least 2,200,000) and function, this complex may correspond to the terminal knobs present at the 5′ ends of nascent pre-rRNAs. We have termed this large RNP the small subunit (SSU) processome.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Weinberg, R. A. & Penman, S. Small molecular weight monodisperse nuclear RNA. J. Mol. Biol. 38, 289–304 (1968)
Nakamura, T., Prestayko, A. W. & Busch, H. Studies on nucleolar 4 to 6S ribonucleic acid of Novikoff hepatoma cells. J. Biol. Chem. 243, 1368–1375 (1968)
Venema, J. & Tollervey, D. Ribosome synthesis in Saccharomyces cerevisiae. Annu. Rev. Genet. 33, 261–311 (1999)
Maxwell, E. S. & Fournier, M. J. The small nucleolar RNAs. Ann. Rev. Biochem. 64, 897–934 (1995)
Watkins, N. J. et al. A common core RNP structure shared between the small nucleolar box C/D RNPs and the spliceosomal U4 snRNP. Cell 103, 457–466 (2000)
Lubben, B., Marshallsay, C., Rottman, N. & Luhrmann, R. Isolation of U3 snoRNP from CHO cells: a novel 55 kDa protein binds to the central part of U3 snoRNA. Nucleic Acids Res. 21, 5377–5385 (1993)
Rigaut, G., Shevchenko, A., Rutz, B., Wilm, M. & Seraphin, B. A generic protein purification method for protein complex characterization and proteome exploration. Nature Biotechnol. 17, 1030–1032 (1999)
Wiederkehr, T., Pretot, R. F. & Minvielle-Sebastia, L. Synthetic lethal interactions with conditional poly(A) polymerase alleles identify LCP5, a gene involved in 18S rRNA maturation. RNA 4, 1357–1372 (1998)
Billy, E., Wegierski, T., Nasr, F. & Filipowicz, W. Rcl1p, the yeast protein similar to the RNA 3′-phosphate cyclase, associates with U3 snoRNP and is required for 18S rRNA biogenesis. EMBO J. 19, 2115–2126 (2000)
Wegierski, T., Billy, E., Nasr, F. & Filipowicz, W. Bms1p, a G-domain-containing protein, associates with Rcl1p and is required for 18S rRNA biogenesis in yeast. RNA 7, 1254–1267 (2001)
Chung, S., McLean, M. R. & Rymond, B. C. Yeast ortholog of the Drosophila crooked neck protein promotes spliceosome assembly through stable U4/U6.U5 snRNP addition. RNA 5, 1042–1054 (1999)
Shou, W. et al. Exit from mitosis is triggered by Tem1-dependent release of the protein phosphatase Cdc14 from nucleolar RENT complex. Cell 97, 233–244 (1999)
Kamakaka, R. & Rine, J. Sir- and silencer-independent disruption of silencing in Saccharomyces by Sas10p. Genetics 149, 903–914 (1998)
Fournier, M. J. & Maxwell, E. S. The nucleolar snRNAs: catching up with the spliceosomal snRNAs. Trends Biochem. Sci. 18, 131–135 (1993)
Venema, J. & Tollervey, D. Processing of pre-ribosomal RNA in Saccharomyces cerevisiae. Yeast 11, 1629–1650 (1995)
Miller, O. L. Jr & Beatty, B. R. Visualization of nucleolar genes. Science 164, 955–957 (1969)
Mougey, E. B. et al. The terminal balls characteristic of eucaryotic rRNA transcription units in chromatin spread are rRNA processing complexes. Genes Dev. 7, 1609–1619 (1993)
Lee, S. J. & Baserga, S. J. Functional separation of pre-rRNA processing steps revealed by truncation of the U3 small nucleolar ribonucleoprotein component, Mpp10. Proc. Natl Acad. Sci. USA 94, 13536–13541 (1997)
Wehner, K. A. & Baserga, S. J. The σ70-like motif: a eukaryotic RNA binding domain unique to a superfamily of proteins required for ribosome biogenesis. Mol. Cell 9, 329–339 (2002)
Warner, J. R. Nascent ribosomes. Cell 107, 133–136 (2001)
Knop, M. et al. Epitope tagging of yeast genes using a PCR-based strategy: more tags and improved practical routines. Yeast 15, 963–972 (1999)
Longtine, M. S. et al. Additional modules for versatile and economical PCR-based gene deletion and modification in Saccharomyces cerevisiae. Yeast 14, 953–961 (1998)
Schneider, B. L., Seufert, W., Steiner, B., Yang, Q. H. & Futcher, A. B. Use of polymerase chain reaction epitope tagging for protein tagging in Saccharomyces cerevisiae. Yeast 11, 1265–1274 (1995)
Ross-MacDonald, P. et al. Large-scale analysis of the yeast genome by transposon tagging and gene disruption. Nature 402, 413–418 (1999)
Dunbar, D. A., Dragon, F., Lee, S. J. & Baserga, S. J. A nucleolar protein related to ribosomal protein L7 is required for an early step in large ribosomal subunit biogenesis. Proc. Natl Acad. Sci. USA 97, 13027–13032 (2000)
Dunbar, D. A., Wormsley, S., Agentis, T. M. & Baserga, S. J. Mpp10p, a U3 small nucleolar ribonucleoprotein component required for pre-18S rRNA processing in yeast. Mol. Cell. Biol. 17, 5803–5812 (1997)
Lee, S. J. & Baserga, S. J. Imp3p and Imp4p: two specific components of the U3 small nucleolar ribonucleoprotein that are essential for pre-18S rRNA processing. Mol. Cell. Biol. 19, 5441–5452 (1999)
Hughes, J. M. X. & Ares, M. Jr Depletion of U3 small nucleolar RNA inhibits cleavage in the 5′ external transcribed spacer of yeast pre-ribosomal RNA and prevents formation of 18S ribosomal RNA. EMBO J. 10, 4231–4239 (1991)
Samarsky, D. A. & Fournier, M. J. Functional mapping of the U3 small nucleolar RNA from the yeast Saccharomyces cerevisiae. Mol. Cell. Biol. 18, 3431–3444 (1998)
Osheim, Y. N. & Beyer, A. L. Electron microscopy of RNP complexes on nascent RNA using the Miller chromatin spreading method. Methods Enzymol. 180, 481–509 (1989)
Acknowledgements
We thank A. Djikeng, G. Dreyfuss, P. Gordon, J. Laney, T. Serio and T. Stone for assistance and advice. We also thank P. Glazer, M. Snyder and J. Steitz for critical reading of the manuscript. F.D. was supported by the Anna Fuller Fund for Molecular Oncology. This work was supported by federal grants from the NIH to D.F.H and S.J.B. and the NSF to A.L.B., and by The Patterson Trust to S.J.B.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Rights and permissions
About this article
Cite this article
Dragon, F., Gallagher, J., Compagnone-Post, P. et al. A large nucleolar U3 ribonucleoprotein required for 18S ribosomal RNA biogenesis. Nature 417, 967–970 (2002). https://doi.org/10.1038/nature00769
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature00769
This article is cited by
-
RRP9 promotes gemcitabine resistance in pancreatic cancer via activating AKT signaling pathway
Cell Communication and Signaling (2022)
-
Transcriptomic analysis of formic acid stress response in Saccharomyces cerevisiae
World Journal of Microbiology and Biotechnology (2022)
-
Transcriptomic analysis reveals process of autolysis of Kluyveromyces marxianus in vacuum negative pressure and the higher temperature
International Microbiology (2022)
-
Nanopore sequencing reveals endogenous NMD-targeted isoforms in human cells
Genome Biology (2021)
-
Puf6 primes 60S pre-ribosome nuclear export at low temperature
Nature Communications (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.