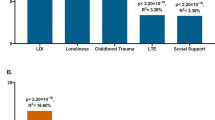
Abstract
The hypothesis that the S allele of the 5-HTTLPR serotonin transporter promoter region is associated with increased risk of depression, but only in individuals exposed to stressful situations, has generated much interest, research and controversy since first proposed in 2003. Multiple meta-analyses combining results from heterogeneous analyses have not settled the issue. To determine the magnitude of the interaction and the conditions under which it might be observed, we performed new analyses on 31 data sets containing 38 802 European ancestry subjects genotyped for 5-HTTLPR and assessed for depression and childhood maltreatment or other stressful life events, and meta-analysed the results. Analyses targeted two stressors (narrow, broad) and two depression outcomes (current, lifetime). All groups that published on this topic prior to the initiation of our study and met the assessment and sample size criteria were invited to participate. Additional groups, identified by consortium members or self-identified in response to our protocol (published prior to the start of analysis) with qualifying unpublished data, were also invited to participate. A uniform data analysis script implementing the protocol was executed by each of the consortium members. Our findings do not support the interaction hypothesis. We found no subgroups or variable definitions for which an interaction between stress and 5-HTTLPR genotype was statistically significant. In contrast, our findings for the main effects of life stressors (strong risk factor) and 5-HTTLPR genotype (no impact on risk) are strikingly consistent across our contributing studies, the original study reporting the interaction and subsequent meta-analyses. Our conclusion is that if an interaction exists in which the S allele of 5-HTTLPR increases risk of depression only in stressed individuals, then it is not broadly generalisable, but must be of modest effect size and only observable in limited situations.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Moussavi S, Chatterji S, Verdes E, Tandon A, Patel V, Ustun B . Depression, chronic diseases, and decrements in health: results from the World Health Surveys. Lancet 2007; 370: 851–858.
Ustun TB, Ayuso-Mateos JL, Chatterji S, Mathers C, Murray CJ . Global burden of depressive disorders in the year 2000. Br J Psychiatry 2004; 184: 386–392.
Sullivan PF, Neale MC, Kendler KS . Genetic epidemiology of major depression: review and meta-analysis. Am J Psychiatry 2000; 157: 1552–1562.
Bierut LJ, Heath AC, Bucholz KK, Dinwiddie SH, Madden PA, Statham DJ et al. Major depressive disorder in a community-based twin sample: are there different genetic and environmental contributions for men and women? Arch Gen Psychiatry 1999; 56: 557–563.
Kendler KS, Neale MC, Kessler RC, Heath AC, Eaves LJ . The lifetime history of major depression in women. Reliability of diagnosis and heritability. Arch Gen Psychiatry 1993; 50: 863–870.
Kendler KS, Gardner CO, Prescott CA . Are there sex differences in the reliability of a lifetime history of major depression and its predictors? Psychol Med 2001; 31: 617–625.
McGuffin P, Katz R, Rutherford J . Nature, nurture and depression: a twin study. Psychol Med 1991; 21: 329–335.
McGuffin P, Katz R, Watkins S, Rutherford J . A hospital-based twin register of the heritability of DSM-IV unipolar depression. Arch Gen Psychiatry 1996; 53: 129–136.
Ripke S, Wray NR, Lewis CM, Hamilton SP, Weissman MM, Breen G et al. A mega-analysis of genome-wide association studies for major depressive disorder. Mol Psychiatry 2013; 18: 497–511.
Flint J, Kendler KS . The genetics of major depression. Neuron 2014; 81: 484–503.
Hyde CL, Nagle MW, Tian C, Chen X, Paciga SA, Wendland JR et al. Identification of 15 genetic loci associated with risk of major depression in individuals of European descent. Nat Genet 2016; 48: 1031–1036.
Dunn EC, Brown RC, Dai Y, Rosand J, Nugent NR, Amstadter AB et al. Genetic determinants of depression: recent findings and future directions. Harv Rev Psychiatry 2015; 23: 1–18.
Kendler KS, Kessler RC, Walters EE, MacLean C, Neale MC, Heath AC et al. Stressful life events, genetic liability, and onset of an episode of major depression in women. Am J Psychiatry 1995; 152: 833–842.
Cohen-Woods S, Craig IW, McGuffin P . The current state of play on the molecular genetics of depression. Psychol Med 2013; 43: 673–687.
Caspi A, Sugden K, Moffitt TE, Taylor A, Craig IW, Harrington H et al. Influence of life stress on depression: moderation by a polymorphism in the 5-HTT gene. Science 2003; 301: 386–389.
Lesch KP, Greenberg MD, Higley JD, Bennett A, Murphy DL . Serotonin transporter, personality, and behavior: toward a disection of gene-gene and gene-environment interaction. In: Benjamin J, Ebstein RP, Belmaker RH (eds). Molecular Genetics and the Human Personality. American Psychiatric Association. APA: Washington, DC, 2002, pp 109–136.
Lesch KP, Bengel D, Heils A, Sabol SZ, Greenberg BD, Petri S et al. Association of anxiety-related traits with a polymorphism in the serotonin transporter gene regulatory region. Science 1996; 274: 1527–1531.
McClelland GH, Judd CM . Statistical difficulties of detecting interactions and moderator effects. Psychol Bull 1993; 114: 376–390.
Bosker FJ, Hartman CA, Nolte IM, Prins BP, Terpstra P, Posthuma D et al. Poor replication of candidate genes for major depressive disorder using genome-wide association data. Mol Psychiatry 2011; 16: 516–532.
Need AC, Ge D, Weale ME, Maia J, Feng S, Heinzen EL et al. A genome-wide investigation of SNPs and CNVs in schizophrenia. PLoS Genet 2009; 5: e1000373.
Duncan LE, Keller MC . A critical review of the first 10 years of candidate gene-by-environment interaction research in psychiatry. Am J Psychiatry 2011; 168: 1041–1049.
Risch N, Herrell R, Lehner T, Liang KY, Eaves L, Hoh J et al. Interaction between the serotonin transporter gene (5-HTTLPR), stressful life events, and risk of depression: a meta-analysis. JAMA 2009; 301: 2462–2471.
Munafo MR, Durrant C, Lewis G, Flint J . Gene X environment interactions at the serotonin transporter locus. Biol Psychiatry 2009; 65: 211–219.
Karg K, Burmeister M, Shedden K, Sen S . The serotonin transporter promoter variant (5-HTTLPR), stress, and depression meta-analysis revisited: evidence of genetic moderation. Arch Gen Psychiatry 2011; 68: 444–454.
Sharpley CF, Palanisamy SK, Glyde NS, Dillingham PW, Agnew LL . An update on the interaction between the serotonin transporter promoter variant (5-HTTLPR), stress and depression, plus an exploration of non-confirming findings. Behav Brain Res 2014; 273: 89–105.
Brown GW, Harris TO . Depression and the serotonin transporter 5-HTTLPR polymorphism: a review and a hypothesis concerning gene-environment interaction. J Affect Disord 2008; 111: 1–12.
Munafo MR, Flint J . Replication and heterogeneity in gene x environment interaction studies. Int J Neuropsychopharmacol 2009; 12: 727–729.
Uher R, Caspi A, Houts R, Sugden K, Williams B, Poulton R et al. Serotonin transporter gene moderates childhood maltreatment's effects on persistent but not single-episode depression: replications and implications for resolving inconsistent results. J Affect Disord 2011; 135: 56–65.
Culverhouse RC, Bowes L, Breslau N, Nurnberger JI Jr., Burmeister M, Fergusson DM et al. Protocol for a collaborative meta-analysis of 5-HTTLPR, stress, and depression. BMC Psychiatry 2013; 13: 304.
Boyd A, Golding J, Macleod J, Lawlor DA, Fraser A, Henderson J et al. Cohort Profile: the 'children of the 90s'—the index offspring of the Avon Longitudinal Study of Parents and Children. Int J Epidemiol 2013; 42: 111–127.
Stefanis NC, Mandelli L, Hatzimanolis A, Zaninotto L, Smyrnis N, Avramopoulos D et al. Serotonin transporter gene variants and prediction of stress-induced risk for psychological distress. Genes Brain Behav 2011; 10: 536–541.
Edwards BHM, Letcher P, Little K, Macdonald J, Oberklaid F, O'Connor M et al, The Australian Temperament Project: The first 30 years. Australian Institute of Family Studies: Commonwealth of Australia, 2013.
Fergusson DM, Horwood LJ The Christchurch Health and Development Study. In: Joyce P, Nicholls G, Thomas K, Wilkinson T (eds). The Christchurch Experience: 40 Years of Research and Teaching. University of Otago: Christchurch, 2013, pp79–87.
Bierut LJ, Madden PA, Breslau N, Johnson EO, Hatsukami D, Pomerleau OF et al. Novel genes identified in a high-density genome wide association study for nicotine dependence. Hum Mol Genet 2007; 16: 24–35.
Cohen-Woods S, Gaysina D, Craddock N, Farmer A, Gray J, Gunasinghe C et al. Depression Case Control (DeCC) Study fails to support involvement of the muscarinic acetylcholine receptor M2 (CHRM2) gene in recurrent major depressive disorder. Hum Mol Genet 2009; 18: 1504–1509.
Surtees PG, Wainwright NW, Willis-Owen SA, Luben R, Day NE, Flint J . Social adversity, the serotonin transporter (5-HTTLPR) polymorphism and major depressive disorder. Biol Psychiatry 2006; 59: 224–229.
Ritchie K, Artero S, Beluche I, Ancelin ML, Mann A, Dupuy AM et al. Prevalence of DSM-IV psychiatric disorder in the French elderly population. Br J Psychiatry 2004; 184: 147–152.
Eley TC, Sugden K, Corsico A, Gregory AM, Sham P, McGuffin P et al. Gene-environment interaction analysis of serotonin system markers with adolescent depression. Mol Psychiatry 2004; 9: 908–915.
Etain B, Lajnef M, Henrion A, Dargel AA, Stertz L, Kapczinski F et al. Interaction between SLC6A4 promoter variants and childhood trauma on the age at onset of bipolar disorders. Sci Rep 2015; 5: 16301.
Penas-Lledo E, Guillaume S, Naranjo ME, Delgado A, Jaussent I, Blasco-Fontecilla H et al. A combined high CYP2D6-CYP2C19 metabolic capacity is associated with the severity of suicide attempt as measured by objective circumstances. Pharmacogenomics J 2015; 15: 172–176.
Otte C, McCaffery J, Ali S, Whooley MA . Association of a serotonin transporter polymorphism (5-HTTLPR) with depression, perceived stress, and norepinephrine in patients with coronary disease: the Heart and Soul Study. Am J Psychiatry 2007; 164: 1379–1384.
Laucht M, Treutlein J, Blomeyer D, Buchmann AF, Schmid B, Becker K et al. Interaction between the 5-HTTLPR serotonin transporter polymorphism and environmental adversity for mood and anxiety psychopathology: evidence from a high-risk community sample of young adults. Int J Neuropsychoph 2009; 12: 737–747.
Zucker RA, Ellis DA, Fitzgerald HE, Bingham CR, Sanford K . Other evidence for at least two alcoholisms.2. Life course variation in antisociality and heterogeneity of alcoholic outcome. Dev Psychopathol 1996; 8: 831–848.
Juhasz G, Gonda X, Hullam G, Eszlari N, Kovacs D, Lazary J et al. Variability in the effect of 5-HTTLPR on depression in a large European population: the role of age, symptom profile, type and intensity of life stressors. PLoS ONE 2015; 10: e0116316.
Anstey KJ, Christensen H, Butterworth P, Easteal S, Mackinnon A, Jacomb T et al. Cohort profile: the PATH through life project. Int J Epidemiol 2012; 41: 951–960.
Scheid JM, Holzman CB, Jones N, Friderici KH, Nummy KA, Symonds LL et al. Depressive symptoms in mid-pregnancy, lifetime stressors and the 5-HTTLPR genotype. Genes Brain Behav 2007; 6: 453–464.
Coventry WL, James MR, Eaves LJ, Gordon SD, Gillespie NA, Ryan L et al. Do 5HTTLPR and stress interact in risk for depression and suicidality? Item response analyses of a large sample. Am J Med Genet B Neuropsychiatr Genet 2010; 153B: 757–765.
Sjoberg RL, Nilsson KW, Nordquist N, Ohrvik J, Leppert J, Lindstrom L et al. Development of depression: sex and the interaction between environment and a promoter polymorphism of the serotonin transporter gene. Int J Neuropsychopharmacol 2006; 9: 443–449.
Aslund C, Leppert J, Comasco E, Nordquist N, Oreland L, Nilsson KW . Impact of the interaction between the 5HTTLPR polymorphism and maltreatment on adolescent depression. A population-based study. Behav Genet 2009; 39: 524–531.
Goldman N, Glei DA, Lin YH, Weinstein M . The serotonin transporter polymorphism (5-HTTLPR): allelic variation and links with depressive symptoms. Depress Anxiety 2010; 27: 260–269.
Volzke H, Alte D, Schmidt CO, Radke D, Lorbeer R, Friedrich N et al. Cohort profile: the study of health in Pomerania. Int J Epidemiol 2011; 40: 294–307.
Oldehinkel AJ, Rosmalen JG, Buitelaar JK, Hoek HW, Ormel J, Raven D et al. Cohort profile update: the TRacking Adolescents' Individual Lives Survey (TRAILS). Int J Epidemiol 2015; 44: 76–76n.
Mandelli L, Serretti A, Marino E, Pirovano A, Calati R, Colombo C . Interaction between serotonin transporter gene, catechol-O-methyltransferase gene and stressful life events in mood disorders. Int J Neuropsychopharmacol 2007; 10: 437–447.
Carli V, Mandelli L, Zaninotto L, Roy A, Recchia L, Stoppia L et al. A protective genetic variant for adverse environments? The role of childhood traumas and serotonin transporter gene on resilience and depressive severity in a high-risk population. Eur Psychiatry 2011; 26: 471–478.
Olsson CA, Foley DL, Parkinson-Bates M, Byrnes G, McKenzie M, Patton GC et al. Prospects for epigenetic research within cohort studies of psychological disorder: a pilot investigation of a peripheral cell marker of epigenetic risk for depression. Biol Psychol 2010; 83: 159–165.
Baune BT, Air TM . Clinical, functional and biological correlates of cognitive dimensions in major depressive disorder - rationale, design, and characteristics of the Cognitive Function and Mood Study (CoFaM-Study). Front Psychiatry 2016; 7: 150, in press.
Peyrot WJ, Middeldorp CM, Jansen R, Smit JH, de Geus EJ, Hottenga JJ et al. Strong effects of environmental factors on prevalence and course of major depressive disorder are not moderated by 5-HTTLPR polymorphisms in a large Dutch sample. J Affect Disord 2013; 146: 91–99.
Reich T . A genomic survey of alcohol dependence and related phenotypes: results from the Collaborative Study on the Genetics of Alcoholism (COGA). Alcohol Clin Exp Res 1996; 20: 133A–137A.
Krug EG, Mercy JA, Dahlberg LL, Zwi AB . The world report on violence and health. Lancet 2002; 360: 1083–1088.
Hovens JG, Giltay EJ, van Hemert AM, Penninx BW . Childhood maltreatment and the course of depressive and anxiety disorders: the contribution of personality characteristics. Depress Anxiety 2016; 33: 27–34.
Fergusson DM, Boden JM, Horwood LJ . Exposure to childhood sexual and physical abuse and adjustment in early adulthood. Child Abuse Negl 2008; 32: 607–619.
R Development Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing: Vienna, Austria, 2008.
Lumley T . rmeta: Meta-analysis. R package version 2.16. 2012.
Veichtbauer W . Conducting meta-analyses in R with the metafor package. J Stat Soft 2010; 36: 48.
SAS/STAT. 9.1 edn. SAS Institute Inc.: Cary, NC, USA, 2002-2003.
Open Science C. PSYCHOLOGY. Estimating the reproducibility of psychological science. Science 2015; 349: aac4716.
Fergusson DM, Horwood LJ, Miller AL, Kennedy MA . Life stress, 5-HTTLPR and mental disorder: findings from a 30-year longitudinal study. Br J Psychiatry 2011; 198: 129–135.
Moffitt TE, Caspi A . Bias in a protocol for a meta-analysis of 5-HTTLPR, stress, and depression. BMC Psychiatry 2014; 14: 179.
Button KS, Ioannidis JP, Mokrysz C, Nosek BA, Flint J, Robinson ES et al. Power failure: why small sample size undermines the reliability of neuroscience. Nat Rev Neurosci 2013; 14: 365–376.
Ioannidis JP . Why most discovered true associations are inflated. Epidemiology 2008; 19: 640–648.
Gonda X, Eszlari N, Kovacs D, Anderson IM, Deakin JF, Juhasz G et al. Financial difficulties but not other types of recent negative life events show strong interactions with 5-HTTLPR genotype in the development of depressive symptoms. Transl Psychiatry 2016; 6: e798.
Brown GW, Ban M, Craig TK, Harris TO, Herbert J, Uher R . Serotonin transporter length polymorphism, childhood maltreatment, and chronic depression: a specific gene-environment interaction. Depress Anxiety 2013; 30: 5–13.
Mandelli L, Petrelli C, Serretti A . The role of specific early trauma in adult depression: a meta-analysis of published literature. Childhood trauma and adult depression. Eur Psychiatry 2015; 30: 665–680.
Acknowledgements
Acknowledgment of funding: Funding supporting the participation of the contributing studies was as follows:
ALSPAC: Grant 102215/2/13/2 from The Wellcome Trust and grant MC_UU_12013/6 from the UK Medical Research Council. The University of Bristol also provides core support for ALSPAC. LB receives funding as an Early Career Research Fellow from the Leverhulme Trust. MRM is a member of the UK Centre for Tobacco and Alcohol Studies, a UK Clinical Research Council Public Health Research: Centre of Excellence. Funding from British Heart Foundation, Cancer Research UK, Economic and Social Research Council, Medical Research Council, and the National Institute for Health Research, under the auspices of the UK Clinical Research Collaboration, is gratefully acknowledged. ASPIS: EKBAN 97 from the General Secretariat of Research and Technology, Greek Ministry of Development. ATP: Grants DP130101459, DP160103160 and APP1082406 from the Australian Research Council and The National Health and Medical Research Council of Australia. CHDS: Grant HRC 11/792 from the Health Research Council of New Zealand. CoFaMS: Grant APP1060524 to BTB from the National Health and Medical Research Council of Australia. We acknowledge the University of Adelaide for the provision of seed funding in support of this project. COGA: Grant U10AA008401 from the National Institutes of Health, NIAAA and NIDA. COGEND: National Institutes of Health grants P01CA089392 from NCI and R01DA036583 from NIDA. DeCC: Grant G0701420 from the UK Medical Research Council, and a UK MRC Population Health Scientist fellowship (G1002366) and an MQ Fellows Award (MQ14F40) to Helen L Fisher. EPIC-Norfolk: Grants G9502233, G0300128, C865/A2883 from the UK Medical Research Council and Cancer Research UK. ESPRIT Montpellier: An unconditional grant from Novartis and from the National Research Agency (ANR Project 07 LVIE004). G1219: A project grant from the WT Grant Foundation and G120/635, a Career Development Award from the UK Medical Research Council to Thalia Eley. The GENESiS project was supported by Grant G9901258 from the UK Medical Research Council. This study presents independent research part- funded by the National Institute for Health Research (NIHR) Biomedical Research Centre at South London and Maudsley NHS Foundation Trust and King’s College London. The views expressed are those of the author(s) and not necessarily those of the NHS, the NIHR or the Department of Health. GAN12-France: Research Protocol C0829 from INSERM; Research Protocol GAN12 from Assistance Publique des Hôpitaux de Paris; ANR-11-IDEX- 0004 from Investissements d’Avenir program managed by the ANR, and RTRS Sante Mentale from Fondation FondaMental. GENESIS: Grant PHRC UF 7653 & ANR NEURO 2007 ‘GENESIS’ from CHU Montpellier & Agence Nationale de la Recherche. Heart and Soul: Epidemiology Merit Review Program from the Department of Veterans Affairs; National Institutes of Health grant R01HL-079235 from NHLBI; Generalist Physician Faculty Scholars Program from the Robert Woods Johnson foundation; Paul Beeson Faculty Scholars Program from the American Federation for Aging Research; and a Young Investigator Award from the Bran and Behavior Research Foundation. MARS: Grant LA 733/2-1 from German Research Foundation (DFG) and the Federal Ministry for Education and Research as part of the 'National Genome Research Network'. MLS: National Institutes of Health grants R01 AA07065 and R37 AA07065 from NIAAA. MoodInFlame: Grant EU-FP7-HEALTH-F2-2008-222963 from the European Union. Muenster Neuroimaging Study: Grant FOR2107, DA1151/5-1 from the German Research Foundation (DFG). NEWMOOD: Grants LSHM-CT-2004-503474 from Sixth Framework Program of the European Union; KTIA_NAP_13-1-2013-0001, KTIA_13_NAP-A-II/14 from National Development Agency Hungarian Brain Research Program; KTIA_NAP_13-2-2015-0001 from MTA-SE-NAP B Genetic Brain Imaging Migraine Research Group, Hungarian Academy of Sciences, Semmelweis University; support from Hungarian Academy of Sciences, MTA-SE Neuropsychopharmacology and Neurochemistry Research Group; and support from the National Institute for Health Research Manchester Biomedical Research Centre. NESDA/NTR: The Netherlands Organization for Scientific Research (NWO) and MagW/ZonMW grants Middelgroot-911-09-032, Spinozapremie 56-464-14192, Geestkracht program of the Netherlands Organization for Health Research and Development (ZonMW 10-000-1002), Center for Medical Systems Biology (CSMB, NWO Genomics), Genetic influences on stability and change in psychopathology from childhood to young adulthood (ZonMW 912-10-020), NBIC/BioAssist/RK(2008.024), Biobanking and Biomolecular Resources Research Infrastructure (BBMRI -NL, 184.021.007), VU University's Institute for Health and Care Research (EMGO+) and Neuroscience Campus Amsterdam (NCA); the European Science Council (ERC Advanced, 230374). Part of the genotyping and analyses were funded by the Genetic Association Information Network (GAIN) of the Foundation for the National Institutes of Health, Rutgers University Cell and DNA Repository (NIMH U24 MH068457-06), the Avera Institute, Sioux Falls, South Dakota (USA) and the National Institutes of Health (NIH R01 HD042157-01A1, MH081802, Grand Opportunity grants 1RC2 MH089951 and 1RC2 MH089995). PATH: Program Grant Number 179805 from the National Health and Medical Research Council of Australia. POUCH: Grants 20FY01-38 and 20-FY04-37 of the Perinatal Epidemiologic Research Initiative Program Grant from the March of Dimes Foundation; National Institutes of Health grant R01 HD34543 from NICHD and NINR; grant 02816-7 from the Thrasher Research Foundation; and grant U01 DP000143-01 from the Centers for Disease Control and Prevention. QIMRtwin: Grants 941177, 971232, 339450, 443011 from the National Health and Medical Research Council of Australia; AA07535, AA07728, AA10249 from US Public Health Service; National Institutes of Health grant K99DA023549-01A2 from NIDA. Additional support was provided by Beyond Blue. SALVe 2001 and SALVe 2006: Grants FO2012-0326, FO2013-0023, FO2014-0243 from The Brain Foundation (Hjärnfonden); SLS-559921 from Söderström-Königska Foundation; 2015-00897 from Swedish Council for Working Life and Social Research; and M15-0239 from Åke Wiberg's Foundation. Additional funding was provided by Systembolagets Råd för Alkoholforskning, SRA and Svenska Spel Research Council. SEBAS: National Institutes of Health grants R01 AG16790, R01 AG16661 and R56 AG01661 from NIA and grant P2CHD047879 from NICHD; and additional financial support from the Graduate School of Arts and Sciences at Georgetown University. SHIP/TREND: This work was supported by the German Federal Ministry of Education and Research within the framework of the e:Med research and funding concept (Integrament) Grant No. 01ZX1314E. Study of Health in Pomerania is part of the Community Medicine Research net of the University of Greifswald, Germany, which is funded by the Federal Ministry of Education and Research Grant Nos. 01ZZ9603, 01ZZ0103 and 01ZZ0403; the Ministry of Cultural Affairs; and the Social Ministry of the Federal State of Mecklenburg-West Pomerania. Genome-wide data were supported by the Federal Ministry of Education and Research Grant No. 03ZIK012 and a joint grant from Siemens Healthcare, Erlangen, Germany and the Federal State of Mecklenburg-West Pomerania. The Greifswald Approach to Individualised Medicine (GANI_MED) was funded by the Federal Ministry of Education and Research Grant No. 03IS2061A and the German Research Foundation Grant No. GR 1912/5-1. TRAILS: Grants GB-MW 940-38-011, ZonMW Brainpower 100-001-004, Investment grant 175.010.2003.005, GB-MaGW 480-07-001 and Longitudinal Survey and Panel Funding 481-08-013 from the Netherlands Organization for Scientific Research (NWO). Additional funding was provided by the Dutch Ministry of Justice, the European Science Foundation, BBMRI-NL and the participating centres (UMCG, RUG, Erasmus MC, UU, Radboud MC, Parnassia Bavo group): VAHCS: Grants APP1063091, 1008271 and 1019887 from Australia’s National Health and Medical Research Council of Australia (NHMRC). The coordinating team: National Institutes of Health grants R21 DA033827 and R01 DA026911 from NIDA.
Other acknowledgments:
We wish to thank the following for making the participation of each group possible: ALSPAC: All the families who took part in this study, the midwives for their help in recruiting them, and the whole ALSPAC team, which includes interviewers, computer and laboratory technicians, clerical workers, research scientists, volunteers, managers, receptionists and nurses. ATP: All collaborators who have contributed to the Australian Temperament Project, especially Professors Ann Sanson, Margot Prior, Frank Oberklaid, John Toumbourou and Ms Diana Smart. We would also like to sincerely thank the participating families for their time and invaluable contribution to the study. The ATP study is located at The Royal Children’s Hospital Melbourne (Australia) and is a collaboration between Deakin University, The University of Melbourne, the Australian Institute of Family Studies, The University of New South Wales, The University of Otago (New Zealand) and the Royal Children's Hospital; further information available at www.aifs.gov.au/atp. The views expressed in this paper are those of the authors and may not reflect those of their organisational affiliations, nor of other collaborating individuals or organisations. CHDS: Allison L Miller, laboratory team leader, Carney Centre for Pharmacogenomics. CoFaMS: We would like to thank study participants for their time and willingness to take part in the study. COGA: The Collaborative Study on the Genetics of Alcoholism (COGA), Principal Investigators B Porjesz, V Hesselbrock, H Edenberg, L Bierut, includes 11 different centres: University of Connecticut (V Hesselbrock); Indiana University (HJ Edenberg, J Nurnberger Jr, T Foroud); University of Iowa (S Kuperman, J. Kramer); SUNY Downstate (B. Porjesz); Washington University in St Louis (L Bierut, J Rice, K Bucholz, A Agrawal); University of California at San Diego (M Schuckit); Rutgers University (J Tischfield, A Brooks); University of Texas Rio Grand Valley (L Almasy), Virginia Commonwealth University (D Dick), Icahn School of Medicine at Mount Sinai (A Goate) and Howard University (R Taylor). Other COGA collaborators include: L Bauer (University of Connecticut); J McClintick, L Wetherill, X Xuei, Y Liu, D Lai, S O’Connor, M Plawecki, S Lourens (Indiana University); G Chan (University of Iowa; University of Connecticut); J Meyers, D Chorlian, C Kamarajan, A Pandey, J Zhang (SUNY Downstate); J-C Wang, M Kapoor, S Bertelsen (Icahn School of Medicine at Mount Sinai); A Anokhin, V McCutcheon, S Saccone (Washington University); J Salvatore, F Aliev, B Cho (Virginia Commonwealth University); and Mark Kos (University of Texas Rio Grand Valley). A Parsian and M Reilly are the NIAAA Staff Collaborators. We thank John Budde, Maribel Martinez and Oliver Reyes from Alison Goate's lab for their technical assistance in genotyping. COGEND: COGEND is a collaborative research group and a part of the NIDA Genetics Consortium. Lead investigators who directed data collection include Laura Bierut, Naomi Breslau, Dorothy Hatsukami and Eric Johnson. The authors thank Heidi Kromrei and Tracey Richmond for their assistance in data collection. We thank John Budde, Maribel Martinez and Oliver Reyes from Alison Goate's lab for their technical assistance in genotyping. DeCC: All individuals who participated in the DeCC and BaCCs studies and were essential for their successful completion. Specifically, we acknowledge the leadership of Peter McGuffin and Anne Farmer, and the contribution of the following: Cerisse Gunasinghe, Katharine Mead, Joanna Gray, Ylva Dahlin, Amanda Elkin, Audrey Morgan, Joanna O'Leary, Nathan O'Neill, Nicola Reynolds, Zainab Samaan, Abraham Stern, Linda Southwick, Kopal Tandon, Alison Wheatley, Richard Williamson, Debra Woolway, Julia Woods, Sarah Ball, Ophelia Beer, Julian Childs, Sam Keating, Rachel Marsh, Penny Machin and Lucy Maddox, who were involved with the data collection and management of the studies. EPIC-Norfolk: All participants, general practitioners and the EPIC-Norfolk study team for their contribution to the research program. G1219: The families of the G1219 study for their time, Robert Plomin, Ian Craig, Pak Sham, Karen Sugden, Alice Gregory and Alejandro Crosico, Matthew Nash, Abram Sterne, Richard Williamson and Maria Napolitano for their contributions to the project. GAN12-France: Marion Leboyer, Chantal Henry, Stéphane Jamain, Jean-Pierre Kahn, Sébastien Gard. The GAN12-France study includes three centres : Creteil (M Leboyer, C Henry), Bordeaux (S Gard) and Nancy (JP Kahn) and also the INSERM U955 Translational Psychiatry research team (S Jamain). GENESIS: Catherine Genty and Aurélie Cazals for their assistance in conducting the study. MARS: The participants and their parents for their engagement in the study. We thank Erika Hohm and Katrin Zohsel for their assistance in conducting the study. MLS: Robert Zucker for allowing access to the MLS data. NEWMOOD: Diana Chase, Emma J Thomas, Darragh Downey, Dorottya Pap, Judit Lazary, Zoltan G Toth for their assistance in the recruitment and data acquisition and Hazel Platt for her assistance in genotyping. POUCH: Participants in the POUCH Study and the Project Director, Bertha Bullen and Nicole Jones. QIMRtwin: The staff from the Queensland Institute of Medical Research who were involved in the genotyping, the twins (drawn from the Australian NH&MRC Twin Registry) for their cooperation and Professors Ian Hickie, Grant Montgomery, and Naomi Wray for their encouragement. SALVe 2001: Professor Lars Oreland, Professor Jerzy Leppert, Professor Leif Lindström, Professor John Öhrvik, associate professor Rickard L Sjöberg and PhD Per-Olof Alm. SEBAS: The staff at the Surveillance and Research Division (Health Promotion Administration at the Ministry of Health and Welfare in Taiwan) who were instrumental in the design and implementation of the SEBAS and supervised all aspects of the fieldwork and data processing. SHIP/TREND: The participants taking part in the SHIP and TREND study. The contribution to data collection performed by study nurses, study physicians, interviewers and laboratory workers is gratefully acknowledged. We are also appreciative of the important support of IT and computer scientists, health information managers and administration staff. TRAILS: Everyone who participated in this research and made it possible. U Bologna: Dr Raffaella Calati for contribution in paper writing, Dr Elena Marino and Dr Adele Pirovano for their contribution to genetic analyses, and Professor Cristina Colombo from the Scientific institute San Raffaele of Milan, Department of Psychiatry, for her precious supervision of the study. U Molise: Dr Vladimir Carli (Karolinska Institutet, Stockholm, Sweden), Dr Leonardo Zaninotto and Professor Alessandro Serretti (University of Bologna, Italy), Dr Laura Recchia (University of Molise, Campobasso, Italy), Dr Valentina Gatta and Dr Liborio Stoppia (G d'Annunzio University, Chieti-Pescara, Italy) for their valuable contribution to the study. The coordinating team: Sherri Fisher, James Childress, Brandie Thurman for administrative support and Katharina Domschke for contributions to the design of the analysis plan.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
IM Anderson has received consultancy/speaking fees from Servier, Alkermes, Lundbeck/Otsuka, Takeda and Janssen, and grant support from Servier and AstraZeneca. V Arolt declares that over the last 3 years he has received compensations for his contributions as member of advisory boards and for presentations for the following companies: AstraZeneca, Eli Lilly, Janssen-Organon, Lundbeck, Otsuka, Servier and Trommsdorff. These co-operations have no relevance to the work that is covered in the manuscript. T Banaschewski served in an advisory or consultancy role for Hexal Pharma, Lilly, Medice, Novartis, Otsuka, Oxford outcomes, PCM scientific, Shire and Viforpharma. He received conference attendance support and conference support or received speaker’s fees from Lilly, Medice, Novartis and Shire. He is/has been involved in clinical trials conducted by Lilly, Shire & Viforpharma. F Bellivier has received honoraria or research or educational conference grants from Bristol-Myers Squibb, Otsuka, Eli Lilly, Servier, Takeda, Sanofi Aventis, Lundbeck, AstraZeneca, the European Space Agency and has received peer review research funding from French Ministry of research, Assistance Publique – Hôpitaux de Paris, the National Institute for Research (INSERM) and the NARSAD. LJ Bierut, AM Goate and JC Wang are listed as inventors on Issued Us Patent 8 080 371, 'Markers for Addiction' covering the use of certain SNPs in determining the diagnosis, prognosis and treatment of addiction. NL Saccone is the spouse of SF Saccone, who is also listed as an inventor on the patent. JFW Deakin variously performed consultancy, speaking engagements and research for Bristol-Myers Squibb, AstraZeneca, Eli Lilly, Schering Plough, Janssen-Cilag and Servier (all fees are paid to the University of Manchester to reimburse them for the time taken); he has share options in P1vital. JI Nurnberger is an investigator for Assurex and Janssen and a consultant for Janssen. E Olié has received a research grant from AstraZeneca and speaking fees from Otsuka, AstraZeneca, Lundbeck. She is involved in clinical trail conducted by Janssen. The remaining authors declare no potential conflicts of interest: TM Air, KJ Anstey, R Araya, C Åslund, G Bagdy, BT Baune, F Bellivier, DI Boomsma, L Bowes, M Burmeister, R Burns, EM Byrne, C Coffey, S Cohen-Woods, P Courtet, WF Coventry, RC Culverhouse, U Dannlowski, EJC de Geus, T Eley, S Easteal, B Etain, DM Fergusson, HL Fisher, K Gawronski, D Glei, N Goldman, X Gonda, HJ Grabe, S Guillaume, A Hatzimanolis, C Holzman, AC Horton, J Horwood, JJ Hottenga, I Jaussent, C Jawahar, C Jennen-Steinmetz, EO Johnson, G Juhasz, M Kennedy, JR Kramer, M Lajnef, M Laucht, KJ Lester, K Little, Y Ma, L Mandelli, NG Martin, CM Middeldorp, M Munafò, M Nauck, E Nederhof, KW Nilsson, AJ Oldehinkel, C Olsson, J Ormel, C Otte, GC Patton, BWJH Penninx, P Petschner, WJ Peyrot, K Ritchie, M Sarchiapone, JM Scheid, C Schwahn, A Serretti, G Sinnamon, JH Smit, D Stacey, NC Stefanis, PG Surtees, Y Tian, C Toben, S Van der Auwera, S Villafuerte, H Völzke, N Wainwright, M Weinstein, M Whooley, G Willemsen, HM zu Schwabedissen.
Additional information
Supplementary Information accompanies the paper on the Molecular Psychiatry website
Supplementary information
PowerPoint slides
Rights and permissions
About this article
Cite this article
Culverhouse, R., Saccone, N., Horton, A. et al. Collaborative meta-analysis finds no evidence of a strong interaction between stress and 5-HTTLPR genotype contributing to the development of depression. Mol Psychiatry 23, 133–142 (2018). https://doi.org/10.1038/mp.2017.44
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mp.2017.44
This article is cited by
-
Methodological concerns in umbrella review of serotonin and depression
Molecular Psychiatry (2024)
-
Difficult lives explain depression better than broken brains
Molecular Psychiatry (2024)
-
CYP2C19-rs4986893 confers risk to major depressive disorder and bipolar disorder in the Han Chinese population whereas ABCB1-rs1045642 acts as a protective factor
BMC Psychiatry (2023)
-
Associations of the serotonin transporter gene polymorphism, 5-HTTLPR, and adverse life events with late life depression in the elderly Lithuanian population
Scientific Reports (2023)
-
The serotonin theory of depression: a systematic umbrella review of the evidence
Molecular Psychiatry (2023)