Abstract
Genome-wide association studies (GWASs) have identified several susceptibility loci for bipolar disorder (BD) and shown that the genetic architecture of BD can be explained by polygenicity, with numerous variants contributing to BD. In the present GWAS (Phase I/II), which included 2964 BD and 61 887 control subjects from the Japanese population, we detected a novel susceptibility locus at 11q12.2 (rs28456, P=6.4 × 10−9), a region known to contain regulatory genes for plasma lipid levels (FADS1/2/3). A subsequent meta-analysis of Phase I/II and the Psychiatric GWAS Consortium for BD (PGC-BD) identified another novel BD gene, NFIX (Pbest=5.8 × 10−10), and supported three regions previously implicated in BD susceptibility: MAD1L1 (Pbest=1.9 × 10−9), TRANK1 (Pbest=2.1 × 10−9) and ODZ4 (Pbest=3.3 × 10−9). Polygenicity of BD within Japanese and trans-European-Japanese populations was assessed with risk profile score analysis. We detected higher scores in BD cases both within (Phase I/II) and across populations (Phase I/II and PGC-BD). These were defined by (1) Phase II as discovery and Phase I as target, or vice versa (for ‘within Japanese comparisons’, Pbest~10−29, R2~2%), and (2) European PGC-BD as discovery and Japanese BD (Phase I/II) as target (for ‘trans-European-Japanese comparison,’ Pbest~10−13, R2~0.27%). This ‘trans population’ effect was supported by estimation of the genetic correlation using the effect size based on each population (liability estimates~0.7). These results indicate that (1) two novel and three previously implicated loci are significantly associated with BD and that (2) BD ‘risk’ effect are shared between Japanese and European populations.
Similar content being viewed by others
Introduction
Bipolar disorder (BD) is a common psychiatric disorder characterized by mood swings between positive manic/hypomanic and negative/depressive states, with a lifetime prevalence of >1%.1 Family, twin and adoption studies have yielded heritability estimates of ~80%.2
Recent European ancestry-based genome-wide association studies (GWASs), including a meta-analysis of GWASs by the Psychiatric GWAS Consortium Bipolar Disorder Working Group (PGC-BD), have highlighted a number of susceptibility loci for BD.3, 4, 5, 6, 7 However, many BD loci may be unidentified because their effect sizes are so small that most studies to date are not adequately powered to detect them individually.2
Based on these results, BD can be considered a polygenic disorder with susceptibility, which is the result of the accumulative genetic effect of numerous variants with small effect size. Schizophrenia has been studied with great success using polygenic models such as risk profile score (RPS) analysis.8, 9, 10 RPS analysis has also shown that BD risk can be explained with polygenicity.11 In this model, trans population analysis is important because it can provide more information about the shared genetic ‘risk’ that exists across populations than can analyses that compare the results only according to the same ancestry. However, studies of BD subjects with Asian ancestry have tended to be smaller in size, affording less power to detect small effects12, 13 and making it difficult to examine the trans population effects of the BD ‘risk’ variants.
In this study, we conducted a GWAS of BD in the Japanese population and meta-analysis using data acquired from the PGC-BD with the aim of identifying novel BD susceptibility genes. Further, we aimed to assess whether there was evidence for polygenicity of BD (that is, to assess whether there was evidence that BD susceptibility is influenced by a large number of single nucleotide polymorphisms (SNPs) distributed across the genome) by examining within the Japanese samples and between subjects with Japanese and European ancestries (as the trans population analysis).
Materials and methods
Subjects
We conducted two GWASs for BD within the Japanese population because of the different sample collection periods and the use of different DNA chips (Supplementary Table 1). These were subsequently meta-analyzed.
BD subjects
The Phase I GWAS (Illumina HumanOmniExpressExome v1.0 chip) initially included 1612 BD subjects, with 1545 BD cases remaining after genotype quality control (QC); and the Phase II GWAS (Illumina HumanOmniExpressExome v1.2 chip) initially included 1604 BD subjects, with 1419 BD cases remaining after QC (Supplementary Table 1). The diagnosis for each subject followed the DSM-IV-TR criteria for BD and schizoaffective disorder and was reached by the consensus of at least two experienced psychiatrists, based on unstructured interviews with the subject and their family, as well as a review of the subject's medical records. Subjects were excluded if they had also been diagnosed with an intellectual disability.
Non-psychiatric controls
As controls, we used GWAS data for subjects in the BioBank Japan project. For the Phase I study, these comprised 7408 subjects who were genotyped (Illumina HumanOmniExpress v1 chip) in a previous GWAS as case subjects for five non-psychiatric disorders (cerebral aneurysm, esophageal cancer, endometrial cancer, chronic obstructive pulmonary disease and glaucoma) or as healthy volunteers (Supplementary Table 1). The controls for the Phase II study included 54 479 subjects who had also been genotyped (Illumina HumanOmniExpressExome v1.2 chip) as case subjects for 14 non-psychiatric disorders: nephrotic syndrome, cancers (stomach, lung, colorectal, prostate and breast), glaucoma, chronic periodontitis, type 2 diabetes (T2D), dyslipidemia, arrhythmia, cerebral infarction, epilepsy and nephrolithiasis, or as healthy controls (Supplementary Table 1). The controls were not psychiatrically evaluated.
Genotyping, QC and imputation
We genotyped the BD subjects using the Illumina HumanOmniExpressExome v.1.0/v.1.2 BeadChips (Supplementary Table 1). Detailed QC, including population stratification (Supplementary Figure 1), is presented in the Supplementary Texts. We then performed genotype imputation using a subset of the 1000 Genomes Project Phase I dataset as a reference, comprising the Japanese in Tokyo (JPT), Han Chinese in Beijing (CHB), and Southern Han Chinese (CHS) populations14 (Supplementary Texts). Cases and controls for each phase were imputed in the same run. After imputation, we included only SNPs with an imputation quality score R2⩾0.3 and minor allele frequency >1%.
Statistical analysis
GWA analysis and meta-analysis
For the SNP-based association analysis, logistic regression with co-variation of the first two eigenvectors was applied and the combined P values from GWAS Phases I and II and/or the PGC-BD data were calculated using a fixed-effect model using an inverse-variance method (Supplementary Texts). Regional association plots were generated using LocusZoom.15 The significance level was set at 5 × 10−8 (two-sided).
RPS analysis
For the RPS analysis,8, 9, 10 we used the statistical analysis software package PRSice v1.23.16 The P threshold (PT) for selecting the ‘risk’ SNPs was set sequentially at 0.1, 0.2, 0.3, 0.4 and 0.5; SNPs were selected if their P values were between 0 and the chosen value of PT. As including SNPs in the MHC region could inflate the score because of high linkage disequilibrium (LD), we removed SNPs in this region by setting the ‘remove.mhc’ flag in the software. The eligible SNPs for RPS were then selected based on LD clumping (used by the default setting of the software). The variance explained for the RPS was estimated using Nagelkerke’s R2 from a logistic regression model.
To calculate the RPS within the Japanese samples, we set Phase II (with a larger sample size) as the discovery and Phase I as the target (Phase II/Phase I: discovery/target) and analyzed this set, and repeated this with the discovery and target reversed (that is, Phase I/Phase II: discovery/target). Next, to evaluate the trans population effect of the ‘risk’ SNPs from the PGC-BD (subjects of European ancestry), we set the PGC-BD as discovery and (Phase I+II) as the target (PGC-BD/Phase I+II: discovery/target).
There may have been a possible bias in this analysis because our control subjects comprised people with a number of diseases, as described earlier. To check for any heterogeneity caused by this, and to use these results as a negative control, we performed leave-one-disease-out analysis for each disease. For example, a proportion of the Phase I control subjects had cerebral aneurysm, and so we performed RPS for target samples excluding these cerebral aneurysm subjects from the ‘control’ and by defining cerebral aneurysm as the ‘case’ for the negative control as follows:

Each of the 21 disease groups, including two sets of healthy volunteers, was removed in turn from the target ‘control’ samples, and we calculated the polygenic score defined by PGC-BD for each non-psychiatric control.
The significance level for the RPS analysis was set at 0.001, a conservative threshold described by Euesden et al.16
LD score regression
We used LD score regression to examine SNP heritability17 for the meta-analysis (Phases I and II: imputation R2⩾0.3) and/or PGC-BD (imputation INFO⩾0.3), and to assess whether there was population stratification.18 We used pre-computed LD scores for East Asian (for Japanese results) and European (for PGC results) populations based on 1000 Genome Project data listed on the website (https://data.broadinstitute.org/alkesgroup/LDSCORE/eas_ldscores.tar.bz2). We filtered the SNPs to HapMap3 SNPs (‘—merge-alleles’ flags), and set the population prevalence of BD at 0.01 (‘—pop-prev 0.01’ flag) and the sample prevalence at appropriate values (=‘number of cases’/‘number of total subjects’) in each dataset (‘—samp-prev’ flag).
Although LD score regression can assess genetic correlation (if samples from the same population are targeted), it is difficult to calculate correlations between trans population datasets (such as between the Japanese and European populations). In the comparison between Japanese and European BD results, therefore, we used a recently developed program, Popcorn (version 0.9.6), which is based on a concept similar to LD score regression,19 and examined (1) the trans population genetic effect correlation (the correlation coefficient for the per-allele SNP effect sizes, ρge), and (2) the genetic impact correlation (the correlation coefficient for the population-specific allele variance normalized SNP effect sizes, ρgi).19 For the calculation of the trans-population score, we used HapMap3 datasets, representing the European population by the CEU dataset and the Asian population by CHB+JPT datasets. Genetic correlations were then calculated by the GWAS SNPs of the combined GWAS of the Japanese datasets (a meta-analysis of Phases I and II: imputation R2⩾0.3) and PGC-BD (imputation INFO⩾0.3) with the default setting, which removed SNPs below the minor allele frequency (MAF) cutoff of 5% and those with A/T or G/C alleles.
Results
Combined GWAS of the two Japanese datasets (Phases I and II)
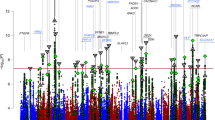
A total of 6 195 093 imputed SNPs in 2964 BD and 61 887 non-psychiatric subjects passed our stringent QC. From these, we generated the Manhattan and quantile-quantile (QQ) plots shown in Figure 1 and Supplementary Figure 2. The genomic inflation factor (λGC) for the meta-analysis appeared at 1.047. In addition, the intercept from the LD score regression analysis was 1.054, indicating that most of the inflation was due to polygenicity rather than bias.18
Manhattan plot of the meta-analyses for (a) the Japanese samples (Phases I and II) and (b) the Japanese samples (Phases I and II) and Psychiatric GWAS Consortium (PGC-BD). Horizontal line indicates threshold for genome-wide significance (P<5 × 10−8). Manhattan plot for (a) the Japanese samples (Phases I and II) and (b) the Japanese samples (Phases I and II) and PGC-BD.
One region reached genome-wide significance (5 × 10−8; Figure 1), namely the association at a SNP on 11q12.2 of rs28456 in the fatty acid desaturase 2 gene (FADS2) (P=6.4 × 10−9, odds ratio (OR)=1.18; Table 1). This had hits in multiple genes, including FADS1/2/3 (Table 1 and Supplementary Table 2). It is of interest that (1) this region is a known locus for the regulation of plasma blood lipid traits20 and n-3/n-6 polyunsaturated fatty acids21, 22, 23 and (2) the top SNP (rs28456) has an expression quantitative trait locus (eQTL) effect on FADS1 in the brain (cerebellum) samples (P=7.8 × 10−10, beta=−0.62 as the effect of the minor allele ‘G (risk for BD)’ relative to major allele ‘A’) according to the Genotype-Tissue Expression (GTEx) database (http://www.gtexportal.org/home/, Data Source: GTEx Analysis Release V6p (dbGaP Accession phs000424.v6.p1). In addition, other datasets (BRAINEAC: http://www.braineac.org/) supported the eQTL effect in the brain samples (FADS2 in temporal cortex, Pbest=1.7 × 10−5; FADS1 in cerebral cortex, Pbest=1.3 × 10−5).
The non-psychiatric controls of Phase II included phenotypes related to lipid and/or glucose metabolism (N=25 228), such as those for dyslipidemia and T2D. We therefore performed sensitivity analysis to exclude the possibility that this signal was derived from an association with these subjects. In this analysis, we confirmed that the effect size did not change substantially compared with the non-psychiatric controls excluding those with dyslipidemia or T2D (rs28456, OR=1.15), although the P value was attenuated (P=7.4 × 10−7; Supplementary Table 3).
To validate the previous findings, we checked the associations of susceptibility genes with genome-wide significance reported by the PGC-BD3 and other individual GWASs (Supplementary Table 4 and Supplementary Figure 3).4, 6, 7 Although, there were no significant associations (that is, with P<0.05) of these indexed SNPs (except for the SNP in ODZ4 rs12576775, P=0.042) with BD in the Japanese population, the surrounding SNPs showed P values of <0.05, particularly for ODZ4 (known as TENM4, encoding teneurin transmembrane protein 4), TRANK1 (encoding tetratricopeptide repeat and ankyrin repeat containing 1) and DHH (encoding desert hedgehog proteins). Specifically, the SNPs in these three genes showed a strengthened association after merging PGC-BD results (see below).
We calculated the SNP heritability of Phase I (979 751 SNPs), Phase II (959 428 SNPs), and the combined Japanese (Phase I/II, 903 223 SNPs) datasets by the LD score regression method. The SNP heritability on the total liability scale (h2) was 0.263 (s.e.=0.0642), 0.129 (s.e.=0.0530) and 0.148 (s.e.=0.0288) for Phase I, Phase II and the combined Phase I/II, respectively. The discrepancy of the h2 between Phases I and II might be due to the smaller size in Phase I (1545 BD vs 7408 controls), therefore the h2~0.15 was the best estimate for the BD in the Japanese population. Whereas, the h2 for the PGC-BD was estimated around 0.25 (1 059 316 SNPs, h2=0.245, s.e.=0.0204), that was larger than that of Japanese sample. Based solely on the current results, we could not interpret the difference of the h2 values between the Japanese samples and the PGC-BD; further replication with larger sample size will be required.
Meta-analysis of the Japanese (Phases I and II) and PGC-BD results
To maximize the sample size, we conducted a meta-analysis combining the Phases I, II and the PGC-BD datasets,3 which produced a combined sample size of 10 445 cases and 71 137 controls. Manhattan and QQ plots are shown in Figure 1 and Supplementary Figure 2, respectively. Again, we found a significant association with 11q.12.2 (rs174576, P=1.34 × 10−10, OR=1.13; Figure 2, Table 2, and Supplementary Table 5). When we set ‘control’ samples excluding dyslipidemia/T2D subjects, rs174576 still reached genome-wide significance (P=4.9 × 10−9; OR=1.12, 95% confidence interval: 1.08–1.16), indicating that any possible bias derived from samples with dyslipidemia/T2D did not influence the significance. Notably, rs174576 has eQTL effect in GTEx dataset (FADS1 in cerebrellum, P=2.3 × 10−9; beta=−0.59 as the effect of the minor allele ‘A (risk for BD)’ relative to major allele ‘C’) and BRAINEAC dataset (FADS1 in cerebral cortex, Pbest=2.5 × 10−6; FADS2 in temporal cortex, Pbest=1.2 × 10−5).
Regional plots of the top hit in the association results based on the meta-analysis of Japanese (Phases I and II) and the PGC-BD results. Blue lines indicate the recombination rate for the ASN population in the 1000 Genome Project. The Y axis is −log10(P-values) of the SNPs and the X axis is chromosomal position (hg19). The linkage disequilibrium (r2) between the top and the remaining SNPs is indicated by color. (a) FADS gene cluster, (b) NFIX, (c) MAD1L1, (d) TRANK1, (e) ODZ4 (known as TENM4), (f) MLL2~DHH. PGC-BD, Psychiatric GWAS Consortium for BD; SNPs, single nucleotide polymorphism.
In this analysis, we found four additional regions that showed significant associations at the genome-wide level, one of which, NFIX (encoding nuclear family I/X, rs4926298, P=5.8 × 10−10) is another novel BD susceptibility gene (Figure 2, Table 2 and Supplementary Table 5).
ODZ4, MAD1L1 (encoding MAD1 mitotic arrest deficient-like 1) and TRANK1, are known BD/psychosis susceptibility genes; these enhanced the association of this meta-analysis of the combined Japanese datasets and the PGC-BD dataset (Figure 2, Table 2 Supplementary Table 5), supporting previous evidence. It is of note that the P value for MLL2 (encoding lysine (K)-specific methyltransferase 2D, located near DHH) did not quite reach that for genome-wide significance, but was just below this level (rs10875914, P=6.3 × 10−8; Figure 2, Table 2 and Supplementary Table 5).
Trans population effect of SNPs on BD
To evaluate any trans population effect, we first checked the SNP-wise correlation of the effect size between the Phase I/II and the PGC-BD datasets for the top hits based on the final meta-analysis (Phase I/II and PGC-BD). As expected, we found a good correlation between the effect sizes of the two datasets (Supplementary Figure 4A). However, the top SNPs selected based on the combined Japanese datasets only (Supplementary Figure 4B) or the PGC-BD dataset only (Supplementary Figure 4C) did not show strong correlations, particularly the SNPs selected from the combined Japanese datasets. Again, we speculate that this may be due to the lower sample size for the Japanese samples resulting in less statistical power.
To analyze the polygenic architecture represented by numerous ‘risk’ SNPs, we calculated the RPSs defined within the Japanese sample as follows. ‘risk’ SNPs were defined according to the association results for Phase II (the dataset with the larger sample size), and we examined whether cases with BD had a higher RPS than controls in Phase I (Phase II/Phase I: discovery/target pair); we then repeated this with the dataset swapped (i.e., Phase I/Phase II: discovery/target pair). In these analyses of the Japanese case-control samples, we found a significant higher RPS in the case (P values 10−20–10−29), explaining ~0.7%–2.3% of the variance (Figure 3, Supplementary Figures 5, 6, and Supplementary Table 6). We then performed a trans population analysis between the European and the Japanese populations. A larger sample size improves the discovery results defining the ‘risk’ SNPs, because more accurate results may be obtained; we therefore set the PGC-BD as discovery and Phase I+II as target. In this comparison, we found a significant higher RPS defined by the European BD in the target Japanese BD samples (P values: 10−12–10−13). However, the variance explained (~0.27%) was smaller than that for the Japanese pair (Figure 3, Supplementary Figures 5, 6, and Supplementary Table 6).
Risk profile score (RPS) analysis. Y axis indicates the variance-explained (Nagaelkerke’s R2). Three discovery/target sets were included: (a) Discovery: Phase II, Target: Phase I, (b) Discovery: Phase I, Target: Phase II, and (c) Discovery: PGC-BD, Target: Phase I+II. *P<10−10, **P<10−20.
To determine whether such results were detected only in BD patients, we performed leave-one-disease-out analyses, removing each disease group from the ‘control’ and treating them as ‘cases.’ These results indicated that most of the profile scores predicted almost no variance (~10−5) in BD risk. The highest variance explained was found in the analysis for nephrotic syndrome (Phase II, N=677: best R2=0.00087) (Supplementary Figure 6 and Supplementary Table 7); however, this result was probably due to the small sample size of subjects with nephrotic syndrome (N=677), as the P value for nephrotic syndrome was 0.0128. We therefore concluded there were no differences in RPSs based on the BD ‘risk’ SNPs between the disease groups making up the controls (Supplementary Figure 6 and Supplementary Table 7). This implies that (1) our controls were minimally biased by sample heterogeneity associated with a polygenic effect, and (2) BD risk based on PGC-BD was shared only by subjects with BD (but not other diseases) in the Japanese population.
Lastly, we estimated trans population genetic correlations using the effect sizes of the SNPs.19 In this analysis of 733 574 SNPs, we detected a significant trans population genetic effect correlation between our Japanese samples and PGC-BD (ρge: liability=0.724, P=5.40 × 10−3) as well as a population genetic impact correlation that accounted for the allele frequencies of the SNPs (ρgi: liability=0.684, P=9.17 × 10−4). Notably, these liabilities were higher than those for rheumatoid arthritis (ρge/ρgi=0.46/0.46) and T2D (ρge/ρgi=0.62/0.61) reported in the original method paper.19
Discussion
We identified two novel loci with significant associations with BD: SNPs in the FADS gene cluster and NFIX. Our findings for ODZ4, MAD1L1 and TRANK1 supported the associations detected previously.
Our meta-analysis found a significant association with BD in a region that included genes encoding fatty acid desaturase (FADS) (Figure 2), where robust associations have been established with blood lipid traits20 and with n-3/n-6 polyunsaturated fatty acids (PUFA).21, 22, 23 The LD matrices with the index BD SNP (rs174576) revealed that all QTL SNPs for lipid traits (HDL/LDL/TG/T-Chol, n-3/n-6 PUFAs) around the FADS region were in strong LD with the index SNP (Supplementary Table 8). It suggested that SNPs located around this region are in the LD block, and most QTL SNPs exhibited significant association with BD. Therefore, we speculate that the lipid abnormality may be involved in the pathophysiology of BD. Notably, epidemiological surveys have revealed an increased incidence of hyperlipidemia in BD patients, at ~5.6% higher than that for the general population (OR=1.75),24 and an approximately two-fold greater risk of metabolic syndrome, including dyslipidemia.25, 26 Although the medication, specifically second-generation antipsychotics, for BD symptoms is the definitive risk for dyslipidemia (which was not taken into account in the studies just cited24, 25, 26), the risk of metabolic abnormality may overlap that of BD. This is supported by surveys, albeit of a small sample size, that have shown an increased prevalence of metabolic syndrome in the first-degree relatives of psychosis subjects, including those with BD,27 and in drug-naive patients with BD.28 As yet, there has been no evidence of a causal relationship of lipid abnormality on BD; further epidemiological or genetic studies are therefore warranted to establish such a relationship.
Another novel locus for BD detected in our GWAS is located near NFIX. In the meta-analysis of our GWAS involving the Japanese subjects (Phases I and II), SNPs in NACC1, downstream of NFIX, showed a stronger association. However, when PGC-BD data was also included, the association of NFIX strengthened, reaching genome-wide significance. NFIX is a member of the nuclear factor one (NF1) family and plays critical roles in transcription and replication.29 Nonsense/frameshift mutations and copy number variants of NFIX cause Sotos syndrome 2 (SOTOS2: OMIM *6145753) or Marshall-Smith syndrome (MRSMS: OMIM *602535), which are characterized by intellectual disability. Although there is no evidence for an association between BD and NFIX, further studies are warranted.
The last locus highlighted was MAD1L1, which contributes to cell cycle control through the regulation of mitosis.30 This gene represents a risk locus for psychosis in a combined population of schizophrenic and BD patients.31 Our results indicate that this gene confers a risk for BD, as well as supporting a pleiotropic effect of variants in MAD1L1.
RPS analysis revealed evidence that the risk profile score defined by European-based BD ‘risk’ SNPs was higher in the BD subjects in the Japanese population. Interestingly, the level of variance explained in this trans population analysis was much lower than that for the ‘within Japanese comparisons’. A previous study of schizophrenia analyzed the trans population effect using our Japanese samples and indicated that the contribution of the European-based score allele was one-half or one-third lower in the Japanese samples.32 The sample size in the present study was larger than that of the schizophrenia GWAS (~500 cases vs ~500 controls),32 but a similar reduction in the contribution (comparing the within-Japanese with the trans population samples) was observed in the two sample sets. This difference in the contribution may result from the combined influences of LD differences and/or a population with unique risk. However, it should be emphasized that BD ‘risk’ effect are shared across populations. A different analysis supported this result by estimating the trans-population genetic effect based on correlations of the effect size. In this analysis, we estimated that the genetic correlation coefficient for BD was ~0.7, which was higher than that for rheumatoid arthritis (~0.5) and T2D (~0.6).19 This indicates that there might be a shared genetic basis in BD involving a higher proportion of subjects than for rheumatoid arthritis or T2D.
In addition, the genetic correlations calculated with and without consideration of the allele frequency (ρge/ρgi=0.724/0.684) had almost the same impact, similar to that in previous reports of rheumatoid arthritis (ρge/ρgi=0.46/0.46) and T2D (ρge/ρgi=0.62/0.61).19 This indicates, and supports the previous speculation,19 that differences in allele frequencies of common SNPs (observed in the Japanese and European populations) did not have a large impact on the trans-population phenotypic differences for BD, and probably for complex diseases. We further speculated that the population-specific effect could be assessed from our results. As an explorative analysis, we calculated the trans population genetic effect (ρge) or impact (ρgi) within the Japanese population using the data from Phases I and II. As expected, we found nearly a perfect correlation (~1.0) in both ρge and ρgi. The difference of ρge /ρgi between the Japanese-European and within-Japanese comparisons was ~0.3. The reason for this difference remains to be explained, but we assume (1) environmental factors or gene-environment interaction have a larger impact on BD in the Japanese population, assuming we detected the correct estimates of SNP heritabilities, with h2 in the Japanese population found to be lower than that in the European PGC-BD dataset or (2) the difference may derive from rare variants (low MAF SNPs (1–5%) or single nucleotide variants (SNVs: MAF<1%)) only seen in one population, which were not selected in this analysis (MAF⩾5%) because MAF in common SNPs does not have large impact, as mentioned above.
Our study had several limitations. First, the subjects we used as controls had not been psychiatrically screened. However, this is reasonable because the prevalence of BD is only ~1% (and may be only ~0.2% in Japan).33 Although most of the control subjects had various non-psychiatric disorders, there were no clear associations between BD and these disorders. We assessed the influence of these diseases on the significant loci using analyses where one disease (non-psychiatric control) was excluded from the controls and compared with BD for the significant SNPs, and confirmed there was little possibility of bias (Supplementary Table 9). We further confirmed the minimal effect of the bias in the leave-one-disease-out RPS analysis (Supplementary Table 7). Second, our samples contained a higher proportion of subjects with BD type II than reported in other studies. The prevalence of BD type II varies across countries, and the proportions of BD I and BD II in our sample simply reflect the lifetime prevalence in Japan (0.1% each).33 There were no significant differences in allele frequencies between BD types I and II for all significant SNPs (that is, SNPs with Pmeta<5 × 10−8). Furthermore, the effect directions were the same for all significant SNPs (Supplementary Table 10). Thus, the proportions of BD types I and II appear to have had minimal influence on our findings. Third, the genotyping of case and control subjects had been performed on different platforms and at different times (Supplementary Table 1). To assess the influence of the batch effect, we checked the concordance of the genotyping determined by these chips (HumanOmniExpress v1, HumanOmniExpressExome v1.0/1.2); the concordance rates (concordance of genotypes between same individuals (three pairs between HumanOmniExpress v1 and HumanOmniExpressExome v1.0, four pairs between HumanOmniExpress v1 and HumanOmniExpressExome v1.2, and 23 pairs between HumanOmniExpressExome v1.0/1.2) on two platforms) in all comparisons were >99.99%, suggesting minimal batch effects in our results.
In conclusion, we identified two novel susceptibility loci (FADS and NFIX) and confirmed loci identified in previous GWASs (ODZ4, MAD1L1, and TRANK1) as susceptibility genes or regions associated with BD. Based on the trans population analysis, we demonstrated a substantial effect of BD ‘risk’ genetic variants across populations. Further analysis will illuminate the shared risk among populations and the population-unique risk.
References
Craddock N, Sklar P . Genetics of bipolar disorder: successful start to a long journey. Trend Genet 2009; 25: 99–105.
Craddock N, Sklar P . Genetics of bipolar disorder. Lancet 2013; 381: 1654–1662.
Psychiatric GWAS Consortium Bipolar Disorder Working Group. Large-scale genome-wide association analysis of bipolar disorder identifies a new susceptibility locus near ODZ4. Nat Genet 2011; 43: 977–983.
Cichon S, Muhleisen TW, Degenhardt FA, Mattheisen M, Miro X, Strohmaier J et al. Genome-wide association study identifies genetic variation in neurocan as a susceptibility factor for bipolar disorder. Am J Hum Genet 2011; 88: 372–381.
Chen DT, Jiang X, Akula N, Shugart YY, Wendland JR, Steele CJ et al. Genome-wide association study meta-analysis of European and Asian-ancestry samples identifies three novel loci associated with bipolar disorder. Mol Psychiatry 2013; 18: 195–205.
Green EK, Hamshere M, Forty L, Gordon-Smith K, Fraser C, Russell E et al. Replication of bipolar disorder susceptibility alleles and identification of two novel genome-wide significant associations in a new bipolar disorder case-control sample. Mol Psychiatry 2013; 18: 1302–1307.
Muhleisen TW, Leber M, Schulze TG, Strohmaier J, Degenhardt F, Treutlein J et al. Genome-wide association study reveals two new risk loci for bipolar disorder. Nat Commun 2014; 5: 3339.
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 2014; 511: 421–427.
Ripke S, O'Dushlaine C, Chambert K, Moran JL, Kahler AK, Akterin S et al. Genome-wide association analysis identifies 13 new risk loci for schizophrenia. Nat Genet 2013; 45: 1150–1159.
International Schizophrenia Consortium, Purcell SM, Wray NR, Stone JL, Visscher PM, O’Donovan MC et al. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 2009; 460: 748–752.
Cross-Disorder Group of the Psychiatric Genomics Consortium, Lee SH, Ripke S, Neale BM, Faraone SV, Purcell SM et al. Genetic relationship between five psychiatric disorders estimated from genome-wide SNPs. Nat Genet 2013; 45: 984–994.
Hattori E, Toyota T, Ishitsuka Y, Iwayama Y, Yamada K, Ujike H et al. Preliminary genome-wide association study of bipolar disorder in the Japanese population. Am J Med Genet 2009; 150B: 1110–1117.
Lee MT, Chen CH, Lee CS, Chen CC, Chong MY, Ouyang WC et al. Genome-wide association study of bipolar I disorder in the Han Chinese population. Mol Psychiatry 2011; 16: 548–556.
1000 Genomes Project Consortium, Abecasis GR, Altshuler D, Auton A, Brooks LD, Durbin RM et al. A map of human genome variation from population-scale sequencing. Nature 2010; 467: 1061–1073.
Pruim RJ, Welch RP, Sanna S, Teslovich TM, Chines PS, Gliedt TP et al. LocusZoom: regional visualization of genome-wide association scan results. Bioinformatics 2010; 26: 2336–2337.
Euesden J, Lewis CM, O’Reilly PF . PRSice: Polygenic Risk Score software. Bioinformatics 2015; 31: 1466–1468.
Finucane HK, Bulik-Sullivan B, Gusev A, Trynka G, Reshef Y, Loh PR et al. Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat Genet 2015; 47: 1228–1235.
Bulik-Sullivan BK, Loh PR, Finucane HK, Ripke S, Yang J et alSchizophrenia Working Group of the Psychiatric Genomics Consortium. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat Genet 2015; 47: 291–295.
Brown BC Asian Genetic Epidemiology Network Type 2 Diabetes Consortium, Ye CJ, Price AL, Zaitlen N . Transethnic Genetic-Correlation Estimates from Summary Statistics. Am J Hum Genet 2016; 99: 76–88.
Global Lipids Genetics Consortium, Willer CJ, Schmidt EM, Sengupta S, Peloso GM, Gustafsson S et al. Discovery and refinement of loci associated with lipid levels. Nat Genet 2013; 45: 1274–1283.
Guan W, Steffen BT, Lemaitre RN, Wu JH, Tanaka T, Manichaikul A et al. Genome-wide association study of plasma N6 polyunsaturated fatty acids within the cohorts for heart and aging research in genomic epidemiology consortium. Circ Cardiovasc Genet 2014; 7: 321–331.
Lemaitre RN, Tanaka T, Tang W, Manichaikul A, Foy M, Kabagambe EK et al. Genetic loci associated with plasma phospholipid n-3 fatty acids: a meta-analysis of genome-wide association studies from the CHARGE Consortium. PLoS Genet 2011; 7: e1002193.
Dorajoo R, Sun Y, Han Y, Ke T, Burger A, Chang X et al. A genome-wide association study of n-3 and n-6 plasma fatty acids in a Singaporean Chinese population. Genes Nutr 2015; 10: 53.
Hsu JH, Chien IC, Lin CH . Increased risk of hyperlipidemia in patients with bipolar disorder: a population-based study. Gen Hosp Psychiatry 2015; 37: 294–298.
McIntyre RS, Danilewitz M, Liauw SS, Kemp DE, Nguyen HT, Kahn LS et al. Bipolar disorder and metabolic syndrome: an international perspective. J Affect Disord 2010; 126: 366–387.
Bai YM, Su TP, Chen MH, Chen TJ, Chang WH . Risk of developing diabetes mellitus and hyperlipidemia among patients with bipolar disorder, major depressive disorder, and schizophrenia: a 10-year nationwide population-based prospective cohort study. J Affect Disord 2013; 150: 57–62.
Mothi SS, Tandon N, Padmanabhan J, Mathew IT, Clementz B, Tamminga C et al. Increased cardiometabolic dysfunction in first-degree relatives of patients with psychotic disorders. Schizophr Res 2015; 165: 103–107.
Guha P, Bhowmick K, Mazumder P, Ghosal M, Chakraborty I, Burman P . Assessment of insulin resistance and metabolic syndrome in drug naive patients of bipolar disorder. Indian J Clin Biochem 2014; 29: 51–56.
Gronostajski RM . Roles of the NFI/CTF gene family in transcription and development. Gene 2000; 249: 31–45.
Avram S, Mernea M, Mihailescu DF, Seiman CD, Seiman DD, Putz MV . Mitotic checkpoint proteins Mad1 and Mad2—structural and functional relationship with implication in genetic diseases. Curr Comput Aided Drug Des 2014; 10: 168–181.
Ruderfer DM, Fanous AH, Ripke S, McQuillin A, Amdur RL et alSchizophrenia Working Group of Psychiatric Genomics Consortium. Polygenic dissection of diagnosis and clinical dimensions of bipolar disorder and schizophrenia. Mol Psychiatry 2014; 19: 1017–1024.
Ikeda M, Aleksic B, Kinoshita Y, Okochi T, Kawashima K, Kushima I et al. Genome-wide association study of schizophrenia in a Japanese population. Biol Psychiatry. 2011; 69: 472–478.
Kawakami N Research of epidemiology on mental health. The final report of the Grant from Ministry of Health, Labour and Welfare, 2006.
Acknowledgements
We thank all the individuals who participated in the study and the contributors of advanced COSMO and Biobank Japan. We also thank Dr Charles Yokoyama at RIKEN Brain Science Institute (BSI) and Dr Makoto Arita at RIKEN Center for Integrative Medical Sciences (IMS) for their critical comments. This work is the result of the Strategic Research Program for Brain Sciences (SRPBS) from Japan Agency for Medical Research and Development (AMED); part of the BioBank Japan Project from the Ministry of Education, Culture, Sports and Technology (MEXT) of Japan; JSPS Kakenhi Grant Numbers JP26117518, JP25293253, JP16H05378, JP26293266 and JP24689046: Grant-in-Aid for Brain/minds from AMED: Grant-in-Aid for Health Labour Sciences Research Grant from the Ministry of Health Labour and Welfare.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Competing interests
During the conduct of the study; Drs Ikeda, Takahashi, Kamatani, Okamoto, Yoshimura, Matsuo, Itokawa, Ozaki and Iwata have received research support from the Ministry of Education, Culture, Sports and Technology (MEXT) of Japan and Japan Agency for Medical Research and Development (AMED); Drs Kusumi, Yoshikawa and Kubo have received research support from MEXT of Japan; Dr Kanba has received research support from AMED and the Ministry of Health Labour and Welfare; Dr Kunugi has received research support from MEXT of Japan, AMED and the Ministry of Health Labour and Welfare; and Dr Kato has received research support from MEXT of Japan, AMED of Japan and Japan Science and Technology Agency. The sponcors had no role in design and conduct of the study; collection, management, analysis and interpretation of the data; and preparation, review or approval of the manuscript. The remaining authors declare no conflicts of interest.
Additional information
The members of for the advanced Collaborative Study of Mood Disorder (COSMO) team are listed before references.
Group Members for the advanced Collaborative Study of Mood Disorder (COSMO) team
The advanced COSMO include authors and co-investigators Masashi Ikeda, MD, PhD, Kenji Kondo, MD, PhD, Takeo Saito, MD, PhD, Kunihiro Kawashima, MD, PhD, Yoshio Yamanouchi, MD, PhD, Tsuyoshi Kitajima, MD, PhD, Ayu Shimasaki, MD, Kohei Kawase, MD, Tomohiro Narita, MD, PhD, Taro Kishi, MD, PhD, Kosei Esaki, MD, PhD, Hiroshi Naitoh, MD, PhD, Nakao Iwata (PI), MD, PhD (Department of Psychiatry, Fujita Health University School of Medicine), Tomoko Toyota, MD, PhD, Kazuo Yamada, MD, PhD, Eiji Hattori, MD, PhD, Yoshimi Iwayama, MS, Takeo Yoshikawa (PI), MD, PhD (Laboratory for Molecular Psychiatry, RIKEN Brain Science Institute), Hiroshi Ujike, MD, PhD, Manabu Takaki, MD, PhD, Shinji Sakamoto, MD, Norihito Yamada, MD, PhD, Yuko Okahisa (PI), MD, PhD (Department of Neuropsychiatry, Okayama University Graduate School of Medicine, Dentistry and Pharmaceutical Sciences), Naoki Hashimoto, MD, PhD, Shin Nakagawa, MD, PhD, Yuki Kako, MD, Teruaki Tanaka, MD, PhD, Yutaka Fujii, MD, PhD, Yuji Kitaichi, MD, PhD, Ichiro Kusumi (PI), MD, PhD (Department of Psychiatry, Hokkaido University Graduate School of Medicine), Takeshi Inoue (PI), MD, PhD (Department of Psychiatry, Tokyo Medical University School of Medicine), Kotaro Hattori, MD, PhD, Noriko Yamamoto, Hiroaki Hori, MD, PhD, Toshiya Teraishi, MD, PhD, Miho Ota, MD, PhD, Hiroshi Kunugi (PI), MD, PhD (Department of Mental Disorder Research, National Institute of Neuroscience, National Center of Neurology and Psychiatry), Branko Aleksic, MD, PhD, Itaru Kushima, MD, PhD, Shohko Kunimoto, PhD, Akira Yoshimi, PhD, Toshiya Inada (co-PI), MD, PhD, Norio Ozaki (PI), MD, PhD (Department of Psychiatry, Nagoya University, Graduate School of Medicine), Katsuaki Suzuki, MD, PhD, Yasuhide Iwata, MD, PhD, Norio Mori (PI), MD, PhD (Department of Psychiatry and Neurology, Hamamatsu University School of Medicine), Yoshio Minabe, MD, PhD (Department of Psychiatry and Neurobiology, Kanazawa University Graduate School of Medical Science), Takahiro A. Kato, MD, PhD, Satomi Katsuki, MD, PhD, Hiroshi Mitsuyasu, MD, PhD, Shigenobu Kanba (PI), MD, PhD (Department of Neuropsychiatry, Kyushu University, Graduate School of Medical Sciences), Hiroaki Kawasaki (PI), MD, PhD (Department of Psychiatry, Fukuoka University, Faculty of Medicine), Tsukasa Sasaki (PI), MD, PhD (Laboratory of Health Education, Graduate School of Education, the University of Tokyo), Mamoru Tochigi, MD, PhD (Department of Psychiatry, Teikyo University School of Medicine), Takeshi Otowa, MD, PhD (Department of NeuroPsychiatry, Graduate School of Medicine, the University of Tokyo), Tohru Ohnuma, MD, PhD, Hajime Baba, MD, PhD, Heii Arai (PI), MD, PhD, (Department of Psychiatry, Juntendo University, School of Medicine), Shusuke Numata, MD, PhD, Jun-ichi Iga, MD, PhD, Tetsuro Ohmori (PI), MD, PhD (Department of Psychiatry, Course of Integrated Brain Sciences, Medical Informatics, Institute of Health Biosciences, The University of Tokushima Graduate School), Motohiro Okada, MD, PhD, Yoshiaki Konishi, MS, Hisashi Tanii (PI), MD, PhD, (Department of Psychiatry, Mie University, Graduate School of Medicine), Shigeto Yamawaki, MD, PhD, Yasumasa Okamoto (PI), MD, PhD, (Department of Psychiatry and Neurosciences, Graduate School of Biomedical and Health Sciences, Hiroshima University), Satoshi Saito, MD, (Department of Psychiatry/ Department of Biological Psychiatry and Neuroscience, Dokkyo Medical University School of Medicine), Kazutaka Shimoda, MD, PhD (Department of Psychiatry, Dokkyo Medical University School of Medicine), Kazufumi Akiyama (PI), MD, PhD (Department of Biological Psychiatry and Neuroscience, Dokkyo Medical University School of Medicine), Hirokazu Fujita, MD, PhD, Shinji Shimodera (PI), MD, PhD (Department of Neuropsychiatry, Kochi Medical school, Kochi University), Masatoshi Takeda, MD, PhD, Ryota Hashimoto (PI), MD, PhD (Molecular Research Center for Children's Mental Development, United Graduate School of Child Development, Osaka University; Department of Psychiatry, Osaka University Graduate School of Medicine), Tadafumi Kato (PI), MD, PhD (Laboratory for Molecular Dynamics of Mental Disorders, RIKEN Brain Science Institute), Tetsufumi, Kanazawa, MD, PhD, Hiroshi Yoneda (PI), MD, PhD (Department of Neuropsychiatry, Osaka Medical College), Yuichiro Watanabe, MD, PhD, Toshiyuki Someya (PI), MD, PhD (Department of Psychiatry, Niigata University Graduate School of Medical and Dental Sciences), Reiji Yoshimura (PI), MD, PhD (Department of Psychiatry, University of Occupational and Environmental Health), Masanari Itokawa (PI), MD, PhD (Center for Medical Cooperation, Tokyo Metropolitan Institute of Medical Science), Masaomi Iyo (PI), MD, PhD (Department of Psychiatry, Chiba University Graduate School of Medicine) and Koji Matsuo (PI), MD, PhD (Division of Neuropsychiatry, Department of Neuroscience, Yamaguchi University Graduate School of Medicine). The contributors did not receive financial compensation.
Supplementary Information accompanies the paper on the Molecular Psychiatry website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Ikeda, M., Takahashi, A., Kamatani, Y. et al. A genome-wide association study identifies two novel susceptibility loci and trans population polygenicity associated with bipolar disorder. Mol Psychiatry 23, 639–647 (2018). https://doi.org/10.1038/mp.2016.259
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mp.2016.259
This article is cited by
-
Incorporating genetic similarity of auxiliary samples into eGene identification under the transfer learning framework
Journal of Translational Medicine (2024)
-
Progress and Implications from Genetic Studies of Bipolar Disorder
Neuroscience Bulletin (2024)
-
Similarity and diversity of genetic architecture for complex traits between East Asian and European populations
BMC Genomics (2023)
-
CEGA: a method for inferring natural selection by comparative population genomic analysis across species
Genome Biology (2023)
-
Methylation in MAD1L1 is associated with the severity of suicide attempt and phenotypes of depression
Clinical Epigenetics (2023)