Abstract
MicroRNAs (miRNAs) represent an important class of small regulatory RNAs that control gene expression posttranscriptionally by targeting mRNAs for degradation or translation inhibition. Early studies have revealed a complex role for miRNAs in major biological processes such as development, differentiation, growth and metabolism. MiR-137 in particular, has been of great interest due to its critical role in brain function and putative involvement in the etiology of both neuropsychiatric disorders and cancer. Several lines of evidence suggest that development, differentiation and maturation of the nervous system is strongly linked to the expression of miR-137 and its regulation of a large number of downstream target genes in various pathways. Dysregulation of this molecule has also been implicated in major mental illnesses through its position in a variant allele highly associated with schizophrenia in the largest mega genome-wide association studies. Interestingly, miR-137 has also been shown to act as a tumor suppressor, with numerous studies finding reduced expression in neoplasia including brain tumor. Restoration of miR-137 expression has also been shown to inhibit cell proliferation, migration and metastasis, and induce cell cycle arrest, differentiation and apoptosis. These properties of miR-137 propose its potential for prognosis, diagnosis and as a therapeutic target for treatment of several human neurological and neoplastic disorders. In this review, we provide details on the discovery, targets, function, regulation and disease involvement of miR-137 with a broad look at recent discovery in this area.
Similar content being viewed by others
Introduction
MicroRNAs (miRNAs) are a large class of non-coding RNAs regulating many biological processes in a wide range of organisms including plants, animal species and virus.1 These molecules exert their regulatory roles by modulation of gene expression through degradation of mRNA or suppression of protein synthesis.2 Over the last few years, miRNAs have been the focus of many studies, providing evidence that these small RNAs have a significant role in the modification of numerous biological pathways.3 It is also evident that any abnormality in miRNA function can cause cellular malfunction, which in turn may lead to a disorder.4
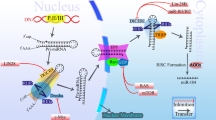
MiRNA genes are initially transcribed into a primary RNA transcript (pri-miR) before their stem loop structure undergoes processing in the nucleus to form a ~70 nt long intermediate termed the precursor miRNA (pre-miR). This is performed by the microprocessor complex comprising the RNase III Drosha and a double-stranded RNA-binding protein, DiGeorge syndrome critical region gene 8 (DGCR8).5, 6 Pre-miRs are then transported to the cytoplasm through the nuclear pores by Exportin-5 complex,7 where further processing occurs by a second RNase III known as Dicer and its associated trans-activation response RNA binding protein (TRBP) to yield double stranded RNAs, 18–24 nt in length.8, 9, 10 These are subsequently unwound and the mature single-stranded miRNA is incorporated into the argonaute protein-containing RNA-induced silencing complex (RISC). In this complex the miRNA functions as a template for homology directed mRNA targeting of the 3′ untranslated region (UTR).11, 12 The intermolecular base pairing occurs primarily at 3′ UTR segments with sequence complementarity to the miRNA ‘seed’ region consisting of 6 nt from positions 2 to 7 or 813 (Figure 1).
Schematic of miRNA biogenesis pathway and function. MicroRNA (miRNA) is initially transcribed by RNA pol II transcription factor and form a hairpin structure termed pri-miRs. Then Drosha and DGCR8 complex cleave the molecule producing pre-miRs in the nucleus. Exportin-5 transfers the double strand miRNA into the cytoplasm where miRNA undergo further editing by TRBP and Dicer complex to form miRNA duplex, which are eventually unwound to reveal the mature miRNA. MiRNA interact within a RISC complex and provide guidance for targeting mRNAs for degradation or translational repression.
MiRNAs are able to posttranscriptionally regulate a large number of genes by either mRNA degradation or translational inhibition.14 Although some miRNAs are widely expressed others have stage, tissue or cell type-specific expression patterns.15 MiRNA genes can be exonic, intronic or reside within intergenic transcripts.16 Significantly, mature miRNA can bind to their target 3′ UTRs despite mismatched pairing, enabling each miRNA to regulate hundreds of mRNA targets.17 Multiple miRNA can also regulate a single gene,18, 19 such that different miRNAs are able to cooperate in a given pathway and share their regulatory responsibilities.
Bioinformatic studies estimate that miRNA regulate expression of up to 80% of genes in human20 encompassing a wide range of cellular functions such as development,21 cell proliferation,22 differentiation,23 apoptosis,24 signal transduction25 and cell cycle.26 MiR-137 is one of several miRNAs that have a significant role in cellular biology, localizing on human chromosome 1p22 within the gene for the non-protein-coding RNA AK094607.27 MiR-137 has a critical regulatory role in brain function, with several reports displaying an association with proliferation and differentiation during development.28 The miR-137 host gene is strongly associated with the etiology of many psychiatric disorders including schizophrenia (SZ)29 and bipolar disorder (BD).30 Dysfunction of miR-137 has also been shown to contribute to several human cancer types such as neuroblastoma31 and glioblastoma multiforme.32
In this review, we summarize our current understanding of miR-137 in cell biology and its association with human disorders with an eye on the potential use of miR-137 as a prognostic/diagnostic signature and target for therapeutic development.
Discovery and targets
MiR-137 is a classic 23-nt microRNA located within a long non-coding host gene, MIR137HG. This transcriptional unit on chromosome 1 is over 62 kb long and is predicted to produce four splice variants ranging from over 2.5 kb to below 1 kb. There is also a 15-bp variable tandem repeat localized in the primary transcript of miR-137, 5′ to the pre-miR-137, shown to change the posttranscriptional processing of miR-137 through alteration of its secondary structure27 (Figure 2).
Genomic location, transcripts and sequence of miR-137 in human. The gene is 61 kb long situated in short arm of chromosome 1 producing four transcripts. The pri-miR is 102 nt in length which is cleaved to a 23-nt mature miRNA (miR-137).
The first report of miR-137 dates back to 2002, where a study aimed to identify functional miRNAs and their tissue-specific expression pattern in mammalians. Lagos-Quintana et al.33 cloned microRNAs from nine different adult mouse tissues including heart, liver, small intestine, colon, cortex, cerebellum and midbrain, with 34 novel miRNAs, including miR-137, being identified specifically in midbrain cortical tissue. Later, another study by Landgraf and colleagues validated this miRNA expression in various human and rodent tissue, along with over 250 other miRNA. This miRNA was also found to be enriched in the nervous system.34
Using a bioinformatic approach, over 1000 putative genes have been predicted to be targets of miR-137.35 Around 5% of these have been experimentally validated (Figure 3); mostly via gene expression profiling, luciferase expression reporter and western blot assay in a wide variety of transfected cell lines. These functional analyses are summarized in Table 1, and highlight target genes involved in a large number of pathways including neural development, cell cycle, differentiation and proliferation.
Experimentally validated target genes of miR-137. Approximately 50 genes have been identified to be directly regulated by miR-137 and most confirmed by luciferase assay and western blot. The target genes of miR-137 are involved in different biological pathways mainly cell cycle, proliferation and differentiation.
Function
A number of studies have demonstrated that microRNA networks are responsible for posttranscriptional modulation of gene expression. They are present in almost all tissue types and mediate a vast range of biological processes (Table 1). MiR-137 is a brain-enriched miRNA in mouse and human33, 36, 37 with a high expression in the cortical brain regions, and hippocampus and low expression in cerebellum and brain stem.37 This is consistent with evidence from several studies suggesting that it regulates cell proliferation and differentiation, in both the embryonic and adult brain (Figure 4). Sun et al. demonstrated that miR-137 promoted cell differentiation while inhibiting proliferation of embryonic neuronal stem cell by downregulating the LSD1 gene through a regulatory loop with the transcriptional co-repressor TLX.28 Work by Tarantino et al. established that miR-137 was increased upon differentiation and functional analysis demonstrated that miR-137 could directly target Jarid1b, also known as KDM5b, resulting in differentiation of mouse embryonic stem cells (ESCs).38 MiR-137 was found to be expressed at the mitotic phase of the cell cycle and highly upregulated during differentiation of ESCs into neural cells.39 This upregulation led to repression of two ES cell transcription factors, Klf4 and Tbx3, which were directly targeted by miR-137 (Figure 4a). Similar findings were obtained from a study showing that miR-137 promoted neural differentiation of various stem cells including adult mouse neural stem cells, mouse oligodendroglioma-derived stem cells and human glioblastoma multiforme-derived stem cells.32 In contrast, a study confirming the role of miR-137 in modulation of proliferation and differentiation concluded the opposite effects in adult neural stem cells (aNSCs) by showing that miR-137 enhanced proliferation and inhibited differentiation through posttranscriptional suppression of Ezh2.40 These results suggest that there might be a context-dependent role for miR-137 in neural stem cells. Also, miR-137 was identified to have an important role in regulating neuronal maturation. Smrt et al. reported that Mib1, known to be associated in neurodevelopment, is negatively modulated by miR-137 through which dendritic morphogenesis, phenotypic maturation and spine development were affected in newborn neurons of the adult hippocampus41 (Figure 4b). We recently analyzed the microRNA and mRNA expression in midbrain and forebrain in the rat embryo and observed upregulation of miR-137 across brain development.42 We further identified two neural development-associated genes, Wnt7a and Gpr88, as direct targets of miR-137 using luciferase assay. Interestingly, a recent study revealed that embryonic development is dependent on the presence of at least one functional allele of MIR137,43 with embryonic lethality occurring in all homozygous (−/−) embryos for miR-137 allele. No phenotype was observed in miR-137 heterozygous mice presumably as a consequence of dosage compensation from the remaining allele, as there was no difference in mature miRNA expression compared with the homozygotes.
The impact of miR-137 on neural differentiation, proliferation and maturation in ESCs and newborn neurons. (a) Upregulation of miR-137 can suppress target mRNAs such as TLX, LSD1, KDM5B and Tbx3 resulting in the neural stem cell move to cell differentiation, whereas low levels of this miRNA cause an increase in target genes expression that lead to cell proliferation.28, 38, 39 (b) MiR-137 regulate dendritic complexity and spine numbers by targeting Mib1 gene in newborn neurons.41 ESC, embryonic stem cells; miRNA, microRNA.
Taken together, these results suggest that miR-137 has an important role during neural development through regulation of target genes associated with neural stem cell proliferation and differentiation. These studies further support miR-137s contribution to the regulation of transition from pluripotentency to differentiated states. Given miR-137s involvement in these processes as well as apoptosis, it is perhaps not surprising that its depletion is also associated with the development and progression of cancers (discussed in detail below).
Regulation
MiRNAs are important regulators of gene expression, however, they can also modify other epiphenomena by modulating the expression of target gene networks in epigenetic mechanisms. Some miRNAs, known as epi-miRNAs, have even been implicated more directly through their capacity to guide some of the enzymes associated with modulation of epigenetic machinery. MiRNA expression can also be affected by epigenetic modulation of their host gene promoters.31
The MIR137 gene is itself highly regulated by large CpG island embedded in the upstream promoter region of its gene located on chromosome 1p22.44 These have been shown to be modified by DNA-hypermethylating agents, which result in suppression of miR-137 transcription.44 For instance, Chen et al. established that miR-137 is suppressed epigenetically in uveal melanoma cell lines and treatment with 5-aza-dC, a DNA methylation inhibitor and/or TSA, a histone deacetylase inhibitor, could restore the expression of miR-137.45 Another study on miR-137 has indicated that it was also silenced as a result of aberrant DNA methylation in oral squamous cell carcinoma.44 Similarly, evidence from a report in glioblastoma validated that miR-137 was silenced by hypermethylation.32 Consistent with previous studies, Zhu and colleagues found that levels of miR-137 were suppressed in non-small cell lung carcinoma (NSCLC) but restoration of activity was observed after treatment with DNA-hypomethylating agents.46 Real-time results showed that miR-137 transcripts increased up to eight-fold following 5-aza-dC treatment, and up to nine-fold in cells treated with both 5-aza-dC and TSA, indicating the role of epigenetic modification of regulatory sequences in CpG islands in miR-137 silencing. Further supporting data from studies on colorectal cancer,47 gastric cancer48, 49 and squamous cell carcinoma of the head and neck50, 51 demonstrated methylation-silencing of miR-137. In mouse adult neuronal stem cells, the expression of miR-137 is affected through epigenetic regulation mediated by the DNA methyl-CpG-binding protein MeCP2,52 and transcription factor Sox2.40 It was shown that direct binding of MeCP2 and Sox2 to the regulatory region of miR-137, in the 2.5 kb upstream region, is essential for proper regulation of miR-137 in aNSCs.40 These findings strongly implicate GpC methylation and silencing as an important mechanism in the deregulation of miR-137.
MiR-137 activity is also involved in the modulation of genes interacting in epigenetic pathways. For example, Szulwach et al. discovered that miR-137 modulated the levels of Ezh2 directly in aNSCs.40 Ezh2 is a H3-K27 methyltransferase and component of the polycomb group (PcG) proteins, which are known to remodel chromatin and induce epigenetic silencing. PcG proteins function by creating and maintaining repressive chromatin environments, allowing cell/tissue-specific genes to be primed for expression.53, 54 Overexpression of miR-137 was able to posttranscriptionally suppress the expression of Ezh2 leading to a global decrease in histone H3 trimethyl lysine 27.40 MiR-137 was also shown to target the downstream transcriptional co-repressor, histone lysine-specific demethylase 1 (LSD1).28 This is important in TLX signaling, which can induce feedback repression of miR-137 through the recruitment of LSD1 to the miRNA genes promoter. Similarly Jarid1b, also known as KDM5b (expressed in ESCs and in the very early stages of embryonic development) was shown to be a direct target of miR-137.39 Jarid1b specifically demethylates lysine 4 of histone H3 (meH3K4), which leads to inhibition of RNA polymerase and transcription factor binding within promoter elements,55), thus repressing gene transcription. Collectively, these findings suggest that miR-137 has a significant role in epigenetic regulation and its dysregulation can affect or be affected by epigenetic modulation.
Psychiatric disorders
Prior to any neurological disease association, several groups provided evidence to suggest that miR-137 is associated with the regulation of adult neurogenesis, dendritic development and neuronal maturation40 as well as control of the dynamics between neural stem cell proliferation and differentiation during neural development.28 With this background, it was perhaps not surprising that miR-137 has been reported to be genetically associated with mental illness. This is supported by molecular neuropathology and expression analysis in peripheral blood.
MiR-137 in psychotic disorders
SZ is a severe and heritable psychiatric disorder affecting ~1% of the general population worldwide,56 which shares some genetic risk factors and abnormalities in brain structure with BD.57, 58 Before the discovery of a genetic link with SZ, variant gene-sets associated with increased BOLD activation in functional imaging of SZ patients, were observed to be enriched with miR-137 target genes.59 In the first large-scale mega genome-wide association studies of SZ, involving more than 20 000 samples, the MIR137 host gene (MIR137HG) was implicated as a susceptibility locus through its proximity to the associated tag single-nucleotide polymorphism (SNP) rs1625579.29 This association was also supported in a cohort of Han Chinese60 and further replicated in Europeans.61 In addition, four predicted miR-137 target genes including TCF4, CACNA1C, CSMD1 and C10orf26 were the other associated loci, suggesting that convergent pathways linked by the miRNA may contribute to this complex neuropsychiatric disorder. The 3′ UTR of these targets were subsequently shown to be modulated by miR-137 in the context of a reporter gene assay.62 Pathway analyses indicated several genes with predicted miR-137 target sites were significantly enriched among smaller association P-values for SZ. Further upstream, rs1198588 was shown to be a significant local expression quantitative trait locus (eQTL) for MIR137 and DPYD—a protein-coding gene in the same linkage disequilibrium block as MIR137 and also predicted target of miR-137.61 More recently in the landmark mega genome-wide association studies (almost 37 000 cases and 113 000 controls) 108 independent SZ risk loci reasserted the MIR137 association (rs1702294, P=3.4 × 10−19).63 Again, gene set analysis of this data also showed the enrichment of putative miR-137 targets for SZ associations.63
In candidate-based phenotypic studies, the risk allele was shown to be associated with lower P300 amplitude.64 Carriers also displayed less positive symptom (lower OPCRIT scores <10−5) and more cognitive deficits involving episodic memory and attentional control in a cohort of SZ, schizoaffective disorder and bipolar affective disorder.65 Similarly, carrier considered high risk for SZ and BD, had reduced response in the right posterior medical frontal gyrus region in functional magnetic resonance imaging. Moreover, the SZ high-risk group had differential activation in the left amygdala and left pre/postcentral gyrus, suggesting the susceptibility allele may affect SZ more specifically.30 Van Erp and colleagues performed a study on 111 subjects including 44 SZ patients and 61 healthy controls to realize the functional role of miR-137 locus risk genotype.66 Using and Sternberg item recognition paradigm (SIRP) data from participants, they found that those with the rs1625579 TT genotype had significantly higher left dorsolateral prefrontal cortex activation than subjects with the GG/GT genotypes, suggesting the association of miR-137 risk genotype with the SZ risk phenotype dorsolateral prefrontal cortex hyperactivation. In support of the functional impact of miR-137 risk variant, it was established that it was predictive of phenotypic variability. A significantly earlier age-at-onset of psychosis was found for ‘TT’ homozygotes compared with protective G-allele carriers.67 Also, brain structure of SZ subjects was affected by the risk genotype, such that those with the miR-137 risk variant had lower white matter integrity, also decreased hippocampi and larger lateral ventricle volumes, while patients carrying the protective allele showed no difference from healthy controls in this regard. In post-mortem analysis, miR-137 has not been shown to be differentially expressed in SZ including studies in our own lab.68, 69 Guella and colleagues, however, showed that while miR-137 was not differentially expressed in the disorder, the levels were significantly reduced in the common risk variant when stratified by the rs1625579 genotype.70 Our team investigated the MIR137 SNP association with specific phenotypic characteristics of SZ indicating that the risk genotype was able to subtype SZ.71 Grade of Membership analysis yielded two patient categories including cognitive deficit group and cognitive spared group, with ‘G’ allele predicting cognitive deficit group status in patients with significant negative symptoms. This was surprising given that the T allele is associated with the risk of SZ. Further investigation of the risk haplotype associated with the tag SNP rs1625579 using promoterless luciferase reporter assay, suggest that miR-137 expression levels are affected by other functionally implicated SNPs including rs2660304 which are in linkage disequilibrium.72, 73 Overexpression of miR-137 in a human neural stem cell line led to differential expression of the genes involved in pathways implicated in SZ including major histocompatibility complex, synapses, FMRP interacting RNAs and calcium channels after the 48 h, providing more support for a role in the etiology of the disorder.74
Major depression
MiR-137 has also been shown to have anti-depressive effects in a post-stroke depression rat model.75 Zhao et al. found that miR-137 levels were significantly lower in the brain and peripheral blood compared with controls. Exogenous delivery of miR-137 in vivo led to improvement of behavioral changes in post-stroke depression rats.75 These impacts were shown to be as a result of miR-137’s suppressive role on Grin2A protein through binding to Grin2A mRNA. MicroRNA expression profiling in prefrontal cortex of depressed suicide subjects demonstrated that miR-137 is downregulated in patients suggesting the involvement of miR-137 in mood disorders.76 In addition, a risk allele within CACNA1C, a target gene of miR-137, is associated with the risk of major depression as well as SZ and BD.77 Other reports identified statistical association between this miR-137 target gene, CACNA1C, and major depressive disorder.78, 79
Autism spectrum disorders
The MIR137 gene has also been implicated in autism spectrum disorders (ASDs). In a study by Devanna and Vernes provided support for MIR137 as an autism candidate by showing that the RORa gene is regulated directly by miR-137 through targeting its 3′ UTR in a site-specific manner.80 RORa, a novel autism candidate gene, is known as a transcription regulator encoding a ligand-dependent orphan nuclear receptor whose target genes are associated with susceptibility to ASDs and was found to be important in pathways implicated in ASDs.81 Further analysis demonstrated that two synaptic genes, SHANK3 and NRXN1, previously identified as autism candidate genes were also putatively targeted by miR-137.81 SHANK3 is involved in synapse formation and function and NRXN1 was found as a risk factor for both autism and SZ with a SNP associated with changes in frontal white matter volume—an endophenotype common to both disorders.82 Findings from other genome-wide association studies have identified SNPs in the human miR-137 gene locus that are associated with an increased risk of SZ and ASDs.61, 83 These findings were supported with reports that autism spectrum disorders are related to microdeletions of this region.37, 84
Huntington’s disease
Recent discoveries implicated miRNA in posttranscriptional dysregulation in Huntington’s disease (HD) and many crucial neural-specific miRNAs are aberrantly repressed in this disorder.85 REST is one of the first transcription factors to be shown to modulate miRNA transcription, appearing to be an important regulator of neuron-specific miRNAs. Evidence from work on HD, using Hdh7/7 and Hdh109/109 cells, used extensively as a cellular model of molecular dysfunction in HD, identified several miRNAs, including miR-137, whose aberrant repression was directly mediated by REST.86 MiR-137 was upregulated in Hdh109/109 cells by REST, whereas REST knock down resulted in decreased miR-137 expression, suggesting that REST is a direct regulator of this miRNA. In the presence of muHtt, mutated form of Htt causing HD, REST protein translocates to the nucleus where it represses transcription of its target genes that are significant for nervous system development and function such as miR-137.
Rett syndrome
A genome-wide analysis in a mouse model of Rett syndrome found that upregulation of miRNAs, including miR-137, were implicated in its pathoetiology.87 This disorder is caused by the mutations in MECP2 gene, which normally negatively regulate the expression of the aberrant miRNAs in brain by directly targeting their methylated promoters. Combined ChIP-chip and expression analyses suggested a model in which Mecp2 directly binds to DNA methylated regions within promoter of miRNA and acts as a transcriptional repressor. Furthermore, in addition to miR-137, several Mecp2-regulated miRNAs are implicated in regulation of dendritogenesis and synapse formation/maturation,88, 89 therefore dysregulation of synapse-localized miRNAs such as miR-137 may contribute to neurological disorders including Rett syndrome. Mecp2-regulated miRNAs, including miR-137, in neurons therefore may act as a critical mechanistic link between nucleus-localized Mecp2 and cytoplasmic/synaptic proteins.
Intellectual disability
Chromosomal microdeletion of 1p21.3 encompassing miR-137 and DPYD have also been observed in ID, with a corresponding reduction of precursor and mature miR-137 and significantly increased levels of downstream targets including MITF, EZH2 and KLF4 in patients compared with controls.37
These genetic association, expression and functional studies support a role for miR-137 pathways in neural processes and neurocognitive disorders. This is consistent with molecular systems analysis of miR-137 implicating pathways in neurogenesis and brain development. High expression of miR-137 have been reported to be present at the synapse where it has a role in controlling dendritic spine maturation and density.41 Alteration of dendritic spine number, density and volume have been shown in SZ and ASDs, with dendritic spine reduced in SZ, but increased in ASD.90, 91 MiR-137 therefore appears to be a central network hub with significant implications for its dysregulation in the etiology of multiple psychiatric and neurocognitive disorders.
Cancer
Several miRNAs have been implicated in the pathogenesis of human cancer. These oncogenenic-miRNA (Onco-miRs) or tumor suppressor-miRs (Ts-miRNAs) are associated with the early stages of carcinogenesis and may also participate in progression and metastasis.92 This is supported by miRNA expression-profiling studies showing a reduction in miR-137 expression in the pathogenesis of numerous malignancies. For instance, in glioblastoma multiforme, which is the most frequent and malignant of primary brain tumors, a cohort of microRNAs, including miR-137, were identified to be significantly downregulated.32 By contrast its overexpression led to inhibition of cell proliferation and invasion in glioma cell lines, perhaps as a result of inverse regulation of Cox-2.93, 94 Li et al. further established that this miR-137 negatively regulated cell viability, anchorage-independent growth and invasion through modulation of CSE1L expression.94 Functional studies in several melanoma cell lines also revealed that miR-137 was able to inhibit melanoma cell invasion, migration, proliferation and induced apoptosis via downregulation of multiple target genes including transcription factor MITF and the oncogenes c-Met, YB1, EZH2 and PAK2, suggesting miR-137 involvement in several significant pathways in melanoma development and progression.95, 96 Lower miR-137 expression contributed to poor survival in stage IV melanoma patients.95 MiR-137 appears to also act as a tumor suppressor in colon cancer.97 MSI1, a stem cell regulator with high expression in colorectal cancer was downregulated by miR-137 in colon tumors leading to reduction in cell growth, colony formation and tumorsphere growth in vitro as well as inhibition of tumor growth in vivo. Restoration of miR-137 in these cell lines negatively regulated two significant oncogenic signaling pathways, Wnt and Notch, which are known to be associated with colorectal cancer progression and modulated by MSI1. Quantitative PCR analysis of a large group of miRNAs in colorectal cancer also indicated that miR-137 is downregulated compared with normal tissue.98 Similarly, genome-wide miRNA expression profiling of a large cohort of breast tumor samples observed significant change in the levels of miR-137 as well as 12 other miRNA compared with normal tissue.99 Moreover, examination of a large number of miRNAs by microarray and then qPCR assays placed miR-137 among the significantly downregulated miRNAs in three different lung adenocarcinoma tumors.100 Bi et al. explored the regulatory function of miR-137 in progression of NSCLC discovering that miR-137 expression was negatively correlated with cell proliferation, cell migration and invasion, and was able to induce cell apoptosis in NSCLC cell lines through regulation of a direct target gene PXN,101 which is implicated in many human tumors102, 103 and involved in various physiological processes such as gene expression, tissue remodeling, cell proliferation and survival.104, 105 MiR-137 expression was low in lung cancer inducing G1 cell cycle arrest and suppressed cell growth both in vivo and in vitro.46 Overexpression of miR-137 was reported to decrease the expression of Cdc42 and Cdk6 and their immediate downstream effectors including cyclin D1, p-ERK1/2 and p-Rb, suggesting that miR-137 acts through the inhibition of Cdc42 and Cdk6 in lung cancer cells. In addition, expression profiles of human miRNAs in some tumor-derived cell lines,106 neuroblastoma107 and high-grade astrocytic tumors108 validated the decreases in miR-137 expression in tumor cells compared with controls, further supporting the role of miR-137 in pathogenesis of cancer. A role for miR-137 as a tumor suppressor is evident in several other malignancies including, pancreatic cancer,109 osteosarcoma cancer,110 gastric cancer,48 oral cancer50 and ovarian cancer.111
With the exception of bladder cancer and thyroid cancer,112, 113, 114 these findings show that a reduction of miR-137 is an important influence in the pathogenesis of cancer. In many cases it seems miR-137 downregulation is caused by DNA hypermethylation, suggesting that epigenetic modulation might be more significant than genetic mechanisms in oncological disorders.
Clinical approach
The discovery of miRNA’s role in cellular networks and disease has opened new possibilities in detection, diagnosis and treatment of complex disorders. This concept is also supported in several studies identifying promising miRNA candidates for diagnosis, prognosis and as therapeutic agents for clinical application.115, 116, 117 Restoration a miRNA function has been accomplished previously using synthetic miR-mimics or vector-based overexpression and these strategies could be employed for the clinical modulation of miR-137, which is downregulation in both cancer and psychiatric disorders. More specifically, intellectual disability associated with microdeletion and haploinsuficiency of miR-137 could be treated by exogenous supplementation in this way. Overexpression of mir-137 may also improve outcomes in ASD by reducing RORa expression. In SZ and BD where a common variant is associated with reduced expression, replacement of the miRNA may prevent onset or alleviate some of the symptoms where deficiency is indicated. Interestingly, delivery of miR-137 in vivo led to improvement of behavioral changes in post-stroke depression rats.75
In addition to supplementing with synthetic miR-137 or a transgene expression system, it may be possible to pharmacologically modulate miR-137 expression using small molecules. For example, it was shown that chromatin-demethylating agents, which transactivate miR-137 expression, resulted in suppression of NSCLC proliferation.46 Capecitabine chemoradiotherapy of rectal cancer was also shown to upregulate miR-137 in the most advanced T-stage.118 An indication of the therapeutic benefit of miR-137 was demonstrated in a study by Xiao et al., showing that upregulation of miR-137 increased sensitivities to chemotherapy reagent 5-FU in pancreatic cancer cells and reduced tumor weight and size in vivo.109 Similarly, modulation of miR-137 levels in the early stages or acute phase of neurodevelopmental disorders might help to alleviate these diseases.
There are a number of microRNA being pursued for therapeutic application, highlighting the feasibility of these molecules clinically. For example, delivery of miR- 34 and let-7 mimics inhibited tumor development in mouse models of lung and prostate cancers.119, 120, 121 Some other specific miRNAs currently in development as clinical candidates include miR-155, miR-21 and miR-33.122 Although miR-137 also presents as a promising candidate for therapeutic application, it will require significantly more preclinical testing in vivo, particularly for application in the central nervous system, where the blood–brain barrier remains as a significant hurdle for the delivery of large molecules and gene constructs.
Future perspectives
It is clear that this miR-137 is of considerable interest both in terms of its biological processes and association with major diseases. There are still many unexplored questions that arise from this exciting research, for example, a huge number of miR-137 target genes have been predicated,35 whereas only a small proportion have been investigated and confirmed. Pathway analysis of targets implicated several aspects of nervous system function and development associated with SZ, including ephrin receptor signaling and axonal guidance signaling as well as synaptic long-term potentiation.123, 124, 125 Although recent studies provide compelling link between MIR137 variants, such as rs1625579, and psychiatric disorder, the mechanisms causing dysregulation of miR-137 is not fully understood. Functional analysis of the variants in linkage disequilibrium with this tag SNP should provide new insight into this question. Siegert et al. provided evidence supporting a variant upstream (rs2660304) while also showing that this miRNA is involved in presynaptic plasticity through the regulation of several protein targets by miR-137.73 Surprisingly, developmental disruption of miR-137 in mice led to embryonic lethality,43 whereas heterozygous knockouts had no phenotype. Alternatively, we recently demonstrated that neurodevelopmental suppression miR-137 in the zebrafish resulted in severe sensorimotor deficits.126 This phenotype was not associated with any obvious change in the neuroanatomy or histology of the animals. More fine control of the miR-137 deficit in these models and more neurophysiological analysis will provide new insight to the biological basis of these behavioral phenotypes. Previously we observed that miR-137 expression was increased in telencephalon between E12 and E19 in the rat, whereas levels in the mesencephalon were fairly consistent from the earliest embryonic age.42 Although this suggested there is significant variation in the temporospatial expression pattern for miR-137, more details are required to better understand the expression of this molecule at different stages of development for various areas within cortical and subcortical structures.
Conclusions
In this review, we have outlined the compelling research suggesting that miR-137 is a significant network figure in the regulation of both neurodevelopmental processes and cancer biology. The development and maturation of neurons in embryonic and adult stem cells both in vivo and in vitro seem to be particularly sensitive to the expression of miR-137, with upregulation of miR-137 making a shift from proliferation to differentiation. The role of miR-137 in the induction of cell differentiation and inhibition of proliferation are also observed in cancer biology, suggesting tumor suppressive effects of this miRNA and significance for its repression in cancer biology. Interestingly, epidemiological studies have reported a potential link between SZ and cancer incidence, with these patients having a differential cancer risk.127 In addition to environmental, pharmacological and psychosomatic factors, genetic background was strongly implicated in this association and may possibly involve MIR137. Regardless, this molecule has a significant function in several neuropsychiatric/neurocognitive/oncological disorders and consequently may have great potential as a biomarker and in treatment of human diseases where dysregulation of this gene or its pathways are a factor. However, to reach this goal more exploration is required to completely understand the molecular mechanisms involved in disorders affected by this fascinating small RNA.
References
Bartel DP . MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 2004; 116: 281–297.
Filipowicz W, Bhattacharyya SN, Sonenberg N . Mechanisms of post-transcriptional regulation by microRNAs: are the answers in sight? Nat Rev Genet 2008; 9: 102–114.
Tufekci KU, Meuwissen RL, Genc S . The role of microRNAs in biological processes. Methods Mol Biol 2014; 1107: 15–31.
Esquela-Kerscher A, Slack FJ . Oncomirs - microRNAs with a role in cancer. Nat Rev Cancer 2006; 6: 259–269.
Gregory RI, Yan KP, Amuthan G, Chendrimada T, Doratotaj B, Cooch N et al. The microprocessor complex mediates the genesis of microRNAs. Nature 2004; 432: 235–240.
Lee Y, Ahn C, Han J, Choi H, Kim J, Yim J et al. The nuclear RNase III Drosha initiates microRNA processing. Nature 2003; 425: 415–419.
Denli AM, Tops BB, Plasterk RH, Ketting RF, Hannon GJ . Processing of primary microRNAs by the microprocessor complex. Nature 2004; 432: 231–235.
Kok KH, Ng MH, Ching YP, Jin DY . Human TRBP and PACT directly interact with each other and associate with dicer to facilitate the production of small interfering RNA. J Biol Chem 2007; 282: 17649–17657.
Chendrimada TP, Gregory RI, Kumaraswamy E, Norman J, Cooch N, Nishikura K et al. TRBP recruits the Dicer complex to Ago2 for microRNA processing and gene silencing. Nature 2005; 436: 740–744.
Hammond SM, Bernstein E, Beach D, Hannon GJ . An RNA-directed nuclease mediates post-transcriptional gene silencing in Drosophila cells. Nature 2000; 404: 293–296.
Diederichs S, Haber DA . Dual role for argonautes in microRNA processing and posttranscriptional regulation of microRNA expression. Cell 2007; 131: 1097–1108.
Rana TM . Illuminating the silence: understanding the structure and function of small RNAs. Nat Rev Mol Cell Biol 2007; 8: 23–36.
Bartel DP . MicroRNAs: target recognition and regulatory functions. Cell 2009; 136: 215–233.
Navarro F, Lieberman J . Small RNAs guide hematopoietic cell differentiation and function. J Immunol 2010; 184: 5939–5947.
Pasquinelli AE, Hunter S, Bracht J . MicroRNAs: a developing story. Curr Opin Genet Dev 2005; 15: 200–205.
Rodriguez A, Griffiths-Jones S, Ashurst JL, Bradley A . Identification of mammalian microRNA host genes and transcription units. Genome Res 2004; 14: 1902–1910.
Seitz H . Redefining microRNA targets. Curr Biol 2009; 19: 870–873.
Wu S, Huang S, Ding J, Zhao Y, Liang L, Liu T et al. Multiple microRNAs modulate p21Cip1/Waf1 expression by directly targeting its 3' untranslated region. Oncogene 2010; 29: 2302–2308.
Vo NK, Dalton RP, Liu N, Olson EN, Goodman RH . Affinity purification of microRNA-133a with the cardiac transcription factor, Hand2. Proc. Natl Acad Sci USA 2010; 107: 19231–19236.
Lu J, Clark AG . Impact of microRNA regulation on variation in human gene expression. Genome Res 2012; 22: 1243–1254.
Sayed D, Abdellatif M . MicroRNAs in development and disease. Physiol Rev 2011; 91: 827–887.
Lenkala D, LaCroix B, Gamazon ER, Geeleher P, Im HK, Huang RS . The impact of microRNA expression on cellular proliferation. Hum Genet 2014; 133: 931–938.
Ivey KN, Srivastava D . MicroRNAs as regulators of differentiation and cell fate decisions. Cell Stem Cell 2010; 7: 36–41.
Su Z, Yang Z, Xu Y, Chen Y, Yu Q . MicroRNAs in apoptosis, autophagy and necroptosis. Oncotarget 2015; 6: 8474–8490.
Inui M, Martello G, Piccolo S . MicroRNA control of signal transduction. Nat Rev Mol Cell Biol 2010; 11: 252–263.
Bueno MJ, Malumbres M . MicroRNAs and the cell cycle. Biochim Biophys Acta 2011; 1812: 592–601.
Bemis LT, Chen R, Amato CM, Classen EH, Robinson SE, Coffey DG et al. MicroRNA-137 targets microphthalmia-associated transcription factor in melanoma cell lines. Cancer Res 2008; 68: 1362–1368.
Sun G, Ye P, Murai K, Lang MF, Li S, Zhang H et al. miR-137 forms a regulatory loop with nuclear receptor TLX and LSD1 in neural stem cells. Nat Commun 2011; 2: 529.
Schizophrenia Psychiatric Genome-Wide Association Study C. Genome-wide association study identifies five new schizophrenia loci. Nat Genet 2011; 43: 969–976.
Whalley HC, Papmeyer M, Romaniuk L, Sprooten E, Johnstone EC, Hall J et al. Impact of a microRNA MIR137 susceptibility variant on brain function in people at high genetic risk of schizophrenia or bipolar disorder. Neuropsychopharmacology 2012; 37: 2720–2729.
Althoff K, Beckers A, Odersky A, Mestdagh P, Koster J, Bray IM et al. MiR-137 functions as a tumor suppressor in neuroblastoma by downregulating KDM1A. Int J Cancer 2013; 133: 1064–1073.
Silber J, Lim DA, Petritsch C, Persson AI, Maunakea AK, Yu M et al. miR-124 and miR-137 inhibit proliferation of glioblastoma multiforme cells and induce differentiation of brain tumor stem cells. BMC Med 2008; 6: 14.
Lagos-Quintana M, Rauhut R, Yalcin A, Meyer J, Lendeckel W, Tuschl T . Identification of tissue-specific microRNAs from mouse. Curr Biol 2002; 12: 735–739.
Landgraf P, Rusu M, Sheridan R, Sewer A, Iovino N, Aravin A et al. A mammalian microRNA expression atlas based on small RNA library sequencing. Cell 2007; 129: 1401–1414.
Wright C, Turner JA, Calhoun VD, Perrone-Bizzozero N . Potential impact of miR-137 and its targets in schizophrenia. Front Genet 2013; 4: 58.
Lagos-Quintana M, Rauhut R, Lendeckel W, Tuschl T . Identification of novel genes coding for small expressed RNAs. Science 2001; 294: 853–858.
Willemsen MH, Valles A, Kirkels LA, Mastebroek M, Olde Loohuis N, Kos A et al. Chromosome 1p21.3 microdeletions comprising DPYD and MIR137 are associated with intellectual disability. J Med Genet 2011; 48: 810–818.
Tarantino C, Paolella G, Cozzuto L, Minopoli G, Pastore L, Parisi S et al. miRNA 34a, 100, and 137 modulate differentiation of mouse embryonic stem cells. FASEB J 2010; 24: 3255–3263.
Jiang K, Ren C, Nair VD . MicroRNA-137 represses Klf4 and Tbx3 during differentiation of mouse embryonic stem cells. Stem Cell Res 2013; 11: 1299–1313.
Szulwach KE, Li X, Smrt RD, Li Y, Luo Y, Lin L et al. Cross talk between microRNA and epigenetic regulation in adult neurogenesis. J Cell Biol 2010; 189: 127–141.
Smrt RD, Szulwach KE, Pfeiffer RL, Li X, Guo W, Pathania M et al. MicroRNA miR-137 regulates neuronal maturation by targeting ubiquitin ligase mind bomb-1. Stem Cells 2010; 28: 1060–1070.
Hollins SL, Goldie BJ, Carroll AP, Mason EA, Walker FR, Eyles DW et al. Ontogeny of small RNA in the regulation of mammalian brain development. BMC Genomics 2014; 15: 777.
Crowley JJ, Collins AL, Lee RJ, Nonneman RJ, Farrell MS, Ancalade N et al. Disruption of the microRNA 137 primary transcript results in early embryonic lethality in mice. Biol Psychiatry 2015; 77: e5–e7.
Kozaki K, Imoto I, Mogi S, Omura K, Inazawa J . Exploration of tumor-suppressive microRNAs silenced by DNA hypermethylation in oral cancer. Cancer Res 2008; 68: 2094–2105.
Chen X, Wang J, Shen H, Lu J, Li C, Hu DN et al. Epigenetics, microRNAs, and carcinogenesis: functional role of microRNA-137 in uveal melanoma. Invest Ophthalmol Vis Sci 2011; 52: 1193–1199.
Zhu X, Li Y, Shen H, Li H, Long L, Hui L et al. miR-137 inhibits the proliferation of lung cancer cells by targeting Cdc42 and Cdk6. FEBS Lett 2013; 587: 73–81.
Balaguer F, Link A, Lozano JJ, Cuatrecasas M, Nagasaka T, Boland CR et al. Epigenetic silencing of miR-137 is an early event in colorectal carcinogenesis. Cancer Res 2010; 70: 6609–6618.
Chen Q, Chen X, Zhang M, Fan Q, Luo S, Cao X . miR-137 is frequently down-regulated in gastric cancer and is a negative regulator of Cdc42. Dig Dis Sci 2011; 56: 2009–2016.
Steponaitiene R, Kupcinskas J, Langner C, Balaguer F, Venclauskas L, Pauzas H et al. Epigenetic silencing of miR-137 is a frequent event in gastric carcinogenesis. Mol Carcinog 2016; 55: 376–386.
Langevin SM, Stone RA, Bunker CH, Grandis JR, Sobol RW, Taioli E . MicroRNA-137 promoter methylation in oral rinses from patients with squamous cell carcinoma of the head and neck is associated with gender and body mass index. Carcinogenesis 2010; 31: 864–870.
Langevin SM, Stone RA, Bunker CH, Lyons-Weiler MA, LaFramboise WA, Kelly L et al. MicroRNA-137 promoter methylation is associated with poorer overall survival in patients with squamous cell carcinoma of the head and neck. Cancer 2011; 117: 1454–1462.
Bird A . DNA methylation patterns and epigenetic memory. Genes Dev 2002; 16: 6–21.
Boyer LA, Plath K, Zeitlinger J, Brambrink T, Medeiros LA, Lee TI et al. Polycomb complexes repress developmental regulators in murine embryonic stem cells. Nature 2006; 441: 349–353.
Lee TI, Jenner RG, Boyer LA, Guenther MG, Levine SS, Kumar RM et al. Control of developmental regulators by Polycomb in human embryonic stem cells. Cell 2006; 125: 301–313.
Sims RJ 3rd, Nishioka K, Reinberg D . Histone lysine methylation: a signature for chromatin function. Trends Genet 2003; 19: 629–639.
van Os J, Kapur S . Schizophrenia. Lancet 2009; 374: 635–645.
Lichtenstein P, Yip BH, Bjork C, Pawitan Y, Cannon TD, Sullivan PF et al. Common genetic determinants of schizophrenia and bipolar disorder in Swedish families: a population-based study. Lancet 2009; 373: 234–239.
International Schizophrenia C Purcell SM, Wray NR, Stone JL, Visscher PM, O'Donovan MC et al. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 2009; 460: 748–752.
Potkin SG, Macciardi F, Guffanti G, Fallon JH, Wang Q, Turner JA et al. Identifying gene regulatory networks in schizophrenia. Neuroimage 2010; 53: 839–847.
Guan F, Zhang B, Yan T, Li L, Liu F, Li T et al. MIR137 gene and target gene CACNA1C of miR-137 contribute to schizophrenia susceptibility in Han Chinese. Schizophr Res 2014; 152: 97–104.
Ripke S, O'Dushlaine C, Chambert K, Moran JL, Kahler AK, Akterin S et al. Genome-wide association analysis identifies 13 new risk loci for schizophrenia. Nat Genet 2013; 45: 1150–1159.
Kwon E, Wang W, Tsai LH . Validation of schizophrenia-associated genes CSMD1, C10orf26, CACNA1C and TCF4 as miR-137 targets. Mol Psychiatry 2013; 18: 11–12.
Schizophrenia Working Group of the Psychiatric Genomics C. Biological insights from 108 schizophrenia-associated genetic loci. Nature 2014; 511: 421–427.
Decoster J, De Hert M, Viechtbauer W, Nagels G, Myin-Germeys I, Peuskens J et al. Genetic association study of the P300 endophenotype in schizophrenia. Schizophr Res 2012; 141: 54–59.
Cummings E, Donohoe G, Hargreaves A, Moore S, Fahey C, Dinan TG et al. Mood congruent psychotic symptoms and specific cognitive deficits in carriers of the novel schizophrenia risk variant at MIR-137. Neurosci Lett 2013; 532: 33–38.
van Erp TG, Guella I, Vawter MP, Turner J, Brown GG, McCarthy G et al. Schizophrenia miR-137 locus risk genotype is associated with dorsolateral prefrontal cortex hyperactivation. Biol Psychiatry 2014; 75: 398–405.
Lett TA, Chakravarty MM, Felsky D, Brandl EJ, Tiwari AK, Goncalves VF et al. The genome-wide supported microRNA-137 variant predicts phenotypic heterogeneity within schizophrenia. Mol Psychiatry 2013; 18: 443–450.
Santarelli DM, Beveridge NJ, Tooney PA, Cairns MJ . Upregulation of dicer and microRNA expression in the dorsolateral prefrontal cortex Brodmann area 46 in schizophrenia. Biol Psychiatry 2011; 69: 180–187.
Beveridge NJ, Gardiner E, Carroll AP, Tooney PA, Cairns MJ . Schizophrenia is associated with an increase in cortical microRNA biogenesis. Mol Psychiatry 2010; 15: 1176–1189.
Guella I, Sequeira A, Rollins B, Morgan L, Torri F, van Erp TG et al. Analysis of miR-137 expression and rs1625579 in dorsolateral prefrontal cortex. J Psychiatr Res 2013; 47: 1215–1221.
Green MJ, Cairns MJ, Wu J, Dragovic M, Jablensky A, Tooney PA et al. Genome-wide supported variant MIR137 and severe negative symptoms predict membership of an impaired cognitive subtype of schizophrenia. Mol Psychiatry 2013; 18: 774–780.
Warburton A, Breen G, Bubb VJ, Quinn JP . A GWAS SNP for schizophrenia is linked to the internal MIR137 promoter and supports differential allele-specific expression. Schizophr Bull 2016; 42: 1003–1008.
Siegert S, Seo J, Kwon EJ, Rudenko A, Cho S, Wang W et al. The schizophrenia risk gene product miR-137 alters presynaptic plasticity. Nat Neurosci 2015; 18: 1008–1016.
Collins AL, Kim Y, Bloom RJ, Kelada SN, Sethupathy P, Sullivan PF . Transcriptional targets of the schizophrenia risk gene MIR137. Transl Psychiatry 2014; 4: e404.
Zhao L, Li H, Guo R, Ma T, Hou R, Ma X et al. miR-137, a new target for post-stroke depression? Neural Regen Res 2013; 8: 2441–2448.
Smalheiser NR, Lugli G, Rizavi HS, Torvik VI, Turecki G, Dwivedi Y . MicroRNA expression is down-regulated and reorganized in prefrontal cortex of depressed suicide subjects. PLoS One 2012; 7: e33201.
Green EK, Grozeva D, Jones I, Jones L, Kirov G, Caesar S et al. The bipolar disorder risk allele at CACNA1C also confers risk of recurrent major depression and of schizophrenia. Mol Psychiatry 2010; 15: 1016–1022.
Shi J, Potash JB, Knowles JA, Weissman MM, Coryell W, Scheftner WA et al. Genome-wide association study of recurrent early-onset major depressive disorder. Mol Psychiatry 2011; 16: 193–201.
Casamassima F, Huang J, Fava M, Sachs GS, Smoller JW, Cassano GB et al. Phenotypic effects of a bipolar liability gene among individuals with major depressive disorder. Am J Med Genet B Neuropsychiatr Genet 2010; 153B: 303–309.
Devanna P, Vernes SC . A direct molecular link between the autism candidate gene RORa and the schizophrenia candidate MIR137. Sci Rep 2014; 4: 3994.
Sarachana T, Hu VW . Genome-wide identification of transcriptional targets of RORA reveals direct regulation of multiple genes associated with autism spectrum disorder. Mol Autism 2013; 4: 14.
Voineskos AN, Lett TA, Lerch JP, Tiwari AK, Ameis SH, Rajji TK et al. Neurexin-1 and frontal lobe white matter: an overlapping intermediate phenotype for schizophrenia and autism spectrum disorders. PLoS One 2011; 6: e20982.
Cross-Disorder Group of the Psychiatric Genomics C. Identification of risk loci with shared effects on five major psychiatric disorders: a genome-wide analysis. Lancet 2013; 381: 1371–1379.
Malhotra D, Sebat J . CNVs: harbingers of a rare variant revolution in psychiatric genetics. Cell 2012; 148: 1223–1241.
Johnson R, Zuccato C, Belyaev ND, Guest DJ, Cattaneo E, Buckley NJ . A microRNA-based gene dysregulation pathway in Huntington's disease. Neurobiol Dis 2008; 29: 438–445.
Soldati C, Bithell A, Johnston C, Wong KY, Stanton LW, Buckley NJ . Dysregulation of REST-regulated coding and non-coding RNAs in a cellular model of Huntington's disease. J Neurochem 2013; 124: 418–430.
Wu H, Tao J, Chen PJ, Shahab A, Ge W, Hart RP et al. Genome-wide analysis reveals methyl-CpG-binding protein 2-dependent regulation of microRNAs in a mouse model of Rett syndrome. Proc Natl Acad Sci USA 2010; 107: 18161–18166.
Schratt GM, Tuebing F, Nigh EA, Kane CG, Sabatini ME, Kiebler M et al. A brain-specific microRNA regulates dendritic spine development. Nature 2006; 439: 283–289.
Fiore R, Khudayberdiev S, Christensen M, Siegel G, Flavell SW, Kim TK et al. Mef2-mediated transcription of the miR379-410 cluster regulates activity-dependent dendritogenesis by fine-tuning Pumilio2 protein levels. EMBO J 2009; 28: 697–710.
Glantz LA, Lewis DA . Decreased dendritic spine density on prefrontal cortical pyramidal neurons in schizophrenia. Arch Gen Psychiatry 2000; 57: 65–73.
Hutsler JJ, Zhang H . Increased dendritic spine densities on cortical projection neurons in autism spectrum disorders. Brain Res 2010; 1309: 83–94.
Lu J, Getz G, Miska EA, Alvarez-Saavedra E, Lamb J, Peck D et al. MicroRNA expression profiles classify human cancers. Nature 2005; 435: 834–838.
Chen L, Wang X, Wang H, Li Y, Yan W, Han L et al. miR-137 is frequently down-regulated in glioblastoma and is a negative regulator of Cox-2. Eur J Cancer 2012; 48: 3104–3111.
Li KK, Yang L, Pang JC, Chan AK, Zhou L, Mao Y et al. MIR-137 suppresses growth and invasion, is downregulated in oligodendroglial tumors and targets CSE1L. Brain Pathol 2013; 23: 426–439.
Luo C, Tetteh PW, Merz PR, Dickes E, Abukiwan A, Hotz-Wagenblatt A et al. miR-137 inhibits the invasion of melanoma cells through downregulation of multiple oncogenic target genes. J Invest Dermatol 2013; 133: 768–775.
Hao S, Luo C, Abukiwan A, Wang G, He J, Huang L et al. miR-137 inhibits proliferation of melanoma cells by targeting PAK2. Exp Dermatol 2015; 24: 947–952.
Smith AR, Marquez RT, Tsao WC, Pathak S, Roy A, Ping J et al. Tumor suppressive microRNA-137 negatively regulates Musashi-1 and colorectal cancer progression. Oncotarget 2015; 6: 12558–12573.
Bandres E, Cubedo E, Agirre X, Malumbres R, Zarate R, Ramirez N et al. Identification by real-time PCR of 13 mature microRNAs differentially expressed in colorectal cancer and non-tumoral tissues. Mol Cancer 2006; 5: 29.
Iorio MV, Ferracin M, Liu CG, Veronese A, Spizzo R, Sabbioni S et al. MicroRNA gene expression deregulation in human breast cancer. Cancer Res 2005; 65: 7065–7070.
Dacic S, Kelly L, Shuai Y, Nikiforova MN . miRNA expression profiling of lung adenocarcinomas: correlation with mutational status. Mod Pathol 2010; 23: 1577–1582.
Bi Y, Han Y, Bi H, Gao F, Wang X . miR-137 impairs the proliferative and migratory capacity of human non-small cell lung cancer cells by targeting paxillin. Hum Cell 2014; 27: 95–102.
Kawada I, Hasina R, Lennon FE, Bindokas VP, Usatyuk P, Tan YH et al. Paxillin mutations affect focal adhesions and lead to altered mitochondrial dynamics: relevance to lung cancer. Cancer Biol Ther 2013; 14: 679–691.
Chen J, Gallo KA . MLK3 regulates paxillin phosphorylation in chemokine-mediated breast cancer cell migration and invasion to drive metastasis. Cancer Res 2012; 72: 4130–4140.
Deakin NO, Turner CE . Distinct roles for paxillin and Hic-5 in regulating breast cancer cell morphology, invasion, and metastasis. Mol Biol Cell 2011; 22: 327–341.
Turner CE . Paxillin and focal adhesion signalling. Nat Cell Biol 2000; 2: E231–E236.
Gaur A, Jewell DA, Liang Y, Ridzon D, Moore JH, Chen C et al. Characterization of microRNA expression levels and their biological correlates in human cancer cell lines. Cancer Res 2007; 67: 2456–2468.
Chen Y, Stallings RL . Differential patterns of microRNA expression in neuroblastoma are correlated with prognosis, differentiation, and apoptosis. Cancer Res 2007; 67: 976–983.
Godlewski J, Nowicki MO, Bronisz A, Williams S, Otsuki A, Nuovo G et al. Targeting of the Bmi-1 oncogene/stem cell renewal factor by microRNA-128 inhibits glioma proliferation and self-renewal. Cancer Res 2008; 68: 9125–9130.
Xiao J, Peng F, Yu C, Wang M, Li X, Li Z et al. microRNA-137 modulates pancreatic cancer cells tumor growth, invasion and sensitivity to chemotherapy. Int J Clin Exp Pathol 2014; 7: 7442–7450.
Li ZM, Zhang HY, Wang YX, Wang WB . MicroRNA-137 is downregulated in human osteosarcoma and regulates cell proliferation and migration through targeting FXYD6. J Drug Target 2015; 24: 102–110.
Guo J, Xia B, Meng F, Lou G . miR-137 suppresses cell growth in ovarian cancer by targeting AEG-1. Biochem Biophys Res Commun 2013; 441: 357–363.
Xiu Y, Liu Z, Xia S, Jin C, Yin H, Zhao W et al. MicroRNA-137 upregulation increases bladder cancer cell proliferation and invasion by targeting PAQR3. PLoS One 2014; 9: e109734.
Leonardi GC, Candido S, Carbone M, Colaianni V, Garozzo SF, Cina D et al. microRNAs and thyroid cancer: biological and clinical significance (review). Int J Mol Med 2012; 30: 991–999.
Marini F, Luzi E, Brandi ML . MicroRNA role in thyroid cancer development. J Thyroid Res 2011; 2011: 407123.
Iorio MV, Croce CM . MicroRNA dysregulation in cancer: diagnostics, monitoring and therapeutics. A comprehensive review. EMBO Mol Med 2012; 4: 143–159.
Hutchison ER, Okun E, Mattson MP . The therapeutic potential of microRNAs in nervous system damage, degeneration, and repair. Neuromolecular Med 2009; 11: 153–161.
Calin GA, Croce CM . MicroRNA signatures in human cancers. Nat Rev Cancer 2006; 6: 857–866.
Svoboda M, Izakovicova Holla L, Sefr R, Vrtkova I, Kocakova I, Tichy B et al. Micro-RNAs miR125b and miR137 are frequently upregulated in response to capecitabine chemoradiotherapy of rectal cancer. Int J Oncol 2008; 33: 541–547.
Wiggins JF, Ruffino L, Kelnar K, Omotola M, Patrawala L, Brown D et al. Development of a lung cancer therapeutic based on the tumor suppressor microRNA-34. Cancer Res 2010; 70: 5923–5930.
Liu C, Kelnar K, Liu B, Chen X, Calhoun-Davis T, Li H et al. The microRNA miR-34a inhibits prostate cancer stem cells and metastasis by directly repressing CD44. Nat Med 2011; 17: 211–215.
Trang P, Medina PP, Wiggins JF, Ruffino L, Kelnar K, Omotola M et al. Regression of murine lung tumors by the let-7 microRNA. Oncogene 2010; 29: 1580–1587.
van Rooij E, Purcell AL, Levin AA . Developing microRNA therapeutics. Circ Res 2012; 110: 496–507.
Xu NJ, Henkemeyer M . Ephrin reverse signaling in axon guidance and synaptogenesis. Semin Cell Dev Biol 2012; 23: 58–64.
Armstrong JN, Saganich MJ, Xu NJ, Henkemeyer M, Heinemann SF, Contractor A . B-ephrin reverse signaling is required for NMDA-independent long-term potentiation of mossy fibers in the hippocampus. J Neurosci 2006; 26: 3474–3481.
Contractor A, Rogers C, Maron C, Henkemeyer M, Swanson GT, Heinemann SF . Trans-synaptic Eph receptor-ephrin signaling in hippocampal mossy fiber LTP. Science 2002; 296: 1864–1869.
Giacomotto J, Carroll AP, Rinkwitz S, Mowry B, Cairns MJ, Becker TS . Developmental suppression of schizophrenia-associated miR-137 alters sensorimotor function in zebrafish. Transl Psychiatry 2016; 6: e818.
Hodgson R, Wildgust HJ, Bushe CJ . Cancer and schizophrenia: is there a paradox? J Psychopharmacol 2010; 24: 51–60.
Haflidadottir BS, Bergsteinsdottir K, Praetorius C, Steingrimsson E . miR-148 regulates Mitf in melanoma cells. PLoS One 2010; 5: e11574.
Liu LL, Lu SX, Li M, Li LZ, Fu J, Hu W et al. FoxD3-regulated microRNA-137 suppresses tumour growth and metastasis in human hepatocellular carcinoma by targeting AKT2. Oncotarget 2014; 5: 5113–5124.
Liu S, Cui J, Liao G, Zhang Y, Ye K, Lu T et al. MiR-137 regulates epithelial-mesenchymal transition in gastrointestinal stromal tumor. Tumour Biol 2014; 35: 9131–9138.
Liu M, Lang N, Qiu M, Xu F, Li Q, Tang Q et al. miR-137 targets Cdc42 expression, induces cell cycle G1 arrest and inhibits invasion in colorectal cancer cells. Int J Cancer 2011; 128: 1269–1279.
Kim AH, Parker EK, Williamson V, McMichael GO, Fanous AH, Vladimirov VI . Experimental validation of candidate schizophrenia gene ZNF804A as target for hsa-miR-137. Schizophr Res 2012; 141: 60–64.
Deng Y, Deng H, Bi F, Liu J, Bemis LT, Norris D et al. MicroRNA-137 targets carboxyl-terminal binding protein 1 in melanoma cell lines. Int J Biol Sci 2011; 7: 133–137.
Geekiyanage H, Chan C . MicroRNA-137/181c regulates serine palmitoyltransferase and in turn amyloid beta, novel targets in sporadic Alzheimer's disease. J Neurosci 2011; 31: 14820–14830.
Bier A, Giladi N, Kronfeld N, Lee HK, Cazacu S, Finniss S et al. MicroRNA-137 is downregulated in glioblastoma and inhibits the stemness of glioma stem cells by targeting RTVP-1. Oncotarget 2013; 4: 665–676.
Zhao Y, Li Y, Lou G, Zhao L, Xu Z, Zhang Y et al. MiR-137 targets estrogen-related receptor alpha and impairs the proliferative and migratory capacity of breast cancer cells. PLoS One 2012; 7: e39102.
Kozlowska E, Krzyzosiak WJ, Koscianska E . Regulation of huntingtin gene expression by miRNA-137, -214, -148a, and their respective isomiRs. Int J Mol Sci 2013; 14: 16999–17016.
Liang L, Li X, Zhang X, Lv Z, He G, Zhao W et al. MicroRNA-137, an HMGA1 target, suppresses colorectal cancer cell invasion and metastasis in mice by directly targeting FMNL2. Gastroenterology 2013; 144: 624–635.e4.
Acknowledgements
E Mahmoudi is supported by the University of Newcastle Faculty of Health and Medicine. MJ Cairns is supported by an M.C. Ainsworth Research Fellowship in Epigenetics; a NARSAD Independent Investigator Award; and an NHMRC project grant APP1067137.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Mahmoudi, E., Cairns, M. MiR-137: an important player in neural development and neoplastic transformation. Mol Psychiatry 22, 44–55 (2017). https://doi.org/10.1038/mp.2016.150
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mp.2016.150
This article is cited by
-
M2 Microglia-Derived Exosomes Protect Against Glutamate-Induced HT22 Cell Injury via Exosomal miR-124-3p
Molecular Neurobiology (2024)
-
Integrative Analysis of Morphine-Induced Differential Circular RNAs and ceRNA Networks in the Medial Prefrontal Cortex
Molecular Neurobiology (2023)
-
Analysis of the Relationship between Genetic Factors and the Risk of Schizophrenia
Neuroscience and Behavioral Physiology (2023)
-
The Significant Role of microRNAs in Gliomas Angiogenesis: A Particular Focus on Molecular Mechanisms and Opportunities for Clinical Application
Cellular and Molecular Neurobiology (2023)
-
Exosome-based strategies for diagnosis and therapy of glioma cancer
Cancer Cell International (2022)