Abstract
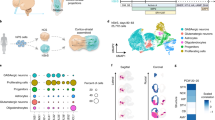
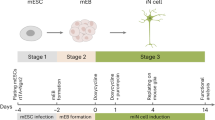
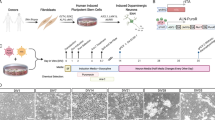
The brain’s serotonergic system centrally regulates several physiological processes and its dysfunction has been implicated in the pathophysiology of several neuropsychiatric disorders. While in the past our understanding of serotonergic neurotransmission has come mainly from mouse models, the development of pluripotent stem cell and induced fibroblast-to-neuron (iN) transdifferentiation technologies has revolutionized our ability to generate human neurons in vitro. Utilizing these techniques and a novel lentiviral reporter for serotonergic neurons, we identified and overexpressed key transcription factors to successfully generate human serotonergic neurons. We found that overexpressing the transcription factors NKX2.2, FEV, GATA2 and LMX1B in combination with ASCL1 and NGN2 directly and efficiently generated serotonergic neurons from human fibroblasts. Induced serotonergic neurons (iSNs) showed increased expression of specific serotonergic genes that are known to be expressed in raphe nuclei. iSNs displayed spontaneous action potentials, released serotonin in vitro and functionally responded to selective serotonin reuptake inhibitors (SSRIs). Here, we demonstrate the efficient generation of functional human serotonergic neurons from human fibroblasts as a novel tool for studying human serotonergic neurotransmission in health and disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Lesch KP, Waider J . Serotonin in the modulation of neural plasticity and networks: implications for neurodevelopmental disorders. Neuron 2012; 76: 175–191.
Deneris ES, Wyler SC . Serotonergic transcriptional networks and potential importance to mental health. Nat Neurosci 2012; 15: 519–527.
Daubert EA, Condron BG . Serotonin: a regulator of neuronal morphology and circuitry. Trends Neurosci 2010; 33: 424–434.
Gaspar P, Cases O, Maroteaux L . The developmental role of serotonin: news from mouse molecular genetics. Nat Rev Neurosci 2003; 4: 1002–1012.
Benekareddy M, Vadodaria KC, Nair AR, Vaidya VA . Postnatal serotonin type 2 receptor blockade prevents the emergence of anxiety behavior, dysregulated stress-induced immediate early gene responses, and specific transcriptional changes that arise following early life stress. Biol Psychiatry 2011; 70: 1024–1032.
Vaidya VA, Vadodaria KC, Jha S . Neurotransmitter regulation of adult neurogenesis: putative therapeutic targets. CNS Neurol Disord Drug Targets 2007; 6: 358–374.
Lin SH, Lee LT, Yang YK . Serotonin and mental disorders: a concise review on molecular neuroimaging evidence. Clin Psychopharmacol Neurosci 2014; 12: 196–202.
Mosienko V, Beis D, Pasqualetti M, Waider J, Matthes S, Qadri F et al. Life without brain serotonin: reevaluation of serotonin function with mice deficient in brain serotonin synthesis. Behav Brain Res 2015; 277: 78–88.
Ming GL, Brustle O, Muotri A, Studer L, Wernig M, Christian KM . Cellular reprogramming: recent advances in modeling neurological diseases. J Neurosci 2011; 31: 16070–16075.
Gage FH, Temple S . Neural stem cells: generating and regenerating the brain. Neuron 2013; 80: 588–601.
Pang ZP, Yang N, Vierbuchen T, Ostermeier A, Fuentes DR, Yang TQ et al. Induction of human neuronal cells by defined transcription factors. Nature 2011; 476: 220–223.
Yang N, Ng YH, Pang ZP, Sudhof TC, Wernig M . Induced neuronal cells: how to make and define a neuron. Cell Stem Cell 2011; 9: 517–525.
Marchetto MC, Brennand KJ, Boyer LF, Gage FH . Induced pluripotent stem cells (iPSCs) and neurological disease modeling: progress and promises. Hum Mol Genet 2011; 20: R109–R115.
Nicholas CR, Chen J, Tang Y, Southwell DG, Chalmers N, Vogt D et al. Functional maturation of hPSC-derived forebrain interneurons requires an extended timeline and mimics human neural development. Cell Stem Cell 2013; 12: 573–586.
Maroof AM, Keros S, Tyson JA, Ying SW, Ganat YM, Merkle FT et al. Directed differentiation and functional maturation of cortical interneurons from human embryonic stem cells. Cell Stem Cell 2013; 12: 559–572.
Liu Y, Weick JP, Liu H, Krencik R, Zhang X, Ma L et al. Medial ganglionic eminence-like cells derived from human embryonic stem cells correct learning and memory deficits. Nat Biotechnol 2013; 31: 440–447.
Caiazzo M, Dell'Anno MT, Dvoretskova E, Lazarevic D, Taverna S, Leo D et al. Direct generation of functional dopaminergic neurons from mouse and human fibroblasts. Nature 2011; 476: 224–227.
Kriks S, Shim JW, Piao J, Ganat YM, Wakeman DR, Xie Z et al. Dopamine neurons derived from human ES cells efficiently engraft in animal models of Parkinson's disease. Nature 2011; 480: 547–551.
Ye W, Shimamura K, Rubenstein JL, Hynes MA, Rosenthal A . FGF and Shh signals control dopaminergic and serotonergic cell fate in the anterior neural plate. Cell 1998; 93: 755–766.
Kiyasova V, Gaspar P . Development of raphe serotonin neurons from specification to guidance. Eur J Neurosci 2011; 34: 1553–1562.
Craven SE, Lim KC, Ye W, Engel JD, de Sauvage F, Rosenthal A . Gata2 specifies serotonergic neurons downstream of sonic hedgehog. Development 2004; 131: 1165–1173.
Hendricks TJ, Fyodorov DV, Wegman LJ, Lelutiu NB, Pehek EA, Yamamoto B et al. Pet-1 ETS gene plays a critical role in 5-HT neuron development and is required for normal anxiety-like and aggressive behavior. Neuron 2003; 37: 233–247.
Zhao ZQ, Scott M, Chiechio S, Wang JS, Renner KJ, Gereau RWt et al. Lmx1b is required for maintenance of central serotonergic neurons and mice lacking central serotonergic system exhibit normal locomotor activity. J Neurosci 2006; 26: 12781–12788.
Alenina N, Bashammakh S, Bader M . Specification and differentiation of serotonergic neurons. Stem Cell Rev 2006; 2: 5–10.
Barberi T, Klivenyi P, Calingasan NY, Lee H, Kawamata H, Loonam K et al. Neural subtype specification of fertilization and nuclear transfer embryonic stem cells and application in parkinsonian mice. Nat Biotechnol 2003; 21: 1200–1207.
Nefzger CM, Haynes JM, Pouton CW . Directed expression of Gata2, Mash1, and Foxa2 synergize to induce the serotonergic neuron phenotype during in vitro differentiation of embryonic stem cells. Stem Cells 2011; 29: 928–939.
Lee SH, Lumelsky N, Studer L, Auerbach JM, McKay RD . Efficient generation of midbrain and hindbrain neurons from mouse embryonic stem cells. Nat Biotechnol 2000; 18: 675–679.
Shimada T, Takai Y, Shinohara K, Yamasaki A, Tominaga-Yoshino K, Ogura A et al. A simplified method to generate serotonergic neurons from mouse embryonic stem and induced pluripotent stem cells. J Neurochem 2012; 122: 81–93.
Dolmazon V, Alenina N, Markossian S, Mancip J, van de Vrede Y, Fontaine E et al. Forced expression of LIM homeodomain transcription factor 1b enhances differentiation of mouse embryonic stem cells into serotonergic neurons. Stem Cells Dev 2011; 20: 301–311.
Kumar M, Kaushalya SK, Gressens P, Maiti S, Mani S . Optimized derivation and functional characterization of 5-HT neurons from human embryonic stem cells. Stem Cells Dev 2009; 18: 615–627.
Tailor J, Kittappa R, Leto K, Gates M, Borel M, Paulsen O et al. Stem cells expanded from the human embryonic hindbrain stably retain regional specification and high neurogenic potency. J Neurosci 2013; 33: 12407–12422.
Malchenko S, Xie J, de Fatima Bonaldo M, Vanin EF, Bhattacharyya BJ, Belmadani A et al. Onset of rosette formation during spontaneous neural differentiation of hESC and hiPSC colonies. Gene 2014; 534: 400–407.
Kim KS . Converting human skin cells to neurons: a new tool to study and treat brain disorders? Cell Stem Cell 2011; 9: 179–181.
Pfisterer U, Wood J, Nihlberg K, Hallgren O, Bjermer L, Westergren-Thorsson G et al. Efficient induction of functional neurons from adult human fibroblasts. Cell Cycle 2011; 10: 3311–3316.
Najm FJ, Lager AM, Zaremba A, Wyatt K, Caprariello AV, Factor DC et al. Transcription factor-mediated reprogramming of fibroblasts to expandable, myelinogenic oligodendrocyte progenitor cells. Nat Biotechnol 2013; 31: 426–433.
Son EY, Ichida JK, Wainger BJ, Toma JS, Rafuse VF, Woolf CJ et al. Conversion of mouse and human fibroblasts into functional spinal motor neurons. Cell Stem Cell 2011; 9: 205–218.
Liu ML, Zang T, Zou Y, Chang JC, Gibson JR, Huber KM et al. Small molecules enable neurogenin 2 to efficiently convert human fibroblasts into cholinergic neurons. Nat Commun 2013; 4: 2183.
Boyer LF, Campbell B, Larkin S, Mu Y, Gage FH . Dopaminergic differentiation of human pluripotent cells. Curr Protoc Stem Cell Biol 2012; Chapter 1: Unit1H 6.
Ladewig J, Mertens J, Kesavan J, Doerr J, Poppe D, Glaue F et al. Small molecules enable highly efficient neuronal conversion of human fibroblasts. Nat Methods 2012; 9: 575–578.
Gentile MT, Nawa Y, Lunardi G, Florio T, Matsui H, Colucci-D'Amato L . Tryptophan hydroxylase 2 (TPH2) in a neuronal cell line: modulation by cell differentiation and NRSF/rest activity. J Neurochem 2012; 123: 963–970.
Marchetto MC, Carromeu C, Acab A, Yu D, Yeo GW, Mu Y et al. A model for neural development and treatment of Rett syndrome using human induced pluripotent stem cells. Cell 2010; 143: 527–539.
Vadodaria KC, Brakebusch C, Suter U, Jessberger S . Stage-specific functions of the small Rho GTPases Cdc42 and Rac1 for adult hippocampal neurogenesis. J Neurosci 2013; 33: 1179–1189.
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S et al STAR: ultrafast universal RNA-seq aligner Bioinformatics 2013; 29: 15–21.
Anders S, Huber W . Differential expression analysis for sequence count data. Genome Biol 2010; 11: R106.
Bardy C, van den Hurk M, Eames T, Marchand C, Hernandez RV et al. Neuronal medium that supports basic synaptic functions and activity of human neurons in vitro. Proc Natl Acad Sci U S A 2015; 112: E2725–E2734.
Chambers SM, Fasano CA, Papapetrou EP, Tomishima M, Sadelain M, Studer L . Highly efficient neural conversion of human ES and iPS cells by dual inhibition of SMAD signaling. Nat Biotechnol 2009; 27: 275–280.
Vierbuchen T, Ostermeier A, Pang ZP, Kokubu Y, Sudhof TC, Wernig M . Direct conversion of fibroblasts to functional neurons by defined factors. Nature 2010; 463: 1035–1041.
Ambasudhan R, Talantova M, Coleman R, Yuan X, Zhu S, Lipton SA et al. Direct reprogramming of adult human fibroblasts to functional neurons under defined conditions. Cell Stem Cell 2011; 9: 113–118.
Broccoli V, Caiazzo M, Dell'Anno MT . Setting a highway for converting skin into neurons. J Mol Cell Biol 2011; 3: 322–323.
Wapinski OL, Vierbuchen T, Qu K, Lee QY, Chanda S, Fuentes DR et al. Hierarchical mechanisms for direct reprogramming of fibroblasts to neurons. Cell 2013; 155: 621–635.
Spaethling JM, Piel D, Dueck H, Buckley PT, Morris JF, Fisher SA et al. Serotonergic neuron regulation informed by in vivo single-cell transcriptomics. FASEB J 2014; 28: 771–780.
Dougherty JD, Maloney SE, Wozniak DF, Rieger MA, Sonnenblick L, Coppola G et al. The disruption of Celf6, a gene identified by translational profiling of serotonergic neurons, results in autism-related behaviors. J Neurosci 2013; 33: 2732–2753.
Diaz SL, Doly S, Narboux-Neme N, Fernandez S, Mazot P, Banas SM et al. 5-HT(2B) receptors are required for serotonin-selective antidepressant actions. Mol Psychiatry 2012; 17: 154–163.
Richardson-Jones JW, Craige CP, Guiard BP, Stephen A, Metzger KL, Kung HF et al. 5-HT1A autoreceptor levels determine vulnerability to stress and response to antidepressants. Neuron 2010; 65: 40–52.
Studer L . Derivation of dopaminergic neurons from pluripotent stem cells. Prog Brain Res 2012; 200: 243–263.
Sanchez-Pernaute R, Studer L, Bankiewicz KS, Major EO, McKay RD . In vitro generation and transplantation of precursor-derived human dopamine neurons. J Neurosci Res 2001; 65: 284–288.
Pfisterer U, Kirkeby A, Torper O, Wood J, Nelander J, Dufour A et al. Direct conversion of human fibroblasts to dopaminergic neurons. Proc Natl Acad Sci USA 2011; 108: 10343–10348.
Liu X, Li F, Stubblefield EA, Blanchard B, Richards TL, Larson GA et al. Direct reprogramming of human fibroblasts into dopaminergic neuron-like cells. Cell Res 2012; 22: 321–332.
Cheng L, Chen CL, Luo P, Tan M, Qiu M, Johnson R et al. Lmx1b, Pet-1, and Nkx2.2 coordinately specify serotonergic neurotransmitter phenotype. J Neurosci 2003; 23: 9961–9967.
Ding YQ, Marklund U, Yuan W, Yin J, Wegman L, Ericson J et al. Lmx1b is essential for the development of serotonergic neurons. Nat Neurosci 2003; 6: 933–938.
Scott MM, Krueger KC, Deneris ES . A differentially autoregulated Pet-1 enhancer region is a critical target of the transcriptional cascade that governs serotonin neuron development. J Neurosci 2005; 25: 2628–2636.
Krueger KC, Deneris ES . Serotonergic transcription of human FEV reveals direct GATA factor interactions and fate of Pet-1-deficient serotonin neuron precursors. J Neurosci 2008; 28: 12748–12758.
Marinelli S, Schnell SA, Hack SP, Christie MJ, Wessendorf MW, Vaughan CW . Serotonergic and nonserotonergic dorsal raphe neurons are pharmacologically and electrophysiologically heterogeneous. J Neurophysiol 2004; 92: 3532–3537.
Hajos M, Gartside SE, Villa AE, Sharp T . Evidence for a repetitive (burst) firing pattern in a sub-population of 5-hydroxytryptamine neurons in the dorsal and median raphe nuclei of the rat. Neuroscience 1995; 69: 189–197.
Allers KA, Sharp T . Neurochemical and anatomical identification of fast- and slow-firing neurones in the rat dorsal raphe nucleus using juxtacellular labelling methods in vivo. Neuroscience 2003; 122: 193–204.
Kocsis B, Varga V, Dahan L, Sik A . Serotonergic neuron diversity: identification of raphe neurons with discharges time-locked to the hippocampal theta rhythm. Proc Natl Acad Sci USA 2006; 103: 1059–1064.
Varga V, Szekely AD, Csillag A, Sharp T, Hajos M . Evidence for a role of GABA interneurones in the cortical modulation of midbrain 5-hydroxytryptamine neurones. Neuroscience 2001; 106: 783–792.
Albert PR, Benkelfat C . The neurobiology of depression—revisiting the serotonin hypothesis. II. Genetic, epigenetic and clinical studies. Philos Trans R Soc Lond B Biol Sci 2013; 368: 20120535.
Charney DS . Monoamine dysfunction and the pathophysiology and treatment of depression. J Clin Psychiatry 1998; 59: 11–14.
Meltzer HY . The role of serotonin in antipsychotic drug action. Neuropsychopharmacology 1999; 21: 106S–115S.
Nagayasu K, Kitaichi M, Nishitani N, Asaoka N, Shirakawa H, Nakagawa T et al. Chronic effects of antidepressants on serotonin release in rat raphe slice cultures: high potency of milnacipran in the augmentation of serotonin release. Int J Neuropsychopharmacol 2013; 16: 2295–2306.
Komlosi G, Molnar G, Rozsa M, Olah S, Barzo P, Tamas G . Fluoxetine (prozac) and serotonin act on excitatory synaptic transmission to suppress single layer 2/3 pyramidal neuron-triggered cell assemblies in the human prefrontal cortex. J Neurosci 2012; 32: 16369–16378.
Fernstrom JD . Role of precursor availability in control of monoamine biosynthesis in brain. Physiol Rev 1983; 63: 484–546.
Acknowledgements
KCV was supported by the Swiss National Science Foundation (SNSF) outgoing postdoctoral fellowship. JM and CB were supported by fellowships Glenn Center for Aging Research and the FP7 Marie Curie, respectively. This research was supported by Lynn and Edward Streim, the Robert and Mary Jane Engman Foundation, the JPB Foundation, and the Leona M and Harry B Helmsley Charitable Trust grant #2012-PG-MED002. We thank Dr Juergen Winkler for a line of primary human fibroblasts, Dr Manching Ku for help with RNA sequencing, Jamie Simon for help with illustrations, Dr Himanish Ghosh for help with semi-automated image analysis and Mary Lynn Gage for editorial comments.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Molecular Psychiatry website
Supplementary information
Rights and permissions
About this article
Cite this article
Vadodaria, K., Mertens, J., Paquola, A. et al. Generation of functional human serotonergic neurons from fibroblasts. Mol Psychiatry 21, 49–61 (2016). https://doi.org/10.1038/mp.2015.161
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mp.2015.161
This article is cited by
-
Transcriptomic profiling across human serotonin neuron differentiation via the FEV reporter system
Stem Cell Research & Therapy (2024)
-
Screens in aging-relevant human ALS-motor neurons identify MAP4Ks as therapeutic targets for the disease
Cell Death & Disease (2024)
-
Molecular pathways of major depressive disorder converge on the synapse
Molecular Psychiatry (2023)
-
Induced neural progenitor cells and iPS-neurons from major depressive disorder patients show altered bioenergetics and electrophysiological properties
Molecular Psychiatry (2022)
-
Sensing serotonin secreted from human serotonergic neurons using aptamer-modified nanopipettes
Molecular Psychiatry (2021)