Abstract
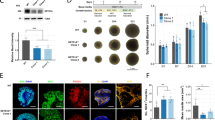
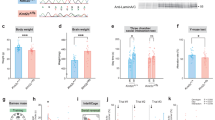
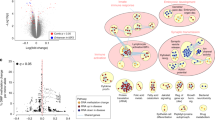
The basic helix-loop-helix PAS (Per, Arnt, Sim) domain transcription factor gene NPAS3 is a replicated genetic risk factor for psychiatric disorders. A knockout (KO) mouse model exhibits behavioral and adult neurogenesis deficits consistent with human illness. To define the location and mechanism of NPAS3 etiopathology, we combined immunofluorescent, transcriptomic and metabonomic approaches. Intense Npas3 immunoreactivity was observed in the hippocampal subgranular zone—the site of adult neurogenesis—but was restricted to maturing, rather than proliferating, neuronal precursor cells. Microarray analysis of a HEK293 cell line over-expressing NPAS3 showed that transcriptional targets varied according to circadian rhythm context and C-terminal deletion. The most highly up-regulated NPAS3 target gene, VGF, encodes secretory peptides with established roles in neurogenesis, depression and schizophrenia. VGF was just one of many NPAS3 target genes also regulated by the SOX family of transcription factors, suggesting an overlap in neurodevelopmental function. The parallel repression of multiple glycolysis genes by NPAS3 reveals a second role in the regulation of glucose metabolism. Comparison of wild-type and Npas3 KO metabolite composition using high-resolution mass spectrometry confirmed these transcriptional findings. KO brain tissue contained significantly altered levels of NAD+, glycolysis metabolites (such as dihydroxyacetone phosphate and fructose-1,6-bisphosphate), pentose phosphate pathway components and Kreb's cycle intermediates (succinate and α-ketoglutarate). The dual neurodevelopmental and metabolic aspects of NPAS3 activity described here increase our understanding of mental illness etiology, and may provide a mechanism for innate and medication-induced susceptibility to diabetes commonly reported in psychiatric patients.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Kamnasaran D, Muir WJ, Ferguson-Smith MA, Cox DW . Disruption of the neuronal PAS3 gene in a family affected with schizophrenia. J Med Genet 2003; 40: 325–332.
Pickard BS, Malloy MP, Porteous DJ, Blackwood DH, Muir WJ . Disruption of a brain transcription factor, NPAS3, is associated with schizophrenia and learning disability. Am J Med Genet B Neuropsychiatr Genet 2005; 136: 26–32.
Pickard BS, Christoforou A, Thomson PA, Fawkes A, Evans KL, Morris SW et al. Interacting haplotypes at the NPAS3 locus alter risk of schizophrenia and bipolar disorder. Mol Psychiatry 2009; 14: 874–884.
Huang J, Perlis RH, Lee PH, Rush AJ, Fava M, Sachs GS et al. Cross-disorder genomewide analysis of schizophrenia, bipolar disorder, and depression. Am J Psychiatry 2010; 167: 1254–1263.
Ferreira MA, O’Donovan MC, Meng YA, Jones IR, Ruderfer DM, Jones L et al. Collaborative genome-wide association analysis supports a role for ANK3 and CACNA1C in bipolar disorder. Nat Genet 2008; 40: 1056–1058.
Macintyre G, Alford T, Xiong L, Rouleau GA, Tibbo PG, Cox DW . Association of NPAS3 exonic variation with schizophrenia. Schizophr Res 2010; 120: 143–149.
Lavedan C, Licamele L, Volpi S, Hamilton J, Heaton C, Mack K et al. Association of the NPAS3 gene and five other loci with response to the antipsychotic iloperidone identified in a whole genome association study. Mol Psychiatry 2009; 14: 804–819.
Brunskill EW, Witte DP, Shreiner AB, Potter SS . Characterization of npas3, a novel basic helix-loop-helix PAS gene expressed in the developing mouse nervous system. Mech Dev 1999; 88: 237–241.
Gilles-Gonzalez MA, Gonzalez G . Signal transduction by heme-containing PAS-domain proteins. J Appl Physiol 2004; 96: 774–783.
Brunskill EW, Ehrman LA, Williams MT, Klanke J, Hammer D, Schaefer TL et al. Abnormal neurodevelopment, neurosignaling and behaviour in Npas3-deficient mice. Eur J Neurosci 2005; 22: 1265–1276.
Erbel-Sieler C, Dudley C, Zhou Y, Wu X, Estill SJ, Han T et al. Behavioral and regulatory abnormalities in mice deficient in the NPAS1 and NPAS3 transcription factors. Proc Natl Acad Sci USA 2004; 101: 13648–13653.
Pieper AA, Wu X, Han TW, Estill SJ, Dang Q, Wu LC et al. The neuronal PAS domain protein 3 transcription factor controls FGF-mediated adult hippocampal neurogenesis in mice. Proc Natl Acad Sci USA 2005; 102: 14052–14057.
Kempermann G, Krebs J, Fabel K . The contribution of failing adult hippocampal neurogenesis to psychiatric disorders. Curr Opin Psychiatry 2008; 21: 290–295.
Reif A, Schmitt A, Fritzen S, Lesch KP . Neurogenesis and schizophrenia: dividing neurons in a divided mind? Eur Arch Psychiatry Clin Neurosci 2007; 257: 290–299.
Pieper AA, Xie S, Capota E, Estill SJ, Zhong J, Long JM et al. Discovery of a proneurogenic, neuroprotective chemical. Cell 2010; 142: 39–51.
Azim E, Jabaudon D, Fame RM, Macklis JD . SOX6 controls dorsal progenitor identity and interneuron diversity during neocortical development. Nat Neurosci 2009; 12: 1238–1247.
Haslinger A, Schwarz TJ, Covic M, Chichung Lie D . Expression of Sox11 in adult neurogenic niches suggests a stage-specific role in adult neurogenesis. Eur J Neurosci 2009; 29: 2103–2114.
Lefebvre V . The SoxD transcription factors -- Sox5, Sox6, and Sox13 -- are key cell fate modulators. Int J Biochem Cell Biol 2010; 42: 429–432.
Reick M, Garcia JA, Dudley C, McKnight SL . NPAS2: an analog of clock operative in the mammalian forebrain. Science 2001; 293: 506–509.
Knight HM, Pickard BS, Maclean A, Malloy MP, Soares DC, McRae AF et al. A cytogenetic abnormality and rare coding variants identify ABCA13 as a candidate gene in schizophrenia, bipolar disorder, and depression. Am J Hum Genet 2009; 85: 833–846.
Balsalobre A, Damiola F, Schibler U . A serum shock induces circadian gene expression in mammalian tissue culture cells. Cell 1998; 93: 929–937.
Huang TS, Grodeland G, Sleire L, Wang MY, Kvalheim G, Laerum OD . Induction of circadian rhythm in cultured human mesenchymal stem cells by serum shock and cAMP analogs in vitro. Chronobiol Int 2009; 26: 242–257.
Wu H, Southam AD, Hines A, Viant MR . High-throughput tissue extraction protocol for NMR- and MS-based metabolomics. Anal Biochem 2008; 372: 204–212.
Reisz-Porszasz S, Probst MR, Fukunaga BN, Hankinson O . Identification of functional domains of the aryl hydrocarbon receptor nuclear translocator protein (ARNT). Mol Cell Biol 1994; 14: 6075–6086.
Nogales-Cadenas R, Carmona-Saez P, Vazquez M, Vicente C, Yang X, Tirado F et al. GeneCodis: interpreting gene lists through enrichment analysis and integration of diverse biological information. Nucleic Acids Res 2009; 37 (Web Server issue): W317–W322.
Axell MZ, Zlateva S, Curtis M . A method for rapid derivation and propagation of neural progenitors from human embryonic stem cells. J Neurosci Methods 2009; 184: 275–284.
Benita Y, Kikuchi H, Smith AD, Zhang MQ, Chung DC, Xavier RJ . An integrative genomics approach identifies hypoxia inducible factor-1 (HIF-1)-target genes that form the core response to hypoxia. Nucleic Acids Res 2009; 37: 4587–4602.
Chi JT, Wang Z, Nuyten DS, Rodriguez EH, Schaner ME, Salim A et al. Gene expression programs in response to hypoxia: cell type specificity and prognostic significance in human cancers. PLoS Med 2006; 3: e47.
Greijer AE, van der Groep P, Kemming D, Shvarts A, Semenza GL, Meijer GA et al. Up-regulation of gene expression by hypoxia is mediated predominantly by hypoxia-inducible factor 1 (HIF-1). J Pathol 2005; 206: 291–304.
Mense SM, Sengupta A, Zhou M, Lan C, Bentsman G, Volsky DJ et al. Gene expression profiling reveals the profound upregulation of hypoxia-responsive genes in primary human astrocytes. Physiol Genomics 2006; 25: 435–449.
Takahashi K, Yamanaka S . Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 2006; 126: 663–676.
Lefebvre V . The SoxD transcription factors—Sox5, Sox6, and Sox13—are key cell fate modulators. Int J Biochem Cell Biol 2010; 42: 429–432.
Brown JP, Couillard-Despres S, Cooper-Kuhn CM, Winkler J, Aigner L, Kuhn HG . Transient expression of doublecortin during adult neurogenesis. J Comp Neurol 2003; 467: 1–10.
Cirelli C, Tononi G . Gene expression in the brain across the sleep-waking cycle. Brain Res 2000; 885: 303–321.
Bartolomucci A, Possenti R, Levi A, Pavone F, Moles A . The role of the vgf gene and VGF-derived peptides in nutrition and metabolism. Genes Nutr 2007; 2: 169–180.
Jethwa PH, Ebling FJ . Role of VGF-derived peptides in the control of food intake, body weight and reproduction. Neuroendocrinology 2008; 88: 80–87.
Thakker-Varia S, Krol JJ, Nettleton J, Bilimoria PM, Bangasser DA, Shors TJ et al. The neuropeptide VGF produces antidepressant-like behavioral effects and enhances proliferation in the hippocampus. J Neurosci 2007; 27: 12156–12167.
Pasinetti GM, Ungar LH, Lange DJ, Yemul S, Deng H, Yuan X et al. Identification of potential CSF biomarkers in ALS. Neurology 2006; 66: 1218–1222.
Selle H, Lamerz J, Buerger K, Dessauer A, Hager K, Hampel H et al. Identification of novel biomarker candidates by differential peptidomics analysis of cerebrospinal fluid in Alzheimer's disease. Comb Chem High Throughput Screen 2005; 8: 801–806.
Ruetschi U, Zetterberg H, Podust VN, Gottfries J, Li S, Hviid Simonsen A et al. Identification of CSF biomarkers for frontotemporal dementia using SELDI-TOF. Exp Neurol 2005; 196: 273–281.
Altar CA, Vawter MP, Ginsberg SD . Target identification for CNS diseases by transcriptional profiling. Neuropsychopharmacology 2009; 34: 18–54.
Malberg JE, Monteggia LM . VGF, a new player in antidepressant action? Sci Signal 2008; 1: pe19.
Huang JT, Leweke FM, Oxley D, Wang L, Harris N, Koethe D et al. Disease biomarkers in cerebrospinal fluid of patients with first-onset psychosis. PLoS Med 2006; 3: e428.
Huang JT, Leweke FM, Tsang TM, Koethe D, Kranaster L, Gerth CW et al. CSF metabolic and proteomic profiles in patients prodromal for psychosis. PLoS One 2007; 2: e756.
Rutter J, Reick M, McKnight SL . Metabolism and the control of circadian rhythms. Annu Rev Biochem 2002; 71: 307–331.
Rutter J, Reick M, Wu LC, McKnight SL . Regulation of clock and NPAS2 DNA binding by the redox state of NAD cofactors. Science 2001; 293: 510–514.
Lamont EW, Legault-Coutu D, Cermakian N, Boivin DB . The role of circadian clock genes in mental disorders. Dialogues Clin Neurosci 2007; 9: 333–342.
Ma D, Chan MK, Lockstone HE, Pietsch SR, Jones DN, Cilia J et al. Antipsychotic treatment alters protein expression associated with presynaptic function and nervous system development in rat frontal cortex. J Proteome Res 2009; 8: 3284–3297.
McQuillin A, Rizig M, Gurling HM . A microarray gene expression study of the molecular pharmacology of lithium carbonate on mouse brain mRNA to understand the neurobiology of mood stabilization and treatment of bipolar affective disorder. Pharmacogenet Genomics 2007; 17: 605–617.
Fernandez-Egea E, Bernardo M, Donner T, Conget I, Parellada E, Justicia A et al. Metabolic profile of antipsychotic-naive individuals with non-affective psychosis. Br J Psychiatry 2009; 194: 434–438.
Herberth M, Koethe D, Cheng TM, Krzyszton ND, Schoeffmann S, Guest PC et al. Impaired glycolytic response in peripheral blood mononuclear cells of first-onset antipsychotic-naive schizophrenia patients. Mol Psychiatry 2010 (e-pub ahead of print).
Martins-de-Souza D, Gattaz WF, Schmitt A, Maccarrone G, Hunyadi-Gulyas E, Eberlin MN et al. Proteomic analysis of dorsolateral prefrontal cortex indicates the involvement of cytoskeleton, oligodendrocyte, energy metabolism and new potential markers in schizophrenia. J Psychiatr Res 2009; 43: 978–986.
Do KQ, Cabungcal JH, Frank A, Steullet P, Cuenod M . Redox dysregulation, neurodevelopment, and schizophrenia. Curr Opin Neurobiol 2009; 19: 220–230.
Acknowledgements
We acknowledge Royal Society of Physicians in Edinburgh Sim Fellowship to BSP and a China Council Studentship to LS. SJC was supported in part by a NARSAD Young Investigator Award.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Molecular Psychiatry website
Supplementary information
Rights and permissions
About this article
Cite this article
Sha, L., MacIntyre, L., Machell, J. et al. Transcriptional regulation of neurodevelopmental and metabolic pathways by NPAS3. Mol Psychiatry 17, 267–279 (2012). https://doi.org/10.1038/mp.2011.73
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mp.2011.73
Keywords
This article is cited by
-
METTL3 exerts synergistic effects on m6A methylation and histone modification to regulate the function of VGF in lung adenocarcinoma
Clinical Epigenetics (2023)
-
Investigation of genetic loci shared between bipolar disorder and risk-taking propensity: potential implications for pharmacological interventions
Neuropsychopharmacology (2021)
-
Angiogenic gene networks are dysregulated in opioid use disorder: evidence from multi-omics and imaging of postmortem human brain
Molecular Psychiatry (2021)
-
Transcriptomic analysis links diverse hypothalamic cell types to fibroblast growth factor 1-induced sustained diabetes remission
Nature Communications (2020)
-
Pathogenic variants in E3 ubiquitin ligase RLIM/RNF12 lead to a syndromic X-linked intellectual disability and behavior disorder
Molecular Psychiatry (2019)