Abstract
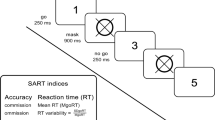
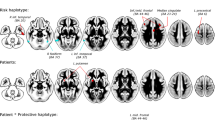
Although therapeutic interventions in attention-deficit/hyperactivity disorder (ADHD) still focus on the dopaminergic system, recent studies indicate a serotonergic dysfunction in this disease as well. In that respect, several variants of the tryptophan hydroxylase gene (TPH2), which codes for the rate-limiting enzyme in the biosynthesis of serotonin (5-HT), have been associated with ADHD. The rs4570625 G-allele polymorphisms of the TPH2 gene have already been related to altered reactivity of the brain during perception tasks with emotional stimuli in healthy adults. Here we investigated the influence of the ADHD related risk alleles for rs4570625 and for rs11178997 on prefrontal brain function during cognitive response control in large samples of adult ADHD patients (n=124) and healthy controls (n=84). Response control was elicited with a Go-NoGo task (continuous performance test; CPT) performed during recording of an ongoing EEG. From the resulting event-related potentials in the Go- and NoGo conditions of the CPT, the NoGo-anteriorization (NGA) has been calculated as a valid neurophysiological parameter for prefrontal brain function. In the current study, ADHD risk alleles of both polymorphisms were found to be associated with a reduction in the NGA in both healthy controls and ADHD patients. These findings are in line with the notion that genetic variations associated with altered serotonergic neurotransmission are also associated with the function of the prefrontal cortex during response inhibition. This mechanism might also be relevant in the pathophysiology of ADHD.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Brookes K, Xu X, Chen W, Zhou K, Neale B, Lowe N et al. The analysis of 51 genes in DSM-IV combined type attention deficit hyperactivity disorder: association signals in DRD4, DAT1 and 16 other genes. Mol Psychiatry 2006; 11: 934–953.
Heiser P, Friedel S, Dempfle A, Konrad K, Smidt J, Grabarkiewicz J et al. Molecular genetic aspects of attention-deficit/hyperactivity disorder. Neurosci Biobehav Rev 2004; 28: 625–641.
Walther DJ, Bader M . A unique central tryptophan hydroxylase isoform. Biochem Pharmacol 2003; 66: 1673–1680.
Cote F, Thevenot E, Fligny C, Fromes Y, Darmon M, Ripoche MA et al. Disruption of the nonneuronal tph1 gene demonstrates the importance of peripheral serotonin in cardiac function. Proc Natl Acad Sci USA 2003; 100: 13525–13530.
Sheehan K, Lowe N, Kirley A, Mullins C, Fitzgerald M, Gill M et al. Tryptophan hydroxylase 2 (TPH2) gene variants associated with ADHD. Mol Psychiatry 2005; 10: 944–949.
Walitza S, Renner TJ, Dempfle A, Konrad K, Wewetzer Ch, Halbach A et al. Transmission disequilibrium of polymorphic variants in the tryptophan hydroxylase-2 gene in attention-deficit/hyperactivity disorder. Mol Psychiatry 2005; 10: 1126–1132.
Canli T, Congdon E, Gutknecht L, Constable RT, Lesch KP . Amygdala responsiveness is modulated by tryptophan hydroxylase-2 gene variation. J Neural Transm 2005; 112: 1479–1485.
Herrmann MJ, Huter T, Muller F, Muhlberger A, Pauli P, Reif A et al. Additive effects of serotonin transporter and tryptophan hydroxylase-2 gene variation on emotional processing. Cereb Cortex 2007; 17: 1160–1163.
Gottesman II, Gould TD . The endophenotype concept in psychiatry: etymology and strategic intentions. Am J Psychiatry 2003; 160: 636–645.
Hariri AR, Weinberger DR . Imaging genomics. Br Med Bull 2003; 65: 259–270.
Lijffijt M, Kenemans JL, Verbaten MN, van Engeland H . A meta-analytic review of stopping performance in attention-deficit/hyperactivity disorder: deficient inhibitory motor control? J Abnorm Psychol 2005; 114: 216–222.
Aron AR, Poldrack RA . The cognitive neuroscience of response inhibition: relevance for genetic research in attention-deficit/hyperactivity disorder. Biol Psychiatry 2005; 57: 1285–1292.
Crean J, Richards JB, de Wit H . Effect of tryptophan depletion on impulsive behavior in men with or without a family history of alcoholism. Behav Brain Res 2002; 136: 349–357.
Walderhaug E, Lunde H, Nordvik JE, Landro NI, Refsum H, Magnusson A . Lowering of serotonin by rapid tryptophan depletion increases impulsiveness in normal individuals. Psychopharmacology (Berl) 2002; 164: 385–391.
Bjork JM, Moeller FG, Dougherty DM, Swann AC, Machado MA, Hanis CL . Serotonin 2a receptor T102C polymorphism and impaired impulse control. Am J Med Genet 2002; 114: 336–339.
Stoltenberg SF, Glass JM, Chermack ST, Flynn HA, Li S, Weston ME et al. Possible association between response inhibition and a variant in the brain-expressed tryptophan hydroxylase-2 gene. Psychiatr Genet 2006; 16: 35–38.
LeMarquand DG, Pihl RO, Young SN, Tremblay RE, Seguin JR, Palmour RM et al. Tryptophan depletion, executive functions, and disinhibition in aggressive, adolescent males. Neuropsychopharmacology 1998; 19: 333–341.
LeMarquand DG, Benkelfat C, Pihl RO, Palmour RM, Young SN . Behavioral disinhibition induced by tryptophan depletion in nonalcoholic young men with multigenerational family histories of paternal alcoholism. Am J Psychiatry 1999; 156: 1771–1779.
Fallgatter AJ, Jatzke S, Bartsch AJ, Hamelbeck B, Lesch KP . Serotonin transporter promoter polymorphism influences topography of inhibitory motor control. Int J Neuropsychopharmacol 1999; 2: 115–120.
Kaiser S, Unger J, Kiefer M, Markela J, Mundt C, Weisbrod M . Executive control deficit in depression: event-related potentials in a Go/Nogo task. Psychiatry Res 2003; 122: 169–184.
Rubia K, Lee F, Cleare AJ, Tunstall N, Fu CH, Brammer M et al. Tryptophan depletion reduces right inferior prefrontal activation during response inhibition in fast, event-related fMRI. Psychopharmacology (Berl) 2005; 179: 791–803.
Fallgatter AJ, Herrmann MJ, Roemmler J, Ehlis AC, Wagener A, Heidrich A et al. Allelic variation of serotonin transporter function modulates the brain electrical response for error processing. Neuropsychopharmacology 2004; 29: 1506–1511.
Fallgatter AJ, Herrmann MJ, Hohoff C, Ehlis AC, Jarczok TA, Freitag CM et al. DTNBP1 (dysbindin) gene variants modulate prefrontal brain function in healthy individuals. Neuropsychopharmacology 2006; 31: 2002–2010.
Fallgatter AJ, Brandeis D, Strik WK . A robust assessment of the NoGo-anteriorisation of P300 microstates in a cued Continuous Performance Test. Brain Topogr 1997; 9: 295–302.
Pascual Marqui RD, Michel CM, Lehmann D . Low resolution electromagnetic tomography: a new method for localizing electrical activity in the brain. Int J Psychophysiol 1994; 18: 49–65.
Strik WK, Fallgatter AJ, Brandeis D, Pascual Marqui RD . Three-dimensional tomography of event-related potentials during response inhibition: evidence for phasic frontal lobe activation. Electroencephalogr Clin Neurophysiol 1998; 108: 406–413.
Fallgatter AJ, Bartsch AJ, Herrmann MJ . Electrophysiological measurements of anterior cingulate function. J Neural Transm 2002; 109: 977–988.
Fallgatter AJ, Ehlis AC, Rosler M, Strik WK, Blocher D, Herrmann MJ . Diminished prefrontal brain function in adults with psychopathology in childhood related to attention deficit hyperactivity disorder. Psychiatry Res 2005; 138: 157–169.
Fallgatter AJ, Ehlis AC, Seifert J, Strik WK, Scheuerpflug P, Zillessen KE et al. Altered response control and anterior cingulate function in attention-deficit/hyperactivity disorder boys. Clin Neurophysiol 2004; 115: 973–981.
Wittchen H-U, Wunderlich U, Gruschwitz S, Zaudig M . SKID I. Strukturiertes Klinisches Interview fur DSM-IV. Achse I: Psychische Storungen. Interviewheft und Beurteilungsheft. Eine deutschsprachige, erweiterte Bearbeitung der amerikanischen Originalversion des SCID I. Hogrefe: Göttingen, 1997.
Fydrich T, Renneberg B, Schmitz B, Wittchen H-U . SKID II. Interviewheft. Strukturiertes Klinisches Interview fur DSM-IV. Achse II (Personlichkeitsstorungen). Eine deutschsprachige, erweiterte Bearbeitung der amerikanischen Originalversion des SCID-II von: First, MB, Spitzer, RL, Gibbon, M, Williams, JBW, Benjamin, L (Version 3/96). Hogrefe: Göttingen, 1997.
Gutknecht L, Jacob C, Strobel A, Kriegebaum C, Müller J, Zeng Y et al. Tryptophan hydroxylase-2 gene variation influences personality traits and disorders related to emotional dysregulation. Int J Neuropsychopharmacol 2007; 10: 309–320.
Gratton G, Coles MG, Donchin E . A new method for off-line removal of ocular artifact. Electroencephalogr Clin Neurophysiol 1983; 55: 468–484.
Lehmann D . Principles of spatial analysis. In: Gevins A, Remond A (eds). Handbook of Electroencephalography and Clinical Neurophysiology. Elsevier: Amsterdam, 1987, pp 309–354.
Bobb AJ, Castellanos FX, Addington AM, Rapoport JL . Molecular genetic studies of ADHD: 1991 to 2004. Am J Med Genet B Neuropsychiatr Genet 2005; 132: 109–125.
Acknowledgements
This study was supported by the Deutsche Forschungsgemeinschaft (KFO 125/1-1).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Baehne, C., Ehlis, AC., Plichta, M. et al. Tph2 gene variants modulate response control processes in adult ADHD patients and healthy individuals. Mol Psychiatry 14, 1032–1039 (2009). https://doi.org/10.1038/mp.2008.39
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mp.2008.39
Keywords
This article is cited by
-
Imaging genetics in attention-deficit/hyperactivity disorder and related neurodevelopmental domains: state of the art
Brain Imaging and Behavior (2017)
-
Attention-deficit hyperactivity disorder in adults: A systematic review and meta-analysis of genetic, pharmacogenetic and biochemical studies
Molecular Psychiatry (2016)
-
Serotonergic modulation of ‘waiting impulsivity’ is mediated by the impulsivity phenotype in humans
Translational Psychiatry (2016)
-
Imaging genetics in adult attention-deficit/hyperactivity disorder (ADHD): a way towards pathophysiological understanding?
Borderline Personality Disorder and Emotion Dysregulation (2014)
-
Stimulation of 5-HT2C Receptors Improves Cognitive Deficits Induced by Human Tryptophan Hydroxylase 2 Loss of Function Mutation
Neuropsychopharmacology (2014)