Abstract
Several lines of evidence indicate that cytomegalovirus infection can be substantially associated with onset of inflammatory bowel disease, especially in patients refractory to immunosuppressive treatment. As cytomegalovirus is widely spread in the population, here we present a quantitative detection system suitable to differentiate clinically relevant cytomegalovirus infection from common latent cytomegalovirus. Using a quantitative real-time PCR approach, cytomegalovirus viral load was evaluated in 917 formalin-fixed and paraffin-embedded colon biopsy samples of 136 patients diagnosed with inflammatory bowel disease. Besides initial cytomegalovirus testing, the PCR system was also used to monitor therapy response after antiviral treatment. Cytomegalovirus DNA was detected in 37 patients (27%) with varying viral loads ranging from 5 to 8.7 × 105 copies/105 cells. Thereof, 13 patients (35%) received an antiviral treatment with 12 of them going into remission (92%). Later, five patients displayed a relapse and three patients who agreed to restart antiviral treatment again showed positive therapy response. A retrospective comparison of viral loads with antiviral therapy response revealed a threshold of 600 cytomegalovirus copies/105 cells as indicative for clinically relevant infection. Of note, sensitivity of cytomegalovirus detection by immunohistochemistry was found to be insufficient to reliably identify antiviral therapy responders. In conclusion, quantitative real-time PCR using formalin-fixed biopsy samples is suitable for detection of cytomegalovirus infection in tissue samples of patients with inflammatory bowel disease. Moreover, it allows the definition of a viral load threshold, predictive for clinical relevance concerning antiviral therapy response.
Similar content being viewed by others
Main
Infections with cytomegalovirus are quite common in humans. Up to 70% of adult people exhibit signs of past infection.1 In immunocompetent hosts, primary cytomegalovirus infections usually remain asymptomatic, while in stem cell and solid organ transplant recipients, infection or reactivation is a serious cause of mortality and morbidity.2 Moreover, several reports indicate that cytomegalovirus infection can be significantly associated with the manifestation of inflammatory bowel disease such as ulcerative colitis and Crohn’s disease.3, 4, 5 Especially in patients refractory to standard anti-inflammatory/immunosuppressive treatment an underlying cytomegalovirus infection should be taken into account.3, 6 However, whether cytomegalovirus exacerbates the disease or just represents a bystander of a severe course of disease is currently unknown.7 Interestingly, extent of cytomegalovirus infections is apparently different in inflammatory bowel disease subtypes with cytomegalovirus, showing a minor association with the onset of Crohn’s disease compared with ulcerative colitis.2
Several methods for analyzing cytomegalovirus infection in tissue are available. Histological diagnosis via detection of inclusion bodies (owl’s eyes) using hematoxylin and eosin-stained tissue sections is considered as the ‘golden standard’.8 Although highly specific, this method lacks sufficient sensitivity and requires a high level of diagnostic experience.2, 8 Additional methods employ immunological cytomegalovirus detection such as immunohistochemistry and the antigenemia assay where the latter one is usually performed with serum samples. More recently, PCR-based cytomegalovirus detection was also applied to evaluate cytomegalovirus infection in tissue of inflammatory bowel disease patients.9 Although highly sensitive, owing to the common cytomegalovirus latency conventional PCR may give rise to the risk of false positive detection, in turn showing limited specificity.7 Therefore, the application of quantitative real-time PCR analysis is increasingly used in order to estimate impact of cytomegalovirus infection.10, 11, 12 For instance, a report published by Roblin et al,13 suggested a cutoff value of >250 cytomegalovirus copies/mg tissue as predictive for resistance to immunosuppressive therapy in patients suffering from ulcerative colitis. Because the study is based on the analysis of fresh frozen sample material, this method is rather inappropriate for day-to-day diagnostics of most outpatient clinics. Moreover, for quantification of intracellular viruses such as cytomegalovirus, normalization of viral loads to total cell number is considered to be more appropriated.14
In the present study, we describe a real-time PCR assay suitable for the sensitive detection and quantification of cytomegalovirus in formalin-fixed and paraffin-embedded colonic biopsy samples. Here normalization of viral loads is referred to cell number, which is determined via a second independent quantitative PCR targeting a human single copy gene. After analyzing 917 biopsy samples derived from 136 inflammatory bowel disease patients, retrospective comparison of viral loads to antiviral therapy response revealed a certain cutoff value predictive for clinically relevant cytomegalovirus infections.
Materials and methods
Patients and samples
Biopsy samples of 136 inflammatory bowel disease patients diagnosed at the Institute of Pathology Vechta and the Department of Gastroenterology of the Hospital St Marien in Vechta in cooperation with the Gastroenterology section of the Private Practice of Internal Medicine in Oldenburg. All samples were captured using standard endoscopic procedures. Diagnostics were based on endoscopic findings and histological evaluations performed by experienced gastroenterologists and pathologists, respectively. At the histological level, the severity of inflammation was determined according to the grading system published by Gupta et al15 and scored as follows: (0) inactive/absent, (1) mild, (2) moderate, and (3) severe, using the histological criteria described. Of the 136 patients, 100 patients had a single case evaluation while 36 patients were monitored more often (at different time points) with a maximum of 6 follow-up analyses. Together, this generates a total of 225 patient evaluations. For the majority of patient evaluations, several consecutive tissue sections (biopsy samples) were analyzed leading to a total of 917 colon biopsy samples enclosed in the study. All samples were formalin fixed (4%) and paraffin embedded.
Histopathology and immunohistochemistry
Hematoxylin and eosin staining was performed using the Tissue-Tek Film staining machine according to the manufacturer’s instructions (Sakura Finetek GmbH, Staufen, Germany). Immunohistochemical detection of cytomegalovirus was performed with the Benchmark Ultra system (Roche diagnostics, Mannheim, Germany) using a mouse monoclonal antibody (clone(s): 8B1.2, IG5.2 and 2D4.2) according the manufacturer’s instructions (DCS, Hamburg, Germany).
Isolation of genomic DNA
Formalin-fixed and paraffin-embedded tissue samples were subjected to de-paraffination followed by DNA extraction using the QiaAMP DNA Mini Kit according to the manual’s instructions (Qiagen, Hilden, Germany). DNA concentrations and sample purity were determined by UV spectroscopy and DNA concentration was set to 40 ng/μl.
Real-time PCR
Real-Time PCR was performed using TaqMan universal PCR Mastermix (Thermo Fisher Scientific, Darmstadt, Germany) in a total volume of 25 μl, including 400 nM of each primer, 100 nM of TaqMan-probe and 160 ng of sample DNA. All samples were analyzed in a 7300 Real-Time PCR System (Thermo Fisher Scientific) with the following conditions: After 2 min at 50 °C and 10 min initial denaturation at 95 °C, samples were submitted to 45 cycles consisting of a 95 °C denaturation step for 25 s and an annealing/extension step at 60 °C for 1 min. For every sample, cytomegalovirus and human glyceraldehyde 3-phosphate dehydrogenase (GAPDH) copy numbers were quantified via TaqMan-Assays using separate reactions. The cytomegalovirus PCR set-up was composed of a primer pair/probe targeting a 69 bp region of the pp65 gene (Forward: 5′-CAGCCACGGGATCGTACTG-3′; Reverse: 5′-GCAGATCTTCCTGGAGGTACAAG-3′; probe: 5′-FAM-CGCGAGACCGTGGAACTGCG-TAMRA-3′). During GAPDH-PCR, a 99 bp product was generated (Forward: 5′-CTCCCCACACACATGCACTTA-3′; Reverse: 5′-CCTAGTCCCAGGGCTTTGATT-3′; probe: 5′-VIC- AAAAGAGCTAGGAAGGACAGGCAACTTGGC-TAMRA-3′). All primers and probes were generated using the Primer Express software 3.0 (Thermo Fisher Scientific) or published elsewhere.16
Calculation of viral loads
For absolute quantification of cytomegalovirus and GAPDH copy numbers, one plasmid containing both target sequences served as external standard (Thermo Fisher Scientific). Tenfold dilution series ranging from 2 × 106 to 2 × 102 copies were used to generate two standard curves by plotting the Ct values to the copy numbers (Supplementary Figure S1). Calculation of absolute cytomegalovirus and GAPDH copy numbers in unknown samples was achieved via the SDS software (version 1.3.1, Thermo Fisher Scientific) by referring to the respective standard curve. As GAPDH is a single copy gene with two alleles per diploid genome, half the amount of GAPDH copy numbers equals analyzed cell number. Finally, viral load was expressed as cytomegalovirus copy numbers/105 cells. All samples and standards were analyzed at least in duplicates. Amplification of GAPDH was also used to control absence of PCR inhibitors in extracted DNA samples. The mean GAPDH-Ct value of 27.6±0.9 that is derived from 115 samples served as a guide level. Samples with GAPDH-Ct-values ≥30.6 were excluded or re-analyzed. Assessment of cytomegalovirus copy numbers using the Qiagen Artus Kit was carried out according to the manufacturer’s instructions (Qiagen).
Antiviral therapy application
Patient selections for antiviral therapy start were made individually based on clinical data and histological as well as molecular diagnostic findings and patients’ preference. All patients included had an active inflammation (inflammation score ≥1), a positive cytomegalovirus detection via PCR and were refractory to standard therapy. Administration of antiviral therapy was carried out according to the respective guidelines initially starting with intravenous Ganciclovir (5 mg/kg) every 12 h for 14 days. Then therapy was shifted to oral administration using 900 mg Valganciclovir twice a day for 2–3 weeks. Afterwards, therapy was continued with administration of 900 mg Valganciclovir once a day for 4–8 weeks. Therapy course was monitored by analyzing changes in viral load numbers, and for inflammatory bowel disease patients, clinical response was evaluated using the partial Mayo score. According to Lewis et al,17 a two-point reduction (or greater) of the partial Mayo score was considered as indicative for clinical response to antiviral treatment.
Statistical analysis
For statistical comparison, the Fisher’s exact test were utilized while P<0.05 was considered significant. Determination of cutoff thresholds was performed using the Cutoff Finder software as described.18
Results
Performance of the cytomegalovirus detection assay
The assay for quantitative cytomegalovirus detection in human tissue consists of two independent PCR reactions for absolute quantification of cytomegalovirus and GAPDH copy numbers, respectively. Calculation of viral loads was carried out as described in the ‘Material and methods’ section and finally expressed as the amount of cytomegalovirus copy numbers/105 cells. In order to estimate assay performance, cytomegalovirus and GAPDH PCRs were extensively tested for assay characteristics such as analytical sensitivity, PCR efficiency, linear dynamic range as well as intra-assay and interassay variability and long-time variability according to the ‘Minimum information for publication of quantitative real-time PCR experiments’ (MIQE) guidelines.19 Results are shown in Supplementary Figure S1 and Supplementary Table S1, which identify the assay as suitable for sensitive cytomegalovirus detection and quantification. Moreover, performance of cytomegalovirus detection was compared with the commercial cytomegalovirus Artus Kit (Qiagen), revealing comparable results with a yet slightly increased sensitivity of the assay described here (Supplementary Figure S2, Supplementary Table S2).
Cytomegalovirus detection in patients
Cytomegalovirus infection was evaluated in 917 colon biopsy samples derived from 136 inflammatory bowel disease patients (male/female: 81/55; median age in years: 44, range in years: 12–88). As shown in Table 1, with 103 cases the percentage of patients diagnosed with ulcerative colitis was 76%, 30 patients had a Crohn’s disease (22%) and three patients were diagnosed with ‘colitis indeterminata’ (2%). In 37 of the 136 patients, cytomegalovirus DNA was detectable (27%). Interestingly, in 33% of ulcerative colitis patients cytomegalovirus was detectable while only 10% of Crohn’s disease cases were shown to be positive for cytomegalovirus. This difference is highly significant as demonstrated by a P-value of P=0.019.
Viral loads of cytomegalovirus-positive patients
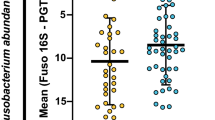
Generally, viral loads showed a broad variation ranging from 5 copies/105 cells to 8.7 × 105 copies/105 cells. The mean viral load covering all cytomegalovirus positive samples was approximately 37 400 (±1.50 × 105) cytomegalovirus copies/105 cells and the median was 900 cytomegalovirus copies/105 cells (Figure 1a). Regarding the total distribution of viral loads, with 13 patients (35%) the majority showed viral loads of <100 cytomegalovirus copies/105 human cells. Six patients (16%) displayed viral loads ranging between 100 and 1000, 11 patients (30%) had 1000–10 000 cytomegalovirus copies/105 cells, 5 patients (14%) had viral loads between 10 000 and 100 000 cytomegalovirus copies/105 cells and 2 patients (5%) revealed >100 000 cytomegalovirus copies/105 cells (Figure 1b). For assay control, a panel composed of 21 cytomegalovirus-negative control samples was analyzed. In none of these, cytomegalovirus DNA could be detected (data not shown). By histological evaluation only in tissue sections with exceptionally high viral loads such as 8.9 × 105 copies/105 cells cytomegalovirus infection could be confirmed by the presence of typical inclusion bodies (owl’s eyes) as shown in Figure 2a and Table 2.
Distribution of viral loads in cytomegalovirus-positive patients. A wide spectrum of cytomegalovirus viral burden was detected in 37 patients ranging from 5 copies/105 cells to 8.7 × 105 cytomegalovirus copies/105 cells. (a) The mean viral load covering all cytomegalovirus-positive samples was approximately 37 400 (±1.50 × 105) cytomegalovirus copies/105 cells and the median (—) was calculated as 900 cytomegalovirus copies/105 cells. (b) Frequency of patients with 1–100, 100–1000, 1000–10 000, 10 000–100 000 and >100 000 cytomegalovirus copies/105 cells. Along with the total sample set (black), distribution of patients according to antiviral treatment status is shown in dark gray and light gray.
Histological detection of cytomegalovirus infection. (a) Detection of viral inclusion (indicated by black arrows) using hematoxylin and eosin-stained tissue sections. (b) Detection of cytomegalovirus infection by immunohistochemistry. A colon tissue section stained with a monoclonal antibody directed against cytomegalovirus is shown. Immunohistochemistry-positive cells are indicated by arrows.
Local manifestation of cytomegalovirus infections
For the majority of patients, multiple consecutive tissue sections related to different intestinal localization sites were evaluated (up to 12, mean: 4 consecutive tissue sections per patient evaluation). Here major differences of viral loads could be detected in different tissue sections of individual patients, indicating that cytomegalovirus infections could be locally quite heterogeneous (Figure 3a). In order to identify the area that is mostly affected by cytomegalovirus, localization sites showing the maximum of viral load for each case were recapitulated in Figure 3b. Owing to follow-up monitoring of a subset of patients, a total of 60 cytomegalovirus evaluations corresponding to the 37 cytomegalovirus-positive patients were included (each evaluation consisted of the cytomegalovirus analysis of multiple biopsy samples derived from different localization sites). Interestingly, within 22 out of the 60 cytomegalovirus evaluations, the peak of viral loads was found to be in the rectum section (37%), while no cytomegalovirus was detectable in the upper colon areas.
Localization of cytomegalovirus infections. (a) Local distribution of cytomegalovirus viral loads in consecutive tissue sections of six exemplary patients is shown. The numbers on x axis indicate different intestinal areas with low numbers referring to the more upper part and higher numbers to the lower part of the colon. On the y axis, the cytomegalovirus load is presented. (b) Cumulative evaluation of cytomegalovirus-positive tissue sections.
Correlation of viral load number with antiviral treatment response
Of the 37 patients with positive cytomegalovirus detection, 13 patients (35%) were subjected to antiviral therapy (Figure 4). Here treatment decisions were based on histological and clinical findings as described in the ‘Materials and Methods’ section, most importantly, no response to standard therapy. As shown in Table 2, all 13 patients revealed an active inflammation with an inflammation score ≥1. With 77% (10/13 patients), the major fraction even had a highly severe grade of inflammation as characterized by an inflammation score of three and the presence of ulceration. However, typical viral inclusions could only be detected in one case, thus again demonstrating its limited sensitivity for cytomegalovirus detection. During antiviral therapy, 12 of the 13 patients (92%) had a positive clinical response as indicated by a ≥2-point reduction of the partial Mayo score (Table 2). For one patient, no therapy response data could be available owing to premature death. Response to antiviral treatment was also monitored by analysis of follow-up biopsies using cytomegalovirus PCR. As exemplarily depicted in Figure 5, after therapy initiation in every case a significant reduction in cytomegalovirus viral load could be detected, pointing to a positive therapy response. However, 5 patients displayed relapses after 105–609 days. Two refused restart of antiviral therapy while the remaining three again showed response to restart of antiviral treatment accompanied by clinical remission.
Clinical course of 136 patients enrolled in the study. *For four patients, no therapy response data were available because of premature death (†) or follow-up monitoring was not feasible. Numbers in dark gray boxes reflect the number of patients with cytomegalovirus viral loads <600 cytomegalovirus copies/105 cells, numbers in light gray boxes show the number of patients with viral burden >600 cytomegalovirus copies/105 cells.
Follow-up monitoring of antiviral treatment efficacy using cytomegalovirus PCR. For three patients, timeline decrease of cytomegalovirus viral loads in response of antiviral treatment is shown. The respective time points are referred to days of diagnostic sample evaluations by PCR. Viral load numbers are indicated and represent the respective peak values of multiple biopsy samples analyzed. Selection of patients was made exemplarily to demonstrate alterations of viral load values in response to antiviral therapy starting from different amounts of initial viral burden.
Twenty-four of the 37 patients with positive cytomegalovirus detection (65%) were subjected to adequate standard therapy without antiviral treatment. Here therapy was mainly composed of immunomodulatory and/or anti-inflammatory treatment strategies according to the respective guidelines. Further information on individual treatments and therapeutic escalation is presented in Supplementary Table S3. As shown in Figure 4, standard therapy led to remission of 5 patients (21%). In contrast, for 14 patients no remission could be achieved (58%). Furthermore, colectomy was the final option for two patients. For three patients, unfortunately, follow-up monitoring was not feasible.
In order to calculate a certain viral load threshold indicative for a clinically relevant cytomegalovirus infection, a retrospective comparison of viral loads with antiviral therapy response was performed using the ‘Cutoff Finder’ software tool.16 Here a threshold of 600 cytomegalovirus copies/105 cells was found to be indicative for a cytomegalovirus infection suitable for antiviral treatment. Applying this cutoff to the sample set revealed a sensitivity and specificity of 100 and 93%, respectively.
Cytomegalovirus detection by immunohistochemistry
For patients with clinically relevant cytomegalovirus infection (as indicated by a therapeutic response to antiviral treatment), cytomegalovirus detection was also performed using immunohistochemistry. In order to compare sensitivity and concordance with cytomegalovirus detection via threshold-guided real-time PCR, here identical tissue sections were used for both methods. As shown in Table 2, cytomegalovirus infection could be verified in 6 out of the 12 patient samples (for 1 case immunohistochemistry could not be performed owing to material limitations). Looking at the correlation between both methods, as expected, in samples with rather high viral load values (>10 000 cytomegalovirus copies/105 cells) cytomegalovirus infection could also be detected by immunohistochemistry. However, regarding samples with viral loads in the range of 1000–10 000 cytomegalovirus copies/105 cells, for only 3 out of the 6 samples cytomegalovirus positivity could be confirmed at immunohistochemistry level. Considering all samples, the concordance between PCR and immunohistochemistry was 100% (3/3) for samples with viral loads of >10 000 but only 33% (3/9) regarding samples with viral burden <10 000 cytomegalovirus copies/105 cells. Of note, samples with ≤1000 cytomegalovirus copies/105 cells were never identified as cytomegalovirus positive by immunohistochemistry.
Altogether, 50% of the therapeutic responders would have been missed by solely relying on cytomegalovirus detection using immunohistochemistry, thus demonstrating the diagnostic power of threshold-guided real-time PCR for identification of clinically relevant cytomegalovirus infections in inflammatory bowel disease patients.
Discussion
Cytomegalovirus infections emerge as a well-known cause of disease complication in patients suffering from inflammatory bowel disease, especially in a therapy-refractory setting.3, 4, 5, 20, 21 For cytomegalovirus detection in tissue, histological evaluation supported by immunohistochemistry is often considered as ‘golden standard’. However, albeit an excellent specificity this method is adversely affected by its rather limited sensitivity.2, 8, 11 Therefore, application of highly sensitive PCR and quantitative real-time PCR is recently increasing to improve cytomegalovirus diagnostics. However, recommendations for PCR-dependent cytomegalovirus diagnostics as published by the ECCO committee are based on usage of fresh material making the analysis hardly feasible for many diagnostic laboratories.21 In order to circumvent this and to further optimize cytomegalovirus diagnostics for inflammatory bowel disease patients, in the present study, we extended application of quantitative real-time cytomegalovirus PCR to the usage of formalin-fixed and paraffin-embedded material. Large testing according to the MIQE guidelines19 as well as performance comparison with the commercial Qiagen Artus Kit revealed our PCR assay to be ideal for cell number-normalized absolute cytomegalovirus quantification. Using this assay, cytomegalovirus infection was evaluated in a comprehensive sample set composed of 917 colonic biopsy samples derived from 136 inflammatory bowel disease patients. Here in 37 patients cytomegalovirus detection was notable (27%). This is a somewhat reduced prevalence as previously reported by Dimitroulia et al9 and Roblin et al13 who found 32 and 38% of inflammatory bowel disease patients to be positive for cytomegalovirus by real-time PCR, respectively. The difference, however, may be related to the respective sample sets used and total sample amount tested as these studies were conducted using smaller case numbers.9, 13 Of note, in studies solely based on histological and immunohistochemical detection methods a significantly lower percentage of cytomegalovirus positivity was reported ranging from 4.5% to 16.6%, thus presumably reflecting the reduced sensitivity of those methods.4, 22, 23
Regarding the extent of cytomegalovirus detection in ulcerative colitis and Crohn’s disease patients separately, a significant difference became obvious (P=0.019). Although in 34 out of our 104 ulcerative colitis patient samples (33%), cytomegalovirus DNA could be detected, only 3 of the 30 Crohn’s disease samples were positive for cytomegalovirus detection (10%). Although the scale of the Crohn’s disease fraction was considerably lower compared with the ulcerative colitis sample amount, these results underscore observations already made by Takahasi and Tange5 and Nakase et al24 together indicating that cytomegalovirus infection may be a rare event in Crohn’s disease.
Our systematic sampling and analysis of consecutive tissue sections related to different colon localization sites allowed the identification of intestinal areas that are predominately affected by cytomegalovirus infections. Besides a generally quite heterogeneous distribution pattern, the rectum part was shown to be prone to cytomegalovirus infections. Of note, the proximal colon areas never displayed peak values of viral loads. Implying that especially in ulcerative colitis the rectum part is usually the most inflamed area of the disease, these results might support a functional link between cytomegalovirus infection and inflammatory extent in ulcerative colitis as previously suggested.1, 7, 13, 25 Another fact arguing for that is based on the relationship between cytomegalovirus infection and therapy response to antiviral treatment. In our study, 13 patients who were positive for cytomegalovirus detection accompanied with a (high) grade of inflammation and who were refractory to standard therapy were subjected to antiviral treatment. Thereof, 12 patients showed positive therapy response as demonstrated by a partial Mayo score reduction ≥2. Although later on, five patients displayed a relapse, three patients who agreed to restart antiviral treatment again showed positive therapy response. This high therapeutic efficacy level is comparable to that found in other studies,12, 13, 26, 27, 28 altogether pointing to a substantial therapeutic benefit of antiviral treatment for inflammatory bowel disease patients with clinically related cytomegalovirus infections. Indeed, recently it was shown that antiviral treatment was associated with a more favorable long-time outcome in ulcerative colitis patients leading to an approximately 50% reduction rate of clinical relapse that require new treatment or colectomy.29 Consistently, Jones et al23 found that antiviral treatment improved surgery-free outcomes especially of inflammatory bowel disease patients with high-grade cytomegalovirus density. On the other hand, some studies failed to uncover a positive aspect of antiviral treatment for patient’s benefit.30, 31 For instance, in a meta-analysis including 9 studies with a total of 176 patients no positive association between antiviral treatment and favorable patient outcome could be demonstrated.32 However, according to the authors the quality of the studies analyzed was rather limited. Moreover, in a recent re-examination of the same data set the informative value of the original investigation was further challenged by implying inappropriate study inclusion.33 In our study, clinical improvement was always accompanied by a drastic reduction of viral load numbers, which also indicates the initial viral loads as being clinically relevant.
Using a web-based cutoff determination tool,18 a retrospective whole data analysis revealed a threshold of >600 cytomegalovirus copies/105 cells as indicative for a clinically relevant cytomegalovirus infection, suitable for antiviral treatment. In this context, it is worthy to note that one relapsed patient who refused re-start of antiviral treatment finally displayed a viral load of 786 cytomegalovirus copies/105 cells that was associated with sustained disease activity. Hence, this is further evidence for the clinical utility of this cutoff level. Applying this threshold to samples revealed an excellent sensitivity of 100% and good specificity of 93%. In comparison to former studies, this is in a highly acceptable range.13, 34
Looking at the patients with positive cytomegalovirus detection but without antiviral treatment, one patient showed remission although initially viral load number were above the 600 cytomegalovirus copies/105 cells threshold. In this case, irrespective of the viral load, cytomegalovirus might have played a minor role or was cleared spontaneously, a phenomenon that was also observed by Matsuoka et al.35 However, whether the remission process could have been further accelerated by antiviral treatment cannot be excluded, too. On the other hand, 6 out of the 14 patients (43%) without remission revealed viral loads higher than the 600 cytomegalovirus copies/105 cells threshold. For two of these patients, even a colectomy was recommended at the end of this study. Considering the high therapeutic efficacy level of antiviral treatment by using the 600 cytomegalovirus copies/105 cells viral load threshold, for those patients an antiviral treatment could have been an alternative therapeutic option to potentially achieve remission.
Interestingly, immunohistochemistry led to insufficient detection of cytomegalovirus infections. Here only patient samples with rather high cytomegalovirus copy numbers (>104 cytomegalovirus copies/105 cells) could be reliably identified. As a consequence, 50% of the patients with clinically relevant cytomegalovirus infection (as confirmed by the positive response to antiviral therapy) would have been missed by solely relying on this method. This also complies with observations made by several other groups and further challenges immunohistochemistry to be appropriated for accurate cytomegalovirus detection in tissues of inflammatory bowel disease patients.12, 13, 25, 36 For reliable cytomegalovirus detection in those patients, we therefore would recommend to test multiple tissue sections showing clear signs of (high grade) inflammation by using threshold-guided real-time PCR.
Although usage of quantitative real time PCR is increasing, findings derived from studies based on classical end point PCR might be one reason for the overall conflicting data situation regarding the role of cytomegalovirus in inflammatory bowel disease.9, 12, 13, 25, 34, 36, 37 A binary ‘yes/no’ PCR result unsuitable to filter out substantial cytomegalovirus infections from clinically irrelevant latency may lead to false correlation clues concerning the association of cytomegalovirus with inflammatory bowel disease appearance.31 In an analogous manner, conclusions drawn from PCR-based analyses using peripheral blood or serum specimen are also difficult to consider as an increasing body of evidence suggests poor correlation between cytomegalovirus levels in serum and colon tissue, especially in mild–moderate forms of inflammatory bowel disease.12, 13, 38, 39 More importantly, cytomegalovirus end-organ disease has been suggested to exist without systemic involvement.40, 41, 42, 43, 44 Therefore, additional studies based on tissue-specific and quantitative cytomegalovirus detection systems such as quantitative real-time PCR would be helpful to further clarify the role of cytomegalovirus in inflammatory bowel disease and, more importantly, to aid stratification of inflammatory bowel disease patients for antiviral treatment. Recently, in a publication by Roblin et al,13 a viral load threshold of 250 cytomegalovirus copies/mg tissue was reported to be predictive for resistance to immunosuppressive therapy in ulcerative colitis patients, showing a sensitivity and specificity of 100 and 66%, respectively. As a consequence, the ECCO has incorporated this threshold into their guidelines for cytomegalovirus diagnostics of therapy-refractory inflammatory bowel disease patients.21 However, as this threshold is based on fresh material, an area-wide adequate day-to-day diagnostic supply would require a high level of transport logistics to assure appropriate sample quality. Furthermore, this normalization strategy for quantitative PCR analysis of intracellular viruses such as cytomegalovirus is difficult.14 In order to circumvent this, here formalin-fixed and paraffin-embedded material was used to define a cell number-normalized cutoff level, thus making it more feasible for implementation of the present cytomegalovirus detection assay in most diagnostic laboratories.
In summary, we show that quantitative real-time PCR is a useful diagnostic tool for objective cytomegalovirus detection in colon tissue of inflammatory bowel disease patients that clearly exceeds immunohistochemistry in terms of sensitivity and diagnostic reliability. As this assay is based on formalin-fixed and paraffin-embedded tissue material, a readily area-wide assay implementation in diagnostic laboratories is warranted. A threshold of 600 cytomegalovirus copies/105 cells was shown to separate relevant cytomegalovirus infection from common latent cytomegalovirus. This cutoff level therefore may serve as a ‘biomarker’ to stratify inflammatory bowel disease patients for antiviral treatment. Interestingly, this threshold is in a comparable range of those found in two recent studies where 0.01 cytomegalovirus copy numbers/cell and 1000 cytomegalovirus copies/105 cells, respectively, were proposed to be of clinical relevance.25, 34 All these findings cooperatively underline the diagnostic power of quantitative real-time PCR for cytomegalovirus detection in inflammatory bowel disease. Nevertheless, further prospective studies are needed to confirm these results. That may lead to the definition of a universal cutoff level (or range) indicative of clinically relevant cytomegalovirus infections in inflammatory bowel disease.
References
Onyeagocha C, Hossain MS, Kumar A et al. Latent cytomegalovirus infection exacerbates experimental colitis. Am J Pathol 2009;175:2034–2042.
Garrido E, Carrera E, Manzano R et al. Clinical significance of cytomegalovirus infection in patients with inflammatory bowel disease. World J Gastroenterol 2013;19:17–25.
Kambham N, Vij R, Cartwright CA et al. Cytomegalovirus infection in steroid-refractory ulcerative colitis: a case-control study. Am J Surg Pathol 2004;28:365–373.
Criscuoli V, Casà A, Orlando A et al. Severe acute colitis associated with CMV: a prevalence study. Dig Liver Dis 2004;36:818–820.
Takahashi Y, Tange T . Prevalence of cytomegalovirus infection in inflammatory bowel disease patients. Dis Colon Rectum 2004;47:722–726.
de Saussure P, Lavergne-Slove A, Mazeron MC et al. A prospective assessment of cytomegalovirus infection in active inflammatory bowel disease. Aliment Pharmacol Ther 2004;20:1323–1327.
Lawlor G, Moss AC . Cytomegalovirus in inflammatory bowel disease: pathogen or innocent bystander? Inflamm Bowel Dis 2010;16:1620–1627.
Wu GD, Shintaku IP, Chien K et al. A comparison of routine light microscopy, immunohistochemistry, and in situ hybridization for the detection of cytomegalovirus in gastrointestinal biopsies. Am J Gastroenterol 1989;84:1517–1520.
Dimitroulia E, Spanakis N, Konstantinidou AE et al. Frequent detection of cytomegalovirus in the intestine of patients with inflammatory bowel disease. Inflamm Bowel Dis 2006;12:879–884.
Machida U, Kami M, Fukui T et al. Real-time automated PCR for early diagnosis and monitoring of cytomegalovirus infection after bone marrow transplantation. J Clin Microbiol 2000;38:2536–2542.
Mills AM, Guo FP, Copland AP et al. A comparison of CMV detection in gastrointestinal mucosal biopsies using immunohistochemistry and PCR performed on formalin-fixed, paraffin-embedded tissue. Am J Surg Pathol 2013;37:995–1000.
Yoshino T, Nakase H, Ueno S et al. Usefulness of quantitative real-time PCR assay for early detection of cytomegalovirus infection in patients with ulcerative colitis refractory to immunosuppressive therapies. Inflamm Bowel Dis 2007;13:1516–1521.
Roblin X, Pillet S, Oussalah A et al. Cytomegalovirus load in inflamed intestinal tissue is predictive of resistance to immunosuppressive therapy in ulcerative colitis. Am J Gastroenterol 2011;106:2001–2008.
Watzinger F, Ebner K, Lion T . Detection and monitoring of virus infections by real-time PCR. Mol Aspects Med 2006;27:254–298.
Gupta RB, Harpaz N, Itzkowitz S et al. Histologic inflammation is a risk factor for progression to colorectal neoplasia in ulcerative colitis. Gastroenterol 2007;133:1099–1105.
Griscelli F, Barrois M, Chauvin S et al. Quantification of human cytomegalovirus DNA in bone marrow transplant recipients by real-time PCR. J Clin Microbiol 2001;39:4362–4369.
Lewis JD, Chuai S, Nessel L et al. Use of the non-invasive components of the Mayo score to assess clinical response in ulcerative colitis. Inflamm Bowel Dis 2008;14:1660–1666.
Budczies J, Klauschen F, Sinn BV et al. Cutoff Finder: a comprehensive and straightforward Web application enabling rapid biomarker cutoff optimization. PLoS ONE 2012;14:e51862.
Bustin SA, Benes V, Garson JA et al. The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem 2009;55:611–622.
Powell RD, Warner NE, Levine RS et al. Cytomegalic inclusion disease and ulcerative colitis: report of a case in a young adult. Am J Med 1961;30:334–340.
Rahier JF, Magro F, Abreu C et al. Second European evidence-based consensus on the prevention, diagnosis and management of opportunistic infections in inflammatory bowel disease. J Crohns Colitis 2014;8:443–468.
Kim JJ, Simpson N, Klipfel N et al. Cytomegalovirus infection in patients with active inflammatory bowel disease. Dig Dis Sci 2010;55:1059–1065.
Jones A, McCurdy JD, Loftus EV Jr et al. Effects of antiviral therapy for patients with inflammatory bowel disease and a positive intestinal biopsy for cytomegalovirus. Clin Gastroenterol Hepatol 2015;13:949–955.
Nakase H, Yoshino T, Honzawa Y et al. Low prevalence of CMV infection in patients with Crohn's disease in comparison with ulcerative colitis: effect of different immune response on prevalence of CMV infection. Dig Dis Sci 2010;55:1498–1499.
Ciccocioppo R, Racca F, Paolucci S et al. Human cytomegalovirus and Epstein-Barr virus infection in inflammatory bowel disease: need for mucosal viral load measurement. World J Gastroenterol 2015;21:1915–1926.
Cottone M, Pietrosi G, Martorana G et al. Prevalence of cytomegalovirus infection in severe refractory ulcerative and Crohn's colitis. Am J Gastroenterol 2001;96:773–775.
Sager K, Alam S, Bond A et al. Review article: cytomegalovirus and inflammatory bowel disease. Aliment Pharmacol Ther 2015;41:725–733.
Vega R, Bertrán X, Menacho M et al. Cytomegalovirus infection in patients with inflammatory bowel disease. Am J Gastroenterol 1999;94:1053–1056.
Maconi G, Lombardini M, Furfaro F et al. Long-term outcome of inflammatory bowel diseases with cytomegalovirus colitis: effect of antiviral treatment. Eur J Gastroenterol Hepatol 2014;26:1146–1151.
Delvincourt M, Lopez A, Pillet S et al. The impact of cytomegalovirus reactivation and its treatment on the course of inflammatory bowel disease. Aliment Pharmacol Ther 2014;39:712–720.
Kim YS, Kim YH, Kim JS et al. Long-term outcomes of cytomegalovirus reactivation in patients with moderate to severe ulcerative colitis: a multicenter study. Gut Liver 2014;8:643–647.
Kopylov U, Eliakim-Raz N, Szilagy A et al. Antiviral therapy in cytomegalovirus-positive ulcerative colitis: a systematic review and meta-analysis. World J Gastroenterol 2014;20:2695–2703.
Wu XW, Yang MF, Li N et al. Unfavorable outcome of antiviral therapy in cytomegalovirus-positive ulcerative colitis may be due to inappropriate study inclusion in meta-analysis. World J Gastroenterol 2015;21:1689–1690.
Ganzenmueller T, Henke-Gendo C, Schlué J et al. Quantification of cytomegalovirus DNA levels in intestinal biopsies as a diagnostic tool for CMV intestinal disease. J Clin Virol 2009;46:254–258.
Matsuoka K, Iwao Y, Mori T et al. Cytomegalovirus is frequently reactivated and disappears without antiviral agents in ulcerative colitis patients. Am J Gastroenterol 2007;102:331–337.
McCoy MH, Post K, Sen JD et al. qPCR increases sensitivity to detect cytomegalovirus in formalin-fixed, paraffin-embedded tissue of gastrointestinal biopsies. Human Pathol 2014;45:48–53.
Zidar N, Ferkolj I, Tepeš K et al. Diagnosing cytomegalovirus in patients with inflammatory bowel disease-by immunohistochemistry or polymerase chain reaction? Virchows Arch 2015;466:533–539.
Kim JW, Boo SJ, Ye BD et al. Clinical utility of cytomegalovirus antigenemia assay and blood cytomegalovirus DNA PCR for cytomegaloviral colitis patients with moderate to severe ulcerative colitis. J Crohns Colitis 2014;8:693–701.
Kou T, Nakase H, Tamaki H et al. Cytomegalovirus infection in patients with ulcerative colitis diagnosed by quantitative real-time PCR analysis. Dig Dis Sci 2006;51:1052–1055.
Gerna G, Lilleri D, Furione M et al. Human cytomegalovirus end-organ disease is associated with high or low systemic viral load in preemptively treated solid-organ transplant recipients. New Microbiol 2012;35:279–287.
Berk T, Gordon SJ, Choi HY et al. Cytomegalovirus infection of the colon: a possible role in exacerbations of inflammatory bowel disease. Am J Gastronterol 1985;80:355–360.
Rachima C, Maoz E, Apter S et al. Cytomegalivirus infection associated with ulcerative colitis in immunocompetent individuals. Postgrad Med J 1998;74:486–489.
Hofkin GA, Ting CD . Case report: recurrence of chronic ulcerative colitis induced by intercurrent cytomegalic virus infection. Md Med J 1995;44:1047–1048.
Kandiel A, Lashner B . CMV-colitis complicates IBD. Am J Gastroenterol 2006;101:1857–1865.
Acknowledgements
We thank Marion Menke for performing immunohistochemistry experiments and for technical assistance.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Modern Pathology website
Rights and permissions
About this article
Cite this article
Wethkamp, N., Nordlohne, EM., Meister, V. et al. Identification of clinically relevant cytomegalovirus infections in patients with inflammatory bowel disease. Mod Pathol 31, 527–538 (2018). https://doi.org/10.1038/modpathol.2017.149
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2017.149
This article is cited by
-
SFED recommendations for IBD endoscopy during COVID-19 pandemic: Italian and French experience
Nature Reviews Gastroenterology & Hepatology (2020)
-
Cytomegalovirus reactivation in inflammatory bowel disease: an uncommon occurrence related to corticosteroid dependence
Modern Pathology (2019)