Abstract
Malignant phyllodes tumors of the breast are poorly understood rare neoplasms with potential for aggressive behavior. Few efficacious treatment options exist for progressed or metastatic disease. The molecular features of malignant phyllodes tumors are poorly defined, and a deeper understanding of the genetics of these tumors may shed light on pathogenesis and progression and potentially identify novel treatment approaches. We sequenced 510 cancer-related genes in 10 malignant phyllodes tumors, including 5 tumors with liposarcomatous differentiation and 1 with myxoid chondrosarcoma-like differentiation. Intratumoral heterogeneity was assessed by sequencing two separate areas in 7 tumors, including non-heterologous and heterologous components of tumors with heterologous differentiation. Activating hotspot mutations in FGFR1 were identified in 2 tumors. Additional recurrently mutated genes included TERT promoter (6/10), TP53 (4/10), PIK3CA (3/10), MED12 (3/10), SETD2 (2/10) and KMT2D (2/10). Together, genomic aberrations in FGFR/EGFR PI-3 kinase and RAS pathways were identified in 8 (80%) tumors and included mutually exclusive and potentially actionable activating FGFR1, PIK3CA and BRAF V600E mutations, inactivating TSC2 mutation, EGFR amplification and PTEN loss. Seven (70%) malignant phyllodes tumors harbored TERT aberrations (six promoter mutations, one amplification). For comparison, TERT promoter mutations were identified by Sanger sequencing in 33% borderline (n=12) and no (0%, n=8) benign phyllodes tumors (P=0.391 and P=0.013 vs malignant tumors, respectively). Genetic features specific to liposarcoma, including CDK4/MDM2 amplification, were not identified. Copy number analysis revealed intratumoral heterogeneity and evidence for divergent tumor evolution in malignant phyllodes tumors with and without heterologous differentiation. Tumors with liposarcomatous differentiation revealed more chromosomal aberrations in non-heterologous components compared with liposarcomatous components. EGFR amplification was heterogeneous and present only in the non-heterologous component of one tumor with liposarcomatous differentiation. The results identify novel pathways involved in the pathogenesis of malignant phyllodes tumors, which significantly increase our understanding of tumor biology and have potential clinical impact.
Similar content being viewed by others
Main
Phyllodes tumors of the breast are rare biphasic fibroepithelial neoplasms, which comprise approximately 2.5% of fibroepithelial tumors of the breast and <1% of all primary breast tumors.1, 2 Clinical behavior is widely variable and correlates in part with histological grade. Phyllodes tumors are classified as benign (~60–75%), borderline (~15–20%) or malignant (~10–20%) based on a constellation of histological features.1, 3, 4, 5 Phyllodes tumor recurrence correlates with increasing grade, with malignant tumors having the highest recurrence risk (up to ~30%).1, 3 Distant metastasis occurs essentially only in malignant phyllodes tumors, with reported rates up to ~22%.3, 6 Prognosis and survival for patients with metastatic malignant phyllodes tumors is poor, with few available treatment options.1, 3, 7, 8 Indeed, the mainstay of malignant phyllodes tumor treatment is surgery with or without radiation, with the role and efficacy of standard chemotherapy being uncertain and without proven benefit beyond palliation.7, 9, 10, 11, 12, 13 A better grasp of the genomic features of these poorly understood tumors is therefore needed to shed light on their pathogenesis and progression in hopes of identifying novel treatment approaches and potentially actionable targets for the subset with aggressive behavior.
The pathogenesis and genetic alterations driving the development and progression of phyllodes tumors in general and of malignant phyllodes tumors in particular remain poorly defined. Comparative genomic hybridization studies have highlighted recurrent large scale copy number alterations, including gains of chromosome 1q, 5p, 7 and 8 and losses of 6q, 9p, 10p and 13q, among others, with the mean number of chromosomal alterations increasing from benign to malignant phyllodes tumors.14, 15, 16, 17, 18, 19, 20 Presumed targets of copy number changes include CDKN2A (9p), RB1 (13q) TP53 (17p) and EGFR (7p), all of which may show aberrations in subsets of phyllodes tumors.14, 20, 21, 22, 23, 24, 25, 26, 27, 28 Immunohistochemical studies have also implicated insulin-like growth factor and platelet-derived growth factor signaling in phyllodes tumor development.29, 30, 31 Recent studies have begun to examine the genomic features of phyllodes tumors at higher resolution.21, 24 Cani et al.21 used targeted next-generation sequencing of 126 genes to identify hotspot mutations in codon 44 of MED12, a subunit of the multiprotein transcriptional regulator Mediator complex, in phyllodes tumors. MED12 mutations, which are also present in fibroadenomas and uterine smooth muscle tumors, have subsequently been identified across the spectrum of benign, borderline and malignant phyllodes tumors, with some, but not all, authors reporting fewer MED12 mutations in malignant compared with benign and borderline tumors.21, 32, 33, 34, 35, 36, 37, 38, 39 The findings not only reveal a common pathway potentially related to dysregulated estrogen signaling in fibroepithelial lesions, but also suggest that some malignant phyllodes tumors may develop or progress along a separate pathway.32, 37 Tan et al.24 performed whole-exome sequencing of 22 phyllodes tumors (10 benign, 8 borderline and only 4 malignant tumors) and found recurrent retinoic acid receptor alpha (RARA) mutations in fibroadenomas, lower grade phyllodes tumors and 1 malignant phyllodes tumor, further supporting this notion. TP53, NF1 and RB1 mutations and IGF1R, MYC and CCND1 amplification appear to be more common in borderline and/or malignant phyllodes tumors compared with benign phyllodes tumors or fibroadenomas and may be drivers for tumor progression.21, 24, 28 In addition, telomerase reverse transcriptase (TERT) was recently shown to be amplified in a subset of malignant phyllodes tumors, and TERT promoter mutations were identified by Sanger sequencing in fibroadenomas and phyllodes tumors.21, 33 Together, these studies have begun to shed light on phyllodes tumor pathogenesis and progression, but the genomic features of malignant phyllodes tumors in particular remain largely unexplored, in part due to the inclusion of only few malignant tumors in prior exome sequencing studies and the utilization of limited sequencing panels in most malignant tumors studied. Given that patients with malignant phyllodes tumors would be most likely to benefit from systemic therapies, additional studies are warranted.
Malignant heterologous differentiation is sufficient for a diagnosis of malignancy irrespective of other histological features and has been suggested, along with stromal overgrowth, to be an important predictor of metastasis.1, 2, 40, 41 Liposarcomatous differentiation is the most common type of malignant heterologous differentiation in phyllodes tumors,3 with tumors showing histological features of well-differentiated, myxoid or pleomorphic liposarcoma. The molecular features of heterologous differentiation, and the genetic relationships of these elements to matched non-heterologous components, have not been studied. Indeed, although intratumoral heterogeneity is known to carry important clinical implications for cancer treatment, progression and metastasis, very little is known about intratumoral heterogeneity of phyllodes tumors, whether it be in tumors with or without heterologous differentiation.20, 42, 43, 44
In this study, we sought to examine the genomic features of malignant phyllodes tumors using capture-based next-generation sequencing of 510 cancer-related genes, with the goal of identifying driver mutations and pathways, as well as potentially actionable drug targets, important for malignant phyllodes tumor development and growth. Malignant phyllodes tumors with and without heterologous differentiation (including five with liposarcomatous differentiation) were analyzed in order to gain insight into intratumoral heterogeneity and characterize the genomic features associated with the malignant heterologous and non-heterologous phenotypes.
Materials and methods
Study Population
This study was approved by the institutional review boards of the University of California San Francisco and Kaiser Permanente. For capture-based next-generation DNA sequencing analysis, 10 primary malignant phyllodes tumors were retrieved from the archives of the Departments of Pathology of the University of California San Francisco, San Francisco General Hospital and Kaiser Permanente, spanning years 2001 to 2013 (MPT1–MPT10). For Sanger DNA sequencing, borderline (n=12) and benign (n=8) phyllodes tumors were retrieved from the archives of the University of California San Francisco and San Francisco General Hospital Departments of Pathology, spanning years 2001–2015. Specimens were fixed in 10% neutral-buffered formalin and embedded in paraffin. All cases were re-reviewed, and diagnoses and tumor grades were independently confirmed by two breast pathologists (GK and Y-YC) using criteria established by the World Health Organization.1 Six malignant phyllodes tumors had areas of malignant heterologous differentiation, including five tumors with liposarcomatous differentiation (focal or patchy in four cases and diffuse in the fifth case) and one tumor with areas reminiscent of myxoid chondrosarcoma. Clinical information of patients with malignant phyllodes tumors was obtained from the electronic medical records of the respective institutions. Selected histological and clinicopathological parameters were recorded, including patient age at diagnosis, tumor size, type of surgical procedure, margin status, treatment, recurrence and/or metastasis and survival data, as well as the presence of stromal overgrowth and the presence and type of malignant heterologous differentiation.
Capture-Based Next-Generation DNA Sequencing
Matched normal and tumor tissues were selected from 10 malignant phyllodes tumors for capture-based next-generation DNA sequencing. Sequencing was performed at the UCSF Clinical Cancer Genomics Laboratory, using an assay that targets the coding regions of 510 cancer-related genes, select introns from 40 genes and TERT promoter, with a total sequencing footprint of 2.8 Mb (UCSF500 panel). Sequencing libraries were prepared from genomic DNA extracted from punch biopsies of both tumor and normal formalin-fixed paraffin-embedded tissue. For malignant phyllodes tumors, stromal areas without epithelium were punched for DNA extraction. Target enrichment was performed by hybrid capture using a custom oligonucleotide library. Sequencing was performed on an Illumina HiSeq 2500. Duplicate sequencing reads were removed computationally to allow for accurate allele frequency determination and copy number calling. The analysis was based on the human reference sequence UCSC build hg19 (NCBI build 37), using the following software packages: BWA: 0.7.10-r789, Samtools: 1.1 (using htslib 1.1), Picard tools: 1.97 (1504), GATK: 2014.4-3.3.0-0-ga3711, CNVkit: 0.3.3, Pindel: 0.2.5a7, SATK: 2013.1-10- gd6fa6c3, Annovar: v2015Mar22, Freebayes: 0.9.20, and Delly: 0.5.9.45, 46, 47, 48, 49, 50, 51, 52, 53, 54 Only insertions/deletions up to 100 bp in length were included in the mutational analysis. Somatic single-nucleotide variants and insertions/deletions were visualized and verified using Integrated Genome Viewer. Genome-wide copy number analysis based on on-target and off-target reads was performed by CNVkit and Nexus Copy Number (Biodiscovery, Hawthorne, CA, USA).55 Large-scale chromosomal changes were defined as those involving entire chromosomes or chromosome arms, and focal changes were defined as those involving <3 megabases. For four malignant phyllodes tumors with liposarcomatous differentiation (MPT1, MPT2, MPT6 and MPT8) and the one malignant phyllodes tumor with myxoid chondrosarcoma-like differentiation (MPT5), DNA from heterologous and non-heterologous areas was extracted and analyzed separately. For one additional malignant phyllodes tumor with diffuse liposarcomatous differentiation (MPT7), DNA from two separate liposarcomatous areas was extracted and analyzed separately. For one malignant phyllodes tumor without heterologous differentiation (MPT9), DNA was extracted separately from two histologically distinct areas (Figure 1).
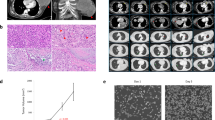
Representative images of malignant phyllodes tumors sequenced in this study. (a–c) This malignant phyllodes tumor (MPT8) shows well-developed biphasic fibroepithelial architecture (a) with predominantly non-heterologous spindled stromal tumor cells (a, b) and multiple scattered areas of liposarcomatous differentiation (a top left, c). Liposarcomatous differentiation is characterized by pleomorphic atypical lipoblasts and other tumor cells with variably sized fat droplets and large hyperchromatic nuclei (c). (d–e) MPT7 shows well-developed biphasic fibroepithelial architecture throughout the tumor, with stroma composed of intimately admixed spindled tumor cells and tumor cells with liposarcomatous differentiation. In focal areas, only few scattered lipoblastic tumor cells are present in a predominant background of spindled tumor cells (e), whereas most areas demonstrate a predominance of liposarcomatous tumor cells (f). (g–i) MPT5 shows areas of well-developed fibroepithelial growth (g, left) with typical spindled tumor cells (h), whereas other areas (g, right) demonstrate stromal overgrowth by plumper tumor cells with moderate-to-high nuclear grade arranged in linear and poorly formed acinar-like formations in a myxoid matrix (i). The histological features are similar to those of extraskeletal myxoid chondrosarcoma. (j–l) MPT9 demonstrates histologically distinct areas without heterologous elements. In some areas of biphasic fibroepithelial growth (j), the spindled tumor cells are less densely packed and show either mild or moderate nuclear atypia (moderate atypia in k). Other areas demonstrate stromal overgrowth by highly cellular proliferations of spindled and giant pleomorphic tumor cells with severe nuclear atypia (l). (m–n), MPT10. Focal areas of residual fibroepithelial growth (m, right) are present, but most of the tumor demonstrates diffuse stromal overgrowth (m, left) by spindled tumor cells with moderate-to-severe atypia, numerous mitotic figures and foci of necrosis (n). (hematoxylin and eosin, all images).
PCR and Sanger DNA Sequencing
Sanger sequencing of a 163 base pair fragment of the TERT promoter spanning the region of hotspot mutations was performed for borderline and benign phyllodes tumors, as previously described.33 Regions of interest were macrodissected on unstained slides, and genomic DNA was isolated using the QIAamp DNA FFPE Tissue Kit, following the manufacturer’s instructions (Qiagen, Valencia, CA, USA). PCR was performed with MyTaq Mix (Bioline) using the following primers: 5′-CAGCGCTGCCTGAAACTC-3′ (TERTp-F) and 5′-GTCCTGCCCCTTCACCTT-3′ (TERTp-R). Following treatment with ExoSAP-IT (Affymetrix, Cleveland, OH, USA), reaction products were diluted and bidirectionally sequenced by Sanger methodology at Elim Biopharmaceuticals (Hayward, CA, USA) using the same primers. Sequences were analyzed using Bioedit and CodonAligner software.
Tissue Microarray Construction
Tissue microarrays were constructed from the 10 malignant phyllodes tumors using three separate 2-mm punch biopsy tissue cores from each tumor area to create triplicates for analysis.
Immunohistochemistry
Immunohistochemistry was performed on tissue microarrays (EGFR, p53, p16, Rb, PTEN, βcatenin) and whole tissue sections (EGFR, BRAF V600E). The following antibodies were used: EGFR (5B7, undiluted (Roche, Ventana Medical Systems, Tucson, AZ, USA), PTEN (6H2.1, 1:200 anti-background (DAKO, Carpinteria, CA, USA), p16 (E6H4, prediluted (Roche, Ventana Medical Systems)), p53 (D0-7, 1:100 (Vector laboratories, Burlingame, CA, USA)), Rb (G3-245, 1:100 (BD Biosciences, San Jose, CA, USA)), BRAF V600E (VE1, prediluted (Roche, Ventana Medical Systems)), and βcatenin (Clone 14, 1:400 (BD Biosciences, Franklin Lakes, NJ, USA)). Antigen retrieval was as follows: for EGFR, CC1 (Roche, Ventana Medical Systems); for PTEN and Rb, Bond epitope retrieval solution 2 (Leica Biosystems, Buffalo Grove, IL, USA); for p16, p53 and βcatenin, Bond epitope retrieval solution 1; and for BRAF V600E, Ventana kit.
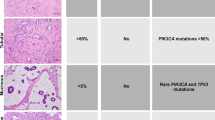
For p53, diffuse (≥90%) strong homogeneous nuclear staining was considered positive, and no nuclear staining was considered negative. Patchy heterogeneous nuclear staining of weak-to-moderate intensity was considered wild-type pattern.56, 57, 58, 59 For p16 and Rb, positive staining was defined as diffuse (≥80%) moderate or strong nuclear/cytoplasmic (p16) or nuclear only (Rb) staining, and lack of staining was considered negative. Cases without positive or negative p16 or Rb staining were classified as decreased staining, which, for p16, was <20% weak-to-moderate staining in all cases. For PTEN, positive staining was defined as ≥90% nuclear/cytoplasmic staining to reflect staining in nearly all tumor cells, with decreased staining defined as <90% and/or weak-to-moderate staining only.60, 61 EGFR staining was scored as 0 (no staining), 1+ (weak membranous/cytoplasmic), 2+ (moderate membranous/cytoplasmic) or 3+ (strong membranous/cytoplasmic) and classified as diffuse (≥90%), non-diffuse (<90%) or focal (non-diffuse staining in 1 of the 3 cores). βcatenin was scored as positive (discernible nuclear staining) or negative (indiscernible nuclear staining).
Statistics
Statistical analysis of differences between groups was performed using Fisher exact test. Differences between means were calculated using t-test. Results were considered statistically significant if P<0.05.
Results
Histological and Clinical Features of Malignant Phyllodes Tumors
The clinicopathological features of the primary malignant phyllodes tumors analyzed in this study are shown in Table 1. Six of the 10 tumors showed areas of malignant heterologous differentiation, with 5 of these demonstrating liposarcomatous differentiation (MPT1, MPT2, MPT6-8) and one showing myxoid chondrosarcoma-like differentiation (MPT5) (Figure 1 and Supplementary Table S1). In four cases with liposarcomatous differentiation and the sole case with myxoid chondrosarcoma-like elements, heterologous components were focal or patchy and spatially distinct from non-heterologous elements, whereas one liposarcomatous tumor (MPT7) demonstrated diffuse liposarcomatous differentiation in a biphasic fibroepithelial background (Figure 1). One patient was known to have Li–Fraumeni syndrome and had a germline TP53c.202G>T (p.68*) mutation (MPT9). No germline mutations were known or detected in the other nine patients. Three patients died of disease, which was preceded in all cases by at least one distant metastasis (MPT3, MPT6 and MPT10) and by local breast recurrence and axillary lymph node metastasis in one patient (MPT3) (Table 1).
The Spectrum of Genomic Aberrations in Malignant Phyllodes Tumors
Sequencing revealed 47 somatic non-synonymous mutations or insertions/deletions in 33 genes, 7 of which were recurrently mutated (pathogenic mutations shown in Figure 2, additional mutations of uncertain pathogenicity in Supplementary Figure S1). The total number of non-synonymous single-nucleotide variants and insertions/deletions in UCSF500 panel genes ranged from 2 per tumor (MPT8 and MPT10) to 10 per tumor (MPT6), and the number of known pathogenic mutations ranged from 1 (MPT10) to 5 (MPT3 and MPT4) (Figure 2 and Supplementary Figure S1). The mean target tumor coverage was 414.3 (range 166.2–927.6; Supplementary Table S2).
Genomic aberrations in malignant phyllodes tumors. Summary of pathogenic genomic aberrations identified in malignant phyllodes tumors. For additional aberrations of uncertain pathogenicity, see Supplementary Figure S1.
Recurrently mutated genes included TERT promoter (6/10), TP53 (4/10), MED12 (3/10), PIK3CA (3/10), FGFR1 (2/10), SETD2 (2/10) and KMT2D/MLL2 (2/10) (Figure 2 and Supplementary Table S3).
Eight (80%) of the 10 malignant phyllodes tumors showed mutations in genes involved in FGFR or EGFR signaling through phosphatidyl inositol-3 kinase (PI-3K) or RAS pathways. These included mutually exclusive activating hotspot mutations of FGFR1 (MPT2 and MPT4), PIK3CA (MPT5–MPT7) and BRAF (MPT3), EGFR amplification (MPT8) and an inactivating frameshift mutation of TSC2 (p.V1475fs) with associated loss of heterozygosity (MPT9). MPT4 had homozygous PTEN deletion in addition to the hotspot FGFR1 mutation (Figures 2, 3, 4). PIK3CA mutations were in hotspots in the helical (c.1624G>A, p.E542K) and kinase (c.3140A>G, H1047R) domains, known to result in constitutive activation and considered pathogenic.62 FGFR1 mutations (c.1638C>A and c.1966A>G) were both located in the protein kinase domain (p.N546K and p.K656E). Both FGFR1 mutations have been previously described in a variety of central nervous system neoplasms and were shown to be activating in vitro.63, 64, 65, 66, 67 MPT3 with BRAF p.V600E mutation was particularly aggressive despite adjuvant chemotherapy, with local recurrence showing skeletal muscle invasion ~6 months after diagnosis and death from disease within ~14 months (Table 1). Immunohistochemical staining using an antibody specific for BRAF V600E confirmed expression of the mutant protein in the tumor cells (Figure 3). Amplification of a narrow region on chromosome 7p including EGFR was only present in the non-heterologous component and not in the liposarcomatous area of MPT8. Immunohistochemistry confirmed strong EGFR expression in non-heterologous areas only (Figure 4).
Malignant phyllodes tumor with BRAF V600E mutation. (a–d) Histological sections of this tumor (MPT3) demonstrate areas with well-developed, moderately cellular biphasic fibroepithelial growth (a, c), as well as areas with stromal overgrowth of pleomorphic spindled tumor cells with scattered osteoclast-like giant cells and distinctly infiltrative border (b, d). No osteoid or bone formation is present. (e) Screenshot of Integrated Genome Viewer, showing somatic BRAF c.1799T>A (p.V600E) mutation in the tumor sample. (f) Immunohistochemical staining for BRAF V600E confirmed expression of the mutant protein in the tumor cells.
Heterogeneous EGFR amplification in malignant phyllodes tumor. (a–c) MPT8 demonstrates EGFR amplification in spindled tumor cells without heterologous differentiation. (a) Copy number analysis reveals EGFR amplification in non-heterologous tumor component. (b) Non-heterologous component (hematoxylin and eosin). (c) Immunohistochemical staining confirms strong membranous and cytoplasmic overexpression of EGFR. (d–f) Foci of liposarcomatous differentiation lack EGFR amplification. (d) Copy number analysis of sequenced liposarcomatous tumor component shows lack of EGFR amplification. (e) Liposarcomatous component (hematoxylin and eosin). (f) Immunohistochemical staining of liposarcomatous area shows lack of EGFR overexpression, with only weak background staining.
All TERT mutations (chr5:g.1,295,228 C>T, c. -124C>T, MPT1-6) were in one of the two previously described hotspots in the promoter.33, 68 One additional tumor (MPT8) revealed high-level TERT amplification (Supplementary Figures S2 and S3). Given the high recurrence of TERT aberrations in malignant phyllodes tumors (7/10, 70%), we sought to determine the frequency of TERT promoter mutations across the histological spectrum of phyllodes tumors. TERT promoter mutations were identified in 4 of the 12 (33%) borderline phyllodes tumors (P=0.391 vs MPT) and 0 of the 8 (0%) benign phyllodes tumors (P=0.013 vs malignant phyllodes tumors) by Sanger sequencing. As in malignant phyllodes tumors, all TERT promoter mutations in borderline tumors were at the previously described hotspot (chr5:g.1,295,228 C>T, c. -124C>T, Supplementary Figure S3). Malignant phyllodes tumors with TERT alterations demonstrated fewer large-scale chromosomal alterations (mean=3.6) than malignant phyllodes tumors lacking TERT aberrations (mean=16; P=0.006).
MED12 mutations were found in three malignant phyllodes tumors, and all affected exon 2. Two tumors (MPT1 and MPT2) had non-synonymous mutations (c.130G>C) at hotspot codon 44 (G44R) and one tumor (MPT3) had an in-frame deletion (c.117_161del, p.39_54del).21, 32, 33, 34, 35, 36, 37, 38, 39, 69, 70, 71 All malignant phyllodes tumors with MED12 mutations also had TERT promoter mutations, whereas 3 of the 6 malignant phyllodes tumors with TERT promoter mutations and the 1 with TERT amplification lacked MED12 mutations (Figure 2).
The histone methyltransferases SETD2 and KMT2D were mutated in two tumors each, and these were non-overlapping mutations. MPT2 and MPT4 had two SETD2 mutations each, though the mutations were too far apart to determine whether they affected different alleles. MPT2 had a SETD2 frameshift mutation (c.4670_4671del, p.T1557fs) and an in-frame deletion (c.5470_5472del, p.1824_1824del). MPT4 had a SETD2 stopgain (c.2120T>G, p.L707*) and frameshift deletion (c.3965delG, p.R1322fs) (Figure 2). Interestingly, both tumors with SETD2 mutations also had FGFR1 mutations and TERT promoter mutations. Mutations in KMT2D consisted of one stopgain (c.12304C>T, p.Q4102*; MPT3) and one missense (c.16385A>G, p. D5462G; MPT6) (Figure 2 and Supplementary Table S3). Homozygous deletion of the DNA methyltransferase DNMT3A was also identified in the tumor (MPT6) with the missense KMT2D mutation.
Aberrations were identified in common tumor-suppressor pathways, including non-overlapping mutations in TP53 (4 tumors), CDKN2A (2 tumors), PTEN (1 tumor) and CHEK2 (1 tumor) (Figure 2). TP53 mutations (MPT5, MPT7, MPT8 and MPT9) were in each case associated with loss of heterozygosity. Tumors with TP53 missense mutations predicted to be pathogenic showed strong diffuse p53 expression, while tumors with frameshift deletions (c.626_627del, p.R209fs) or stopgain mutations (c.202G>T, p.E68*) showed no immunoreactivity for p53 (Supplementary Table S4). Two of the tumors with TP53 mutations also showed RB1 alteration. MPT8 had a frameshift RB1 mutation (c.998_1014del, p.A333fs) in association with loss of heterozygosity in one tumor, and MPT9 had homozygous loss of RB1. Both of these tumors had reduced Rb protein expression as compared with those without RB1 alterations (Supplementary Table S4). Homozygous loss of chromosome 9p21 including the CDKN2A locus was identified in two tumors (MPT3 and MPT6). CDKN2A deletions correlated with p16 immunohistochemistry, with all tumors that harbored hemizygous or homozygous CDKN2A deletions showing either decreased or absent p16 expression, compared with retained diffuse expression in tumors without deletions of CDKN2A (Supplementary Table S4). One tumor (MPT4) showed homozygous PTEN deletion. As was the case for p53, p16 and Rb, genomic aberration of PTEN was associated with reduced protein expression by immunohistochemistry (Supplementary Table S4). One tumor showed a somatic nonsense mutation (p.Y297*) in the DNA damage checkpoint kinase and tumor-suppressor CHEK2, which was associated with loss of heterozygosity. Another tumor (MPT5) showed nonsense mutation and loss of heterozygosity of the CDKN1B tumor suppressor (encoding p27Kip1).
Genomic aberrations specific to malignant heterologous differentiation were not identified. C DK4/MDM2 amplifications characteristic of classic liposarcoma at other sites were not present in any of the tumors.
Intratumoral Heterogeneity in Malignant Phyllodes Tumors With and Without Heterologous Differentiation
Copy number analysis of separate areas in 7 of the 10 malignant phyllodes tumors revealed significant intratumoral genomic heterogeneity in tumors with and without heterologous differentiation. Copy number alterations that were shared between the two separately sequenced components were found in all tumors. However, additional private copy number changes present in only one of the sequenced components were also identified in all cases, suggestive of divergent evolution (Figures 5 and 6,Supplementary Tables S4 and S5). The vast majority of mutations (43/47, 91.5%), on the other hand, were identified in both sequenced areas (Figure 2).
Intratumoral heterogeneity in malignant phyllodes tumors with liposarcomatous differentiation. (a–d) Genome-wide copy number analysis of malignant phyllodes tumors with liposarcomatous differentiation and separately analyzed non-heterologous and liposarcomatous components (MPT1, MPT2, MPT6 and MPT8). Note that more private copy number alterations are present in non-heterologous compared with liposarcomatous components of the paired tumor samples in all cases, in addition to copy number alterations shared between the two components. Thick black arrows—large-scale (chromosome or whole-chromosome arm level) copy number alterations shared between the two components, thin black arrows—smaller-scale (sub-chromosomal arm) copy number alterations shared between the two components, thick red arrows—large-scale (chromosome or whole-chromosome arm level) copy number alterations private to only one of the components, thin red arrows—smaller-scale (sub-chromosomal arm) copy number alterations private to only one of the components. In (d), the long stippled red arrow highlights EGFR amplification (7p11.2), which is present only in the non-heterologous component of this tumor, and the long stippled grey arrows highlight TERT amplification (5p15.33), which is shared between the components. (e) Genome-wide copy number analysis of MPT7 with two separately analyzed liposarcomatous components. Annotations are the same as described for panels (a–d). Copy number changes are shown as matched tumor minus normal log2 ratios. Chromosomes are organized from 1 (left) to X (right).
Intratumoral heterogeneity in malignant phyllodes tumors. (a) Genome-wide copy number analysis of separately analyzed non-heterologous and myxoid chondrosarcoma-like components of MPT5. Note the presence of numerous copy number alterations private to the myxoid chondrosarcoma-like area, in addition to copy number alterations shared between the components. (b) Genome-wide copy number analysis of two separately analyzed non-heterologous components of MPT9, one of which is histologically less atypical (see Figure 1k), and the other of which is highly atypical with pleomorphic giant cells (see Figure 1l). Note the presence of numerous copy number alterations private to the histologically more atypical area, in addition to copy number alterations shared between the components. Thick black arrows—large-scale (chromosome or whole-chromosome arm level) copy number alterations shared between the two components, thin black arrows—smaller-scale (sub-chromosomal arm) copy number alterations shared between the two components, thick red arrows—large-scale (chromosome or whole-chromosome arm level) copy number alterations private to only one of the components, thin red arrows—smaller-scale (sub-chromosomal arm) copy number alterations private to only one of the components. Copy number changes are shown as matched tumor minus normal log2 ratios. Chromosomes are organized from 1 (left) to X (right).
All tumors with liposarcomatous differentiation in which non-heterologous and heterologous components could be evaluated separately (n=4) showed increased private chromosomal changes in the non-heterologous areas compared with liposarcomatous areas. Indeed, only one malignant phyllodes tumor with liposarcomatous differentiation (MPT1) showed private large-scale chromosomal changes (9p loss and 22 loss) in the liposarcomatous component (Figure 5 and Supplementary Table S6). Additional evidence of divergent evolution in tumors with liposarcomatous differentiation included the presence of high-level EGFR amplification in the non-heterologous but not the liposarcomatous component of MPT8 (Figures 2 and 3). The fifth malignant phyllodes tumor with liposarcomatous differentiation (MPT7) showed diffuse liposarcoma throughout the tumor, albeit with differing histological features (Figures 1d–f). No differences in large-scale copy number changes were identified between separately analyzed liposarcomatous areas (Figure 5,Supplementary Table S6). Of the four malignant phyllodes tumors with liposarcomatous differentiation with separately evaluable heterologous and non-heterologous components, a missense KMT2D variant in the non-heterologous but not the liposarcomatous component of MPT6 was the only identified mutation that was not present in both sequenced components (Figure 2). The presence of this mutation in only the non-heterologous area could not be explained by differences in tumor cellularity, depth of coverage or mutant allele frequencies between the two components (Supplementary Table S3).
One malignant phyllodes tumor with liposarcomatous differentiation (MPT6) and aggressive behavior resulting in multiple metastases and death showed a striking complex pattern of numerous alternating gains and losses with interspersed copy number neutral areas affecting chromosome 3 and, to a lesser degree, chromosome 5, suggestive of chromothripsis.72, 73, 74 The aberrations were shared among the non-heterologous and liposarcomatous components, indicating that they arose before the different areas underwent independent evolution, with additional superimposed changes in the non-heterologous component (Supplementary Figure S4 and Figure 5).
Analysis of the malignant phyllodes tumor with myxoid chondrosarcoma-like differentiation (MPT5, Figure 1g–i) also revealed heterogeneity between non-heterologous and heterologous components. However, in contrast to tumors with liposarcomatous differentiation, the heterologous area of MPT5 showed more copy number alterations than the non-heterologous component (Figure 6a,Supplementary Table S6). In addition, TP53 mutation with loss of heterozygosity was identified only in the chondrosarcoma-like component of this tumor, whereas non-heterologous areas showed MDM4 and CDKN1B mutations not present in the chondrosarcoma-like areas (Figure 2a). The differences could not be explained by differences in tumor cellularity, depth of coverage or mutant allele frequencies between the components (Supplementary Table S3).
MPT9, which did not show heterologous differentiation, showed significantly more chromosomal aberrations in histologically more atypical areas compared with less atypical areas (Figures 1j–l and 6b, Supplementary Table S6). Interestingly, this tumor harbored different TSC2 mutations with loss of heterozygosity in the two components, a splice site variant (c.1716+1G>T) in the less atypical area and a TSC2 frameshift deletion (c.4425_4426del, p.V1475fs) in the more progressed area (Figure 2,Supplementary Table S3), suggestive of dysregulation of the same pathway by two different presumed loss-of-function alterations of TSC2.
Discussion
Recent studies have begun to shed light on the molecular pathology and progression of mammary fibroepithelial lesions,21, 24 but these have included only few malignant phyllodes tumors. Given the risk of a subset of malignant phyllodes tumors for metastasis and death with limited treatment options, a deeper knowledge of the genetic features of these poorly understood and potentially aggressive tumors is necessary in order to explore putative therapeutic approaches. This is the first study to utilize an extensive targeted sequencing panel focusing specifically on malignant phyllodes tumors. Using capture-based next-generation sequencing of 510 cancer-related genes, we have identified recurrently mutated oncogenic drivers in malignant phyllodes tumors, notably including frequent and potentially actionable aberrations in FGFR1 and associated PI-3K/RAS pathway genes, and highlighted the presence of intratumoral heterogeneity.
FGFR signaling is deregulated in many cancers and is emerging as a potential target for therapy.75 We describe for the first time FGFR1-activating hotspot mutations N546K and K656E in malignant phyllodes tumors. These mutations have been described in central nervous system tumors, such as pilocytic astrocytomas and rosette-forming glioneuronal tumors of the fourth ventricle, and have been shown to be activating in vitro, with N546K causing receptor autophosphorylation.63, 64, 66, 67, 76 Tsang et al.28 recently reported FGFR1 amplification in a malignant phyllodes tumor, but prior exome sequencing of 18 benign and borderline phyllodes tumors failed to identify FGFR1 aberrations in these tumors.24 Our results implicate FGFR1 as an oncogenic driver in malignant phyllodes tumors and suggest that these aberrations may be involved in tumor progression. Furthermore, together with other mutually exclusive potentially actionable activating mutations in PIK3CA and BRAF (V600E), EGFR amplification and inactivation of PTEN and TSC2, our data suggest a role for the FGFR PI-3K/RAS signaling axis in malignant phyllodes tumor development. Inactivation of the cell cycle inhibitor p27Kip (CDKN1B) is interesting in this context, as PI-3K downregulates p27, and FGFR1 stimulates proliferation of mesenchymal stem cells via p27 inactivation.77, 78 PIK3CA mutations have been previously reported in 5 of the 35 borderline phyllodes tumors and 1 malignant phyllodes tumor in a separate study, although only 2 of these were hotspots (both borderline tumors).24, 28 A case report of a malignant phyllodes tumor with activating NRAS (Q61L) mutation and phosphorylated AKT and mTOR has also been published, and NF1 mutations have been described in borderline and malignant phyllodes tumors.21, 24, 79 EGFR is amplified in a subset of malignant phyllodes tumors,14, 21, 22, 23, 24, 28 and IGF and PDGF signaling has been implicated in phyllodes tumor development, further implicating PI-3K/RAS signaling through receptor tyrosine kinases in tumor development and/or progression.21, 29, 31, 80 All malignant phyllodes tumors with pathogenic FGFR1/PI-3K/RAS pathway aberrations also harbored inactivation of a pathogenic tumor suppressor, indicative of cooperation between oncogenic and inactivated tumor-suppressor pathways.
TERT promoter mutations were highly recurrent in malignant phyllodes tumors (6 of the 10 cases), with another tumor showing TERT amplification. Whereas TERT is not expressed in most somatic tissues, TERT promoter mutations can lead to reactivation of TERT expression and may help explain increased telomerase activity, a hallmark of cancer.68, 81, 82, 83, 84, 85 The identification of TERT promoter mutations in a smaller subset (33%) of borderline phyllodes tumors but not in benign phyllodes tumors suggests that TERT promoter mutations and/or telomerase reactivation may have a role in phyllodes tumor progression. Telomerase expression or activity has not been described in phyllodes tumors, but absent or only low levels of expression and activity have been described in fibroadenomas.86, 87 A recent study identified TERT promoter mutations not only in borderline and malignant phyllodes tumors but also in benign phyllodes tumors and in few fibroadenomas.33 Differences from our findings may be at least partly due to differences in tumor grading and classification.88 Whereas TERT promoter mutations were closely correlated with MED12 mutations in that study, only half of malignant phyllodes tumors with TERT promoter mutations also had MED12 mutations in our series of malignant tumors.33
Recurrent hotspot MED12 exon 2 and RARA ligand-binding domain mutations in fibroepithelial lesions implicates dysregulated hormonal signaling in these tumors.21, 24, 32, 33, 34, 35, 36, 37, 38, 39, 71, 89, 90, 91 It is interesting in this context that we identified an ESR1 mutation (C530L) near a previously characterized cluster of activating hotspot mutations (L536Q, Y537S/N/C, D538G) in hormone-resistant breast cancer (Supplementary Figure S1).92, 93 The identification in malignant phyllodes tumors of only rare RARA mutations (~10%, 2 of the 20 combined cases in a previous study and ours), which are prevalent in benign and borderline phyllodes tumors (41% combined24), suggests the possibility that these aberrations may be selected against during tumor progression, but additional studies are warranted.
Pathogenic aberrations were also identified in the recurrently mutated histone methyltransferases SETD2 and KMT2D and in the DNA methyltransferase DNMT3A. SETD2 and KMT2D act as tumor suppressors in multiple cancers, including phyllodes tumors,94, 95, 96, 97 but were reported in one study to be rare in fibroadenomas.24 The results point to a role for altered chromatin regulation in phyllodes tumors. Studies addressing chromatin state alterations and transcriptional effects in phyllodes tumors may shed light on the role of chromatin modification in tumor development.
The molecular features of heterologous differentiation have not been investigated prior to this study. We found no genomic aberrations specific to liposarcomatous differentiation, including mutations or CDK4/MDM2 amplifications characteristic of soft tissue well-differentiated liposarcoma. Lyle et al. also recently failed to identify CDK4/MDM2 amplifications by fluorescence in situ hybridization in malignant phyllodes tumors with well-differentiated liposarcomatous components.98, 99 The results suggest that liposarcomatous differentiation in malignant phyllodes tumors develops through mechanisms distinct from that of true soft tissue liposarcomas, despite similar histological features.
Finally, our results highlight the presence of intratumoral heterogeneity with evidence of divergent evolution in malignant phyllodes tumors, which is predominantly reflected in copy number alterations. The presence of increased private copy number alterations in non-heterologous compared with liposarcomatous components of malignant phyllodes tumors with liposarcomatous differentiation indicates that liposarcomatous areas of these tumors are more chromosomally stable and that the differentiated liposarcomatous state may fundamentally support chromosomal stability.100 In our experience, liposarcomatous differentiation of metastatic malignant phyllodes tumor foci is rare, even in cases where the primary tumor demonstrates liposarcomatous differentiation, although the rarity of these tumors and especially their metastases precludes a systematic analysis. We speculate that this may be a reflection, at least in part, of the increased chromosomal stability in the liposarcomatous components compared with non-heterologous components of these tumors. In this study, both metastatic foci of the sole malignant tumor with liposarcomatous differentiation that metastasized (MPT6) were indeed composed of only non-heterologous stroma without liposarcomatous differentiation, although conclusions cannot be easily drawn from a single case. Future studies comparing the genomic features of non-heterologous and liposarcomatous components of a larger number of malignant phyllodes tumors with their metastases would be useful to further address this issue. Interestingly in this context, three of the four tumors with separately analyzed liposarcomatous and non-heterologous components (MPT1, MPT6 and MPT8) showed significantly more mitotic activity in non-heterologous areas than in liposarcomatous areas, suggesting that the histological features may broadly reflect the progressed genomic changes. A similar correlation was seen in separately sequenced components of the tumor with myxoid chondrosarcoma-like differentiation (MPT5) and in the tumor with separately sequenced non-heterologous but histologically distinct components (MPT9) (Supplementary Table S1). The identification of EGFR amplification in only the non-heterologous component of a malignant phyllodes tumor with liposarcomatous differentiation highlights potential clinical significance of intratumoral heterogeneity in these tumors.
In summary, we have identified mutually exclusive activating hotspot mutations in FGFR1 and related PI-3K/RAS signaling genes in malignant phyllodes tumors, which, in addition to other drivers, such as TERT promoter, TP53, MED12 and histone methyltransferases SETD2 and KMT2D, are implicated in tumor pathogenesis and/or progression. Genomic features specific to liposarcomatous differentiation were not identified, but heterologous and non-heterologous components of malignant phyllodes tumors show evidence of divergent evolution, which may have clinical implications with regards to clonal selection and tumor progression.
References
Tan PH, Tse G, Lee A et al. Fibroepithelial tumours. In: Lakhani SR, Ellis IO, Schnitt SJ et al. (eds). WHO Classification of Tumours of the Breast, 4th edn. International Agency for Research on Cancer (IARC): Lyon, France, 2012, pp 141–147.
Jacobs TW . Update on fibroepithelial lesions of the breast. Surg Pathol Clin 2009; 2: 301–317.
Tan PH, Thike AA, Tan WJ et al. Predicting clinical behaviour of breast phyllodes tumours: a nomogram based on histological criteria and surgical margins. J Clin Pathol 2012; 65: 69–76.
Kim S, Kim JY, Kim do H et al. Analysis of phyllodes tumor recurrence according to the histologic grade. Breast Cancer Res Treat 2013; 141: 353–363.
Karim RZ, Gerega SK, Yang YH et al. Phyllodes tumours of the breast: a clinicopathological analysis of 65 cases from a single institution. Breast 2009; 18: 165–170.
Onkendi EO, Jimenez RE, Spears GM et al. Surgical treatment of borderline and malignant phyllodes tumors: the effect of the extent of resection and tumor characteristics on patient outcome. Ann Surg Oncol 2014; 21: 3304–3309.
Ward RM, Evans HL . Cystosarcoma phyllodes. A clinicopathologic study of 26 cases. Cancer 1986; 58: 2282–2289.
Tan PH, Jayabaskar T, Chuah KL et al. Phyllodes tumors of the breast: the role of pathologic parameters. Am J Clin Pathol 2005; 123: 529–540.
Chaney AW, Pollack A, McNeese MD et al. Primary treatment of cystosarcoma phyllodes of the breast. Cancer 2000; 89: 1502–1511.
Telli ML, Horst KC, Guardino AE et al. Phyllodes tumors of the breast: natural history, diagnosis, and treatment. J Natl Compr Canc Netw 2007; 5: 324–330.
Guillot E, Couturaud B, Reyal F et al. Management of phyllodes breast tumors. Breast J 2011; 17: 129–137.
Khosravi-Shahi P . Management of non metastatic phyllodes tumors of the breast: review of the literature. Surg Oncol 2011; 20: e143–e148.
National Comprehensive Cancer Network (NCCN) Guidelines Version 1. 2015 Phyllodes Tumor. Fort Washington, PA, USA, 2015. Available from http://www.nccn.org/professionals/physician_gls/f_guidelines.asp.
Tan WJ, Lai JC, Thike AA et al. Novel genetic aberrations in breast phyllodes tumours: comparison between prognostically distinct groups. Breast Cancer Res Treat 2014; 145: 635–645.
Jee KJ, Gong G, Ahn SH et al. Gain in 1q is a common abnormality in phyllodes tumours of the breast. Anal Cell Pathol 2003; 25: 89–93.
Lae M, Vincent-Salomon A, Savignoni A et al. Phyllodes tumors of the breast segregate in two groups according to genetic criteria. Mod Pathol 2007; 20: 435–444.
Lu YJ, Birdsall S, Osin P et al. Phyllodes tumors of the breast analyzed by comparative genomic hybridization and association of increased 1q copy number with stromal overgrowth and recurrence. Genes Chromosomes Cancer 1997; 20: 275–281.
Lv S, Niu Y, Wei L et al. Chromosomal aberrations and genetic relations in benign, borderline and malignant phyllodes tumors of the breast: a comparative genomic hybridization study. Breast Cancer Res Treat 2008; 112: 411–418.
Ang MK, Ooi AS, Thike AA et al. Molecular classification of breast phyllodes tumors: validation of the histologic grading scheme and insights into malignant progression. Breast Cancer Res Treat 2011; 129: 319–329.
Jones AM, Mitter R, Springall R et al. A comprehensive genetic profile of phyllodes tumours of the breast detects important mutations, intra-tumoral genetic heterogeneity and new genetic changes on recurrence. J Pathol 2008; 214: 533–544.
Cani AK, Hovelson DH, McDaniel AS et al. Next-gen sequencing exposes frequent MED12 mutations and actionable therapeutic targets in phyllodes tumors. Mol Cancer Res 2015; 13: 613–619.
Kersting C, Kuijper A, Schmidt H et al. Amplifications of the epidermal growth factor receptor gene (egfr) are common in phyllodes tumors of the breast and are associated with tumor progression. Lab Invest 2006; 86: 54–61.
Tse GM, Lui PC, Vong JS et al. Increased epidermal growth factor receptor (EGFR) expression in malignant mammary phyllodes tumors. Breast Cancer Res Treat 2009; 114: 441–448.
Tan J, Ong CK, Lim WK et al. Genomic landscapes of breast fibroepithelial tumors. Nat Genet 2015; 47: 1341–1345.
Feakins RM, Mulcahy HE, Nickols CD et al. p53 expression in phyllodes tumours is associated with histological features of malignancy but does not predict outcome. Histopathology 1999; 35: 162–169.
Cimino-Mathews A, Hicks JL, Sharma R et al. A subset of malignant phyllodes tumors harbors alterations in the Rb/p16 pathway. Hum Pathol 2013; 44: 2494–2500.
Gatalica Z, Finkelstein S, Lucio E et al. p53 protein expression and gene mutation in phyllodes tumors of the breast. Pathol Res Pract 2001; 197: 183–187.
Tsang JY, Go EM, Tse GM . Identification of clinically relevant alterations in phyllodes tumor of the breast by amplicon-based next-generation sequencing. Breast Cancer Res Treat 2015; 151: 717–719.
Sawyer EJ, Hanby AM, Poulsom R et al. Beta-catenin abnormalities and associated insulin-like growth factor overexpression are important in phyllodes tumours and fibroadenomas of the breast. J Pathol 2003; 200: 627–632.
Sawyer EJ, Hanby AM, Rowan AJ et al. The Wnt pathway, epithelial-stromal interactions, and malignant progression in phyllodes tumours. J Pathol 2002; 196: 437–444.
Feakins RM, Wells CA, Young KA et al. Platelet-derived growth factor expression in phyllodes tumors and fibroadenomas of the breast. Hum Pathol 2000; 31: 1214–1222.
Ng CC, Tan J, Ong CK et al. MED12 is frequently mutated in breast phyllodes tumours: a study of 112 cases. J Clin Pathol 2015; 68: 685–691.
Yoshida M, Ogawa R, Yoshida H et al. TERT promoter mutations are frequent and show association with MED12 mutations in phyllodes tumors of the breast. Br J Cancer 2015; 113: 1244–1248.
Yoshida M, Sekine S, Ogawa R et al. Frequent MED12 mutations in phyllodes tumours of the breast. Br J Cancer 2015; 112: 1703–1708.
Mishima C, Kagara N, Tanei T et al. Mutational analysis of MED12 in fibroadenomas and phyllodes tumors of the breast by means of targeted next-generation sequencing. Breast Cancer Res Treat 2015; 152: 305–312.
Lien HC, Huang CS, Yang YW et al. Mutational analysis of MED12 exon 2 in a spectrum of fibroepithelial tumours of the breast: implications for pathogenesis and histogenesis. Histopathology 2015; 68: 433–441.
Lim WK, Ong CK, Tan J et al. Exome sequencing identifies highly recurrent MED12 somatic mutations in breast fibroadenoma. Nat Genet 2014; 46: 877–880.
Makinen N, Mehine M, Tolvanen J et al. MED12, the mediator complex subunit 12 gene, is mutated at high frequency in uterine leiomyomas. Science 2011; 334: 252–255.
Perot G, Croce S, Ribeiro A et al. MED12 alterations in both human benign and malignant uterine soft tissue tumors. PLoS One 2012; 7: e40015.
Hawkins RE, Schofield JB, Fisher C et al. The clinical and histologic criteria that predict metastases from cystosarcoma phyllodes. Cancer 1992; 69: 141–147.
Cohn-Cedermark G, Rutqvist LE, Rosendahl I et al. Prognostic factors in cystosarcoma phyllodes. A clinicopathologic study of 77 patients. Cancer 1991; 68: 2017–2022.
McGranahan N, Swanton C . Biological and therapeutic impact of intratumor heterogeneity in cancer evolution. Cancer Cell 2015; 27: 15–26.
Michor F, Polyak K . The origins and implications of intratumor heterogeneity. Cancer Prev Res (Phila) 2010; 3: 1361–1364.
Karim RZ, O'Toole SA, Scolyer RA et al. Recent insights into the molecular pathogenesis of mammary phyllodes tumours. J Clin Pathol 2013; 66: 496–505.
Li H, Durbin R . Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 2010; 26: 589–595.
Li H, Handsaker B, Wysoker A et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009; 25: 2078–2079.
Yang H, Wang K . Genomic variant annotation and prioritization with ANNOVAR and wANNOVAR. Nat Protoc 2015; 10: 1556–1566.
Rausch T, Zichner T, Schlattl A et al. DELLY: structural variant discovery by integrated paired-end and split-read analysis. Bioinformatics 2012; 28: i333–i339.
Picard: A set of tools (in Java) for working with next generation sequencing data in the BAM. Broad Institute. Available from http://broadinstitute.github.io/picard.
Garrison E, Marth G . Haplotype-based variant detection from short-read sequencing. arXiv:1207.3907 [q-bio.GN]. Available from: http://arxiv.org/abs/1207.3907.
McKenna A, Hanna M, Banks E et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 2010; 20: 1297–1303.
DePristo MA, Banks E, Poplin R et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat Genet 2011; 43: 491–498.
Van der Auwera GA, Carneiro MO, Hartl C et al. From FastQ data to high confidence variant calls: the Genome Analysis Toolkit best practices pipeline. Curr Protoc Bioinformatics 2013; 43: 11.10.1–33.
Ye K, Schulz MH, Long Q et al. Pindel: a pattern growth approach to detect break points of large deletions and medium sized insertions from paired-end short reads. Bioinformatics 2009; 25: 2865–2871.
Talevich E, Shain AH, Boton T, Bastian BC . CNVkit: copy number detection and visualization for targeted sequencing using off-target reads. PLoS Comput Biol 2016; 12: e1004873.
Yemelyanova A, Vang R, Kshirsagar M et al. Immunohistochemical staining patterns of p53 can serve as a surrogate marker for TP53 mutations in ovarian carcinoma: an immunohistochemical and nucleotide sequencing analysis. Mod Pathol 2011; 24: 1248–1253.
Kaye PV, Haider SA, James PD et al. Novel staining pattern of p53 in Barrett's dysplasia—the absent pattern. Histopathology 2010; 57: 933–935.
Ando K, Oki E, Saeki H et al. Discrimination of p53 immunohistochemistry-positive tumors by its staining pattern in gastric cancer. Cancer Med 2015; 4: 75–83.
McCluggage WG, Soslow RA, Gilks CB . Patterns of p53 immunoreactivity in endometrial carcinomas: 'all or nothing' staining is of importance. Histopathology 2011; 59: 786–788.
Garg K, Broaddus RR, Soslow RA et al. Pathologic scoring of PTEN immunohistochemistry in endometrial carcinoma is highly reproducible. Int J Gynecol Pathol 2012; 31: 48–56.
Djordjevic B, Hennessy BT, Li J et al. Clinical assessment of PTEN loss in endometrial carcinoma: immunohistochemistry outperforms gene sequencing. Mod Pathol 2012; 25: 699–708.
Karakas B, Bachman KE, Park BH . Mutation of the PIK3CA oncogene in human cancers. Br J Cancer 2006; 94: 455–459.
Jones DT, Hutter B, Jager N et al. Recurrent somatic alterations of FGFR1 and NTRK2 in pilocytic astrocytoma. Nat Genet 2013; 45: 927–932.
Lew ED, Furdui CM, Anderson KS et al. The precise sequence of FGF receptor autophosphorylation is kinetically driven and is disrupted by oncogenic mutations. Sci Signal 2009; 2: ra6.
Yoon K, Nery S, Rutlin ML et al. Fibroblast growth factor receptor signaling promotes radial glial identity and interacts with Notch1 signaling in telencephalic progenitors. J Neurosci 2004; 24: 9497–9506.
Gessi M, Moneim YA, Hammes J et al. FGFR1 mutations in Rosette-forming glioneuronal tumors of the fourth ventricle. J Neuropathol Exp Neurol 2014; 73: 580–584.
Gessi M, Abdel Moneim Y, Hammes J et al. FGFR1 N546K mutation in a case of papillary glioneuronal tumor (PGNT). Acta Neuropathol 2014; 127: 935–936.
Heidenreich B, Rachakonda PS, Hemminki K et al. TERT promoter mutations in cancer development. Curr Opin Genet Dev 2014; 24: 30–37.
Je EM, Kim MR, Min KO et al. Mutational analysis of MED12 exon 2 in uterine leiomyoma and other common tumors. Int J Cancer 2012; 131: E1044–E1047.
McGuire MM, Yatsenko A, Hoffner L et al. Whole exome sequencing in a random sample of North American women with leiomyomas identifies MED12 mutations in majority of uterine leiomyomas. PLoS One 2012; 7: e33251.
Pfarr N, Kriegsmann M, Sinn P et al. Distribution of MED12 mutations in fibroadenomas and phyllodes tumors of the breast—implications for tumor biology and pathological diagnosis. Genes Chromosomes Cancer 2015; 54: 444–452.
Przybytkowski E, Lenkiewicz E, Barrett MT et al. Chromosome-breakage genomic instability and chromothripsis in breast cancer. BMC Genomics 2014; 15: 579.
Stephens PJ, Greenman CD, Fu B et al. Massive genomic rearrangement acquired in a single catastrophic event during cancer development. Cell 2011; 144: 27–40.
Kinsella M, Patel A, Bafna V . The elusive evidence for chromothripsis. Nucleic Acids Res 2014; 42: 8231–8242.
Dieci MV, Arnedos M, Andre F et al. Fibroblast growth factor receptor inhibitors as a cancer treatment: from a biologic rationale to medical perspectives. Cancer Discov 2013; 3: 264–279.
Becker AP, Scapulatempo-Neto C, Carloni AC et al. KIAA1549: BRAF gene fusion and FGFR1 hotspot mutations are prognostic factors in pilocytic astrocytomas. J Neuropathol Exp Neurol 2015; 74: 743–754.
Dombrowski C, Helledie T, Ling L et al. FGFR1 signaling stimulates proliferation of human mesenchymal stem cells by inhibiting the cyclin-dependent kinase inhibitors p21(Waf1) and p27(Kip1). Stem Cells 2013; 31: 2724–2736.
Nakamura N, Ramaswamy S, Vazquez F et al. Forkhead transcription factors are critical effectors of cell death and cell cycle arrest downstream of PTEN. Mol Cell Biol 2000; 20: 8969–8982.
Jardim DL, Conley A, Subbiah V . Comprehensive characterization of malignant phyllodes tumor by whole genomic and proteomic analysis: biological implications for targeted therapy opportunities. Orphanet J Rare Dis 2013; 8: 112.
Liu C, Zhang Z, Tang H et al. Crosstalk between IGF-1R and other tumor promoting pathways. Curr Pharm Des 2014; 20: 2912–2921.
Borah S, Xi L, Zaug AJ et al. Cancer. TERT promoter mutations and telomerase reactivation in urothelial cancer. Science 2015; 347: 1006–1010.
Huang FW, Hodis E, Xu MJ et al. Highly recurrent TERT promoter mutations in human melanoma. Science 2013; 339: 957–959.
Hanahan D, Weinberg RA . Hallmarks of cancer: the next generation. Cell 2011; 144: 646–674.
Killela PJ, Reitman ZJ, Jiao Y et al. TERT promoter mutations occur frequently in gliomas and a subset of tumors derived from cells with low rates of self-renewal. Proc Natl Acad Sci USA 2013; 110: 6021–6026.
Horn S, Figl A, Rachakonda PS et al. TERT promoter mutations in familial and sporadic melanoma. Science 2013; 339: 959–961.
Hiyama E, Gollahon L, Kataoka T et al. Telomerase activity in human breast tumors. J Natl Cancer Inst 1996; 88: 116–122.
Hiyama E, Hiyama K, Yokoyama T et al. Immunohistochemical detection of telomerase (hTERT) protein in human cancer tissues and a subset of cells in normal tissues. Neoplasia 2001; 3: 17–26.
Lawton TJ, Acs G, Argani P et al. Interobserver variability by pathologists in the distinction between cellular fibroadenomas and phyllodes tumors. Int J Surg Pathol 2014; 22: 695–698.
Kang YK, Guermah M, Yuan CX et al. The TRAP/Mediator coactivator complex interacts directly with estrogen receptors alpha and beta through the TRAP220 subunit and directly enhances estrogen receptor function in vitro. Proc Natl Acad Sci USA 2002; 99: 2642–2647.
Ross-Innes CS, Stark R, Holmes KA et al. Cooperative interaction between retinoic acid receptor-alpha and estrogen receptor in breast cancer. Genes Dev 2010; 24: 171–182.
Markowski DN, Bartnitzke S, Loning T et al. MED12 mutations in uterine fibroids—their relationship to cytogenetic subgroups. Int J Cancer 2012; 131: 1528–1536.
Robinson DR, Wu YM, Vats P et al. Activating ESR1 mutations in hormone-resistant metastatic breast cancer. Nat Genet 2013; 45: 1446–1451.
Toy W, Shen Y, Won H et al. ESR1 ligand-binding domain mutations in hormone-resistant breast cancer. Nat Genet 2013; 45: 1439–1445.
Ortega-Molina A, Boss IW, Canela A et al. The histone lysine methyltransferase KMT2D sustains a gene expression program that represses B cell lymphoma development. Nat Med 2015; 21: 1199–1208.
Ciriello G, Gatza ML, Beck AH et al. Comprehensive molecular portraits of invasive lobular breast cancer. Cell 2015; 163: 506–519.
Duns G, van den Berg E, van Duivenbode I et al. Histone methyltransferase gene SETD2 is a novel tumor suppressor gene in clear cell renal cell carcinoma. Cancer Res 2010; 70: 4287–4291.
Fontebasso AM, Schwartzentruber J, Khuong-Quang DA et al. Mutations in SETD2 and genes affecting histone H3K36 methylation target hemispheric high-grade gliomas. Acta Neuropathol 2013; 125: 659–669.
Pilotti S, Della Torre G, Lavarino C et al. Molecular abnormalities in liposarcoma: role of MDM2 and CDK4-containing amplicons at 12q13-22. J Pathol 1998; 185: 188–190.
Lyle PL, Bridge JA, Simpson JF et al. Liposarcomatous differentiation in malignant phyllodes tumors is unassociated with MDM2 or CDK4 amplification. Histopathology 2015; 68: 1040–1045.
Horvai AE, DeVries S, Roy R et al. Similarity in genetic alterations between paired well-differentiated and dedifferentiated components of dedifferentiated liposarcoma. Mod Pathol 2009; 22: 1477–1488.
Acknowledgements
This study was funded by the Department of Pathology of the University of California San Francisco.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Modern Pathology website
Supplementary information
FIGURE S2. TERT Amplification in Malignant Phyllodes Tumor
. A-B, MPT8, non-heterologous component (hematoxylin and eosin). C, Copy number analysis of both non-heterologous and liposarcomatous components showed TERT amplification. D, RNAScope analysis (performed using manufacturer’s protocol, probe HsTERT [605511], Advanced Cell Diagnostics, Hayward, CA) showed corresponding high TERT RNA levels in the tumor cells.
FIGURE S3
. TERT Promoter Mutations in Malignant and Borderline Phyllodes Tumors but not Benign Phyllodes Tumors. A, Representative image of malignant phyllodes tumor (MPT6; hematoxylin and eosin) and Integrated Genome Viewer screenshot showing c.-124C>T TERT promoter mutation. B, Representative image of borderline phyllodes tumor (hematoxylin and eosin) and Sanger sequencing trace showing c.-124C>T TERT promoter mutation. C, Representative image of benign phyllodes tumor (hematoxylin and eosin) and Sanger sequencing traces of wild-type TERT promoter in hotspot -146 and -124 positions. D, Summary of TERT promoter mutations in malignant, borderline and benign phyllodes tumors.
FIGURE S4. Localized Complex Pattern of Copy Number Alterations in Malignant Phyllodes Tumor
. A-C, Representative images of MPT6, which demonstrated large areas of stromal overgrowth (A), composed of mitotically active spindled tumor cells with moderate to marked nuclear atypia (B). Smaller scattered tumor foci showed liposarcomatous differentiation, marked by atypical lipoblasts and tumor cells with fat vacuoles (C) (hematoxylin and eosin, all images). D, Copy number analysis of chromosome 3, showing numerous copy number changes in liposarcomatous (left) and non-heterologous (right) components, comprised of a complex pattern of adjacent gains and losses with admixed copy neutral areas. Additional superimposed copy number losses were present in the non-heterologous component (red arrows, right), which were not present in the liposarcomatous component (red arrows, left). This complex pattern of copy number alterations was localized only to chromosome 3, and to a lesser extent chromosome 5, and was not present on other chromosomes (see also Figure 5).
FIGURE S5. Highly Recurrent Copy Number Changes in Malignant Phyllodes Tumors
. Graph showing UCSF500 genes involved highly recurrent copy number changes (present in ≤4 malignant tumors), organized by chromosome. Log2 ratios are subtracted tumor minus matched normal values. In cases with more than one tumor sample sequenced, all log2 ratios are shown. Numbers along the top indicate chromosomes.
Rights and permissions
About this article
Cite this article
Liu, SY., Joseph, N., Ravindranathan, A. et al. Genomic profiling of malignant phyllodes tumors reveals aberrations in FGFR1 and PI-3 kinase/RAS signaling pathways and provides insights into intratumoral heterogeneity. Mod Pathol 29, 1012–1027 (2016). https://doi.org/10.1038/modpathol.2016.97
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2016.97
This article is cited by
-
Spindle cell lesions of the breast: a diagnostic approach
Virchows Archiv (2022)
-
Systemic treatment in advanced phyllodes tumor of the breast: a multi-institutional European retrospective case-series analyses
Breast Cancer Research and Treatment (2022)
-
Droplet-digital PCR reveals frequent mutations in TERT promoter region in breast fibroadenomas and phyllodes tumours, irrespective of the presence of MED12 mutations
British Journal of Cancer (2021)
-
Morphologic and genetic heterogeneity in breast fibroepithelial lesions—a comprehensive mapping study
Modern Pathology (2020)
-
High expression of microRNA20b is associated with malignant clinicopathological features and poor prognosis in breast phyllodes tumor
International Journal of Clinical Oncology (2020)