Abstract
Conventional methods used for histologic classification and grading of endometrial cancer (EC) are not sufficient to predict lymph node metastases. microRNA signatures have recently been related to EC pathologic characteristics or prognosis. The aim of this study was to evaluate whether microRNA profiles of grade 1–2 endometrioid adenocarcinomas can be related to nodal status and used as a tool to adapt surgical staging in early-stage EC. microRNA expression was assessed in nine formalin-fixed paraffin-embedded (FFPE) EC primary tumors with positive lymph node and in 27 FFPE EC primary tumors with negative lymph node, matched for grade, stage, and lymphovascular space involvement status. A microarray analysis showed that there was more than a twofold significant difference in the expression of 12 microRNAs between the two groups. A quantitative reverse transcriptase–PCR assay was used to confirm these results: the expression levels of five microRNAs (microRNA-34c-5p, -375, -184, -34c-3p, and -34b-5p) were significantly lower in the EC primary tumor with positive lymph node compared with those with negative lymph node. A minimal P-value approach revealed that women with a microRNA-375-fold change <0.30 were more likely to have positive lymph node (n=8; 53.3%) compared with those with a microRNA-375-fold change >0.30 (n=1; 4.8%), P=0.001. Furthermore, women with a microRNA 184-fold change <0.30 were more likely to have positive lymph node (n=6; 60.0%) compared with those with a microRNA 184-fold change >0.30 (n=3; 11.5%), P=0.006. This is the first study investigating the relative expression of mature microRNA genes in early-stage grade 1–2 EC primary tumors according to the nodal status. This microRNA expression profile provides a potential basis for further study of the microRNA function in EC and could be used as a diagnostic tool for nodal status.
Similar content being viewed by others
Main
Endometrial cancer (EC) is the most common gynecologic tumor in women in developed countries. The highest estimated incidences in 2012 are in the USA and Canada (19.1/100 000) and northern (12.9/100 000) and western Europe (15.6/100 000).1, 2 Women are often diagnosed at an early stage (stage I of the International Federation of Gynaecology and Obstetrics (FIGO) classification), when the disease is still confined to the uterus, representing almost 75% of the cases. The most frequently occurring histologic subtype is endometrioid adenocarcinoma, which is a hormone-receptor-positive EC, with a good prognosis.3 Women can be classified with early-stage EC at low-, intermediate-, or high-risk for recurrence by a combination of histologic criteria (ie depth of myometrial invasion and histologic type and grade).4
Standard treatments for early-stage EC consist of primary hysterectomy and bilateral salpingo-oophorectomy. In this specific setting, many centers in the United States perform lymph node sampling, lymph node dissection, or sentinel lymph node mapping in many or most EC, showing that indications for lymph node dissections are not well established. European guidelines have recently been modified and lymphadenectomy is no longer recommended in women with low- (FIGO stage IA, grade 1 and grade 2, endometrioid adenocarcinoma), or intermediate- (FIGO stage IA, grade 3 type 1 EC; FIGO stage IB, grade 1 and grade 2, endometrioid adenocarcinoma) risk EC.4 However, several authors have demonstrated that such an approach may lead to under-treatment as 10% of women of low-risk and 15% of intermediate-risk EC have nodal metastases,5, 6 rates that can be higher in the case of lymphovascular space involvement (LVSI) on the primary tumor.7, 8, 9
It is now well established that conventional methods used for histologic classification and grading of EC are not sufficient to predict lymph node metastases. This is probably due to considerable heterogeneity within endometrioid histologic subtypes.10 Hence, additional tools including highly sensitive and specific molecular prognostic biomarkers are needed to select the women for whom complete surgical staging should be performed in order to better adapt adjuvant therapies.
microRNAs (miRNAs) are short length (~22 nucleotides), naturally existing RNAs functioning at the post-transcriptional level either by regulating mRNA degradation or by translating repression through binding of the 3′-untranslated regions (3′-UTRs) of mRNAs.11 The human genome encodes at least 1500 miRNAs, and it is estimated that more than 30% of all mRNA expression is regulated by miRNAs.12 They have been linked to a variety of physiologic and pathologic processes, including carcinogenesis and can act as metastatic activators or suppressors.13, 14 Recently, miRNA signatures have been related to the pathologic characteristics and prognosis of EC.15 However, there are few data about nodal status-associated miRNA regulation in EC, especially in women at low and intermediate risk. miRNAs, with their remarkable stability, can be studied in formalin-fixed paraffin-embedded (FFPE) specimens16 and are promising biomarkers for tumor staging.
The aim of this study was thus to evaluate whether miRNA profiles of grade 1–2 endometrioid adenocarcinomas can be related to nodal status and used as a tool to adapt surgical staging.
Materials and methods
Experimental Design
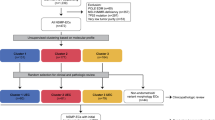
Approval for the present study was obtained from the local Medical Ethics Committee (CPP Ile-de-France V; e-4-15) and written informed consent was obtained from all women who provided the tissue samples used in this study. The experimental design for profiling the miRNA changes of nodal involvement in grade 1–2 early-stage FFPE primary EC tumor specimens is shown in Figure 1.
Flowchart describing the constitution of groups and the experimental design. EC, endometrial cancer; FFPE: formalin-fixed paraffin-embedded; FIGO, international federation of gynaecology and obstetrics; LVSI, lymphovascular space involvement; miRNA, microRNA; RT, reverse transcription; qPCR, quantitative real-time polymerase chain reaction.
Thirty-six women with early-stage EC (ie women with primary tumor confined to the corpus uteri with or without nodal involvement) who underwent primary surgical treatment (including total hysterectomy, bilateral salpingo-oophorectomy, and systematic nodal staging) between January 2003 and December 2012 in Tenon University Hospital—APHP were enrolled in the study. Nine of these women had lymph node (LN) metastases (positive LN) and met the following inclusion criteria: endometrioid adenocarcinoma, grade 1 or 2, FIGO stage I. The remaining 27 women, without LN metastases (negative LN), met the same inclusion criteria and were used as control subjects. One-to-three matching was performed according to the following three criteria: FIGO stage (IA, IB), histologic grade (grade 1, grade 2), and LVSI status (positive, negative). The exclusion criteria were as follows: previous malignancies, history of chemotherapy or radiotherapy, inflammatory disease, or Lynch syndrome (the search for a loss of expression of one of the Mismatch Repair proteins by immunohistochemistry and for tumor instability (microsatellite instability replication error repeats phenotype) were performed when EC occurred before the age 50 years or when there was a suggestive family history).
The clinical and pathologic variables of the women were extracted from maintained EC databases and included age, parity, body mass index (calculated as weight in kilograms divided by the square of height in meters), comorbidities (diabetes, dyslipidemia), 2009 FIGO stage, histologic type, and grade, depth of myometrial invasion, LVSI status, and time between surgery and sample analysis.
The median age was 72 years (interquartile range (IQR): 65–78) in the negative LN group and 68 years (IQR: 64–76) in the positive group (P=0.84). The two groups were comparable for the FIGO stage, histologic grade, and LVSI status. Furthermore, the comorbidities (diabetes and dyslipidemia), parity, body mass index, tumor size, European Society of Medical Oncology (ESMO) risk group of recurrence, and LVSI status did not differ according to the nodal status. The median time between surgery and sample RNA extraction was 32 months (IQR: 21–79) in the negative LN group and 30 months (IQR: 15–60) in the positive group (P=0.95) (Table 1).
Histologic Characteristics
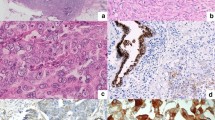
Histologic grade 1 is defined by 5% or less of a solid nonsquamous, nonmorular growth pattern; histologic grade 2 by 6–50% of solid nonsquamous, nonmorular pattern; and histologic grade 3 by more than 50% of solid nonsquamous, nonmorular growth pattern.17 The presence of grade 3 nuclei involving more than 50% of the tumor increases the final tumor grade by 1. A tumor is considered LVSI positive when tumor emboli are found within a space clearly lined by endothelial cells on hematoxylin and eosin-stained sections.18
RNA Extraction from FFPE Tissues
FFPE tissues were obtained from the hysterectomy specimens. FFPE tissue blocks were sectioned on a standard microtome (Leica-microsystems RM 2145) to generate successive 10 μm sections which were evaluated by a pathologist. Regions of invasive carcinoma were confirmed and marked on each slide. For each sample, marked regions from two slides were microdissected using a new, sterile blade and the dissected tissues were placed immediately into an RNase-free microcentrifuge tube.
Total RNA was extracted using the miRNeasy FFPE Kit (Qiagen, Courtaboeuf, France), according to the manufacturer’s instructions. Briefly, a deparaffinization solution was added to the FFPE samples. The tissues were then digested with protease and treated with DNase. After washing, the RNA, including the small miRNA fraction, was eluted with 20 μl distilled water. The concentrations and quality of the RNA recovered were measured using the Nanodrop 1000A spectrophotometer (Nanodrop Technologies, Wilmington, DE, USA). The median ratio of 260/280 was 1.87 (interquartile range (IQR): 1.84–1.90) and the median concentration was 415.8 ng/μl (IQR: 327.1–568.2).
Microarray Hybridization (GEO: GSE75968) and Data Analysis
To offset the high cost of an individual miRNome, the 27 specimens of negative LN status were pooled by 3 in accordance with the matching criteria. In this way the influence of individual differences among the women were limited while maintaining opportunities for exploration of possible statistical variations. Microarray analysis was conducted on 18 distinct specimens: 9 specimens with positive LN and 9 specimens with negative LN. Microarray hybridization on miRNA 4.0 chips (Affymetrix) was conducted at the genomic platform of the Institut Cochin, Paris. After validation of the RNA quality with Bioanalyzer 2100, 1 μg of total RNA was biotin labeled following the FlashTag Biotin HSR RNA labeling kit (Affymetrix).
After overnight hybridization, the miRNA 4.0 chips were washed in the Fluidic Station FS450 (Affymetrix) following a specific protocol and scanned using the GCS3000 7G. The scanned images were then analyzed with Expression Console software (Affymetrix) to obtain raw data (CEL files) and metrics for Quality Control. No apparent outliers were detected. Specific miRNAs analysis was performed using Partek Flow software, version 3.0 Copyright, 2014 (Partek Inc., St Louis, MO, USA). CEL files were imported and normalized using robust multi-array averaging.19
Genes with a nominal P-value ≤0.05 were considered to be differentially expressed. Among these, genes showing a variation of 2 were retained for further analysis.
Validated Target Gene and Enrichment Analysis
To estimate the biologic effects of the differentially expressed miRNAs, lists of validated target genes were determined using currently available databases, including Tarbase and Mirtarbase. Gene Ontology (GO) enrichment analysis was performed on the lists using Genomatix GePS (release 2.4.0; Genomatix BH, Munich, Germany).
Validation of Candidate miRNAs with Reverse Transcription (RT) and Quantitative Real-Time PCR (qRT–PCR)
One microgram of total RNA was used in all RT reactions that were performed with the miScriptII RT Kit, using the 5x miScript HiSpec Buffer method, according to the manufacturer’s instructions (Qiagen) with a Thermo Hybaid PXE 0.2 Thermal Cycler. cDNA samples were stored at −20 °C for further use.
miRNA expression was analyzed by real-time PCR using miScript SYBR Green PCR Kit (Qiagen) according to the manufacturer's instructions with an initial activation step at 95 °C for 15 min, followed by 45 cycles of 94 °C for 15 s, 55 °C for 30 s, and 70 °C for 30 s. A final melting curve analysis was performed to verify that a single product was amplified. All steps were performed in duplicate using a LightCycler 480 System (Roche). The results are expressed as Ct values and normalized on the calculated median Ct of each sample (ΔCt). miRNA primers were from Qiagen. Relative expression was calculated using the comparative Ct method (2−ΔΔCt). SNORD68 and RNU6 were both used as endogenous controls for data normalization.
For the results from the qRT–PCR on miRNA expression, data are expressed as means±s.e.m. Means between two groups were compared using the Mann–Whitney test. P<0.05 was considered to be statistically significant. GraphPad Prism version 5 was used for analysis of tissue samples (GraphPad Software, La Jolla, CA, USA).
Principal Component Analysis
Principal component analysis, a dimensionality reduction technique using a linear transformation applied on multidimensional data, was used to categorize miRNA.20
Principal component analysis and biplot plotting were performed using custom code in R 3.1.3 software, available online.
Optimal miRNA Fold-Change Cutoffs Correlated with Final Nodal Status
We calculate the ΔCt value by minus the Ct value of target microRNA by reference (endogenous control) microRNA. The normalized target microRNA expression level was calculated by 2−ΔCt. We further calculated the fold change of sample of interest to control sample by comparing their normalized target microRNA expression level.
For qualitative analysis, we calculated optimal cutoffs for each miRNA to correlate semiquantitative expression and final nodal status. The optimal fold-change (FC) cutoff was determined by a minimal P-value approach. This involved dichotomizing the FC level into dummy variables with a cutoff every 0.01 units of its range of values. Fisher tests comparing the number of women with or without LN metastases for every dummy variable were then calculated. The cutoff with the minimal P-value was chosen as the optimal cutoff for this variable.
Statistical Analysis
Unless otherwise specified, data were managed with an Excel database and analyzed using R 3.1.3 software, available online.
Results
Distinct miRNA Signatures of Type 1 EC Primary Tumors with Positive LN
To establish a miRNA profile for primary endometrioid adenocarcinoma EC with positive LN, we performed a microarray analysis comparing endometrioid adenocarcinoma EC tumors with positive LN to those with negative LN. We focused our study on the 2560 probes containing sequences for mature miRNAs. Among them, 142 probes had consistent normalized fluorescent intensities between the two groups (positive LN vs negative LN) (P<0.05) (Figure 2a). As illustrated by the volcano plot representation (Figure 2b), there was more than a twofold significant difference in the normalized fluorescence intensity of 12 of these miRNAs between the positive LN vs negative LN groups: seven miRNAs (miR-34c-5p, miR-375, miR-184, miR-34c-3p, miR-34b-5p, miR-148a-3p, and miR-504-5p) had a decreased expression and five miRNAs (miR-129-5p, miR-5001-5p, miR-6068, miR-4467, and miR-6850-5p) had an increased expression in samples from the positive LN group compared with samples from the negative group (Table 2).
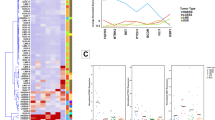
(a) Heat map of microRNA deregulation in positive LN vs negative LN FFPE grade 1–2 EC primary tumor specimen. One hundred and forty-two probes had consistent normalized fluorescent intensities between the two groups, negative LN vs positive LN (P<0.05). (b) Volcano plot. Abscissa is the logarithmic value of the level of variation (LogRatio) and ordinate is the negative logarithm of the statistical value (−log (p)) of fluorescence intensities of the hybridized probes from positive or negative LN samples. There was more than a twofold change in normalized fluorescence intensity of 12 microRNAs (blue triangle) between the positive LN vs negative LN groups (P<0.05): 7 have decreased intensity (on the left) and 5 have increased intensity (on the right). EC, endometrial cancer; FFPE, formalin-fixed paraffin-embedded; LN, lymph node.
GO term enrichment analysis was then performed using the list of all known validated targets for the miRNAs exhibiting at least a twofold change, with a significant value (P<0.05), in the EC primary tumors with positive LN. Genes related to cell transformation, genomic instability, cellular metabolic process, neoplasm invasiveness, neoplasm cell transformation, growth arrest, malignant neoplasms, cancer progression, and microsatellite instability were specifically enriched (Figure 3).
Enrichment analysis. Gene ontology enrichment analysis was performed (Genomatix GePS (release 2.4.0; Genomatix BH, Munich, Germany)) using the list of all known validated targets for the miRNAs exhibiting at least a twofold change, with a significant value (P<0.05), in the EC primary tumors with positive lymph node. EC, endometrial cancer.
Evaluation of miRNA Expression by Real-Time qRT–PCR Analysis
A qRT–PCR assay was used to confirm the expression of the miRNAs that were selected from the previous step. The expression levels of five miRNAs were significantly lower (miRNA-34c-5P, -375, -184, -34c-3p, and -34b-5p; P=0.006, 0.0008, 0.001, and 0.001, respectively) in the EC primary tumors with positive LN compared to those with negative LN. Two miRNAs (miR-148a-3p, miR-504-5p) and four miRNAs (miR-129-5p, miR-5001-5p, miR-6068, and miR-4467) were down- and upregulated, respectively, with no significant difference, in the EC primary tumors with positive LN compared with those with negative LN (Figure 4).
qRT–PCR assay. The expression levels of five microRNAs (miR-34c-5P, miR-375, miR-184, miR-34c-3p, and miR-34b-5p) were significantly lower in the grade 1–2 EC FFPE primary tumor samples with positive LN compared with those with negative LN. Mann–Whitney test, *P<0.05; **P<0.01; ***P<0.001; mean±s.e.m. EC, endometrial cancer; FFPE, formalin-fixed paraffin-embedded; LN, lymph node.
After eigenvalue decomposition of the 10 original dimensions (miR-34c-5p, miR-375, miR-184, miR-34c-3p, miR-34b-5p, miR-148a-3p, miR-504-5p, miR-5001-5p, miR-6068, and miR-4467), four principal components were found to be significant (with an eigenvalue >1). These components explained 75.72% of the variance in the data (Figure 5a). The two principal components explaining the greatest data variance are presented in Table 3. The first principal component, accounting for 32.02% of the variance, was dominated by tumor-suppressor miRNAs, especially of the miR-34 family, including miR-34b-5p, miR-34c-5p, and miR-34c-3p. The second component, accounting for 24.31% of the variance, was dominated by miR-129-5p.
(a) After eigenvalue decomposition of 10 original dimensions (miR-34c-5p, miR-375, miR-184, miR-34c-3p, miR-34b-5p, miR-148a-3p, miR-504-5p, miR-5001-5p, miR-6068, and miR-4467), four principal components were found to be significant (with an eigenvalue >1). These components explained 75.72% of the variance components. (b) Biplot of the first and second components. There is an opposition between microRNAs that have been defined by microarray as being upregulated (miR-129-5p, miR-6068, miR-5001-5p, and miR-4467) and downregulated (miR-34c-3p, miR-34c-5p, miR-34b-5p, miR-375, miR-504-5p, miR-184, and miR-148a-3p) in the positive LN group. PC, principal component; LN, lymph node.
A biplot of the first and second components (those explaining the greatest variance) is shown in Figure 5b, demonstrating an opposition between miRNAs that were defined by microarray as being upregulated miRNAs (miR-129-5p, miR-6068, miR-5001-5p, and miR-4467) and downregulated miRNAs (miR-34c-3p, miR-34c-5p, miR-34b-5p, miR-375, miR-504-5p, miR-184, and miR-148a-3p) in the positive LN group.
Correlation between miRNA Expression and Nodal Status in EC
Optimal cutoffs denoting the strongest correlation between quantitative expression of the miRNAs that were selected from the previous step and final LN status are summarized in Figure 6. The FC cutoffs defined were 0.30, 0.30, 0.0852, 0.09, 0.50, 1.45, and 1.75 for miR-375, miR-184, miR-34c-5p, miR-34b-5p, miR-148a-4p, miR-129-5p, and miR-4467, respectively. miR-375 and miR-184 had the most significant P-values: 0.001 and 0.006, respectively. We compared nodal status according to the cutoffs previously determined: women with EC and an miRNA-375 FC<0.30 were more likely to have positive LN (n=8; 53.3%) compared with those with an miRNA-375 FC>0.30 (n=1; 4.8%), P=0.001; women with EC and an miRNA -184 FC<0.30 were more likely to have positive LN (n=6; 60.0%) compared with those with an miRNA -184 FC>0.30 (n=3; 11.5%), P=0.006 (Table 4).
Optimal fold-change cutoffs denoting a correlation between microRNA expression and lymph node status in histologic grade 1–2 EC primary tumor specimens. (a) miR-34c-5p, (b) miR-375, (c) miR-184, (d) miR-34c-3p, (e) miR-34b-5p, (f) miR-148a-3p, (g) miR-504-5p, (h) miR-129-5p, (i) miR-5001-5p, (j) miR-6068, (k) miR-4467. EC, endometrial cancer.
Discussion
Our results show that in grade 1–2 early-stage EC, women with LN metastases have different miRNA profiles compared with those without metastatic LN. Moreover, we found that final LN status can be accurately predicted using the miRNA expression level.
Nodal status in grade 1–2 early-stage EC is currently a major subject of debate. Although lymphadenectomy is no longer recommended in early-stage EC since the publication of a meta-analysis demonstrating no impact on survival, there are several discrepancies about how to manage women with early-stage/grade 1–2 EC in terms of surgical staging.21 The major limitation of preoperative imaging techniques,3 such as MRI and CT scan, is the poor detection rate of LN metastases.5 Moreover, a prospective multicentre study showed that 12% of women with low- to intermediate-risk EC had LN metastases that would have been overlooked if LN staging had not been performed.6 This partially explains the great heterogeneity of recurrence rates in women with presumed early-stage tumors. Hence, the main challenge for physicians managing women with early-stage grade 1–2 EC is when to opt for lymphadenectomy and, in the absence of data on lymph node status, when to opt for adjuvant therapy.
To address this issue, we assessed miRNA level expression in primary tumors according to LN status. According to our chip analysis, seven miRNAs were downregulated and five upregulated more than twofold between the positive LN vs negative LN groups. Also, according to the enrichment analysis, the validated target genes for these differentially expressed miRNAs were exclusively involved in carcinogenic pathways, underlining the strength of our analysis. These results support data from the current literature showing that miRNAs can act as tumor suppressors or oncogenes in various cancers13 including EC.22,23 Furthermore, recent evidence indicates that several miRNAs, including miR-34, regulate the metastatic process through the expression of epithelial to mesenchymal transition—transcriptions factors or epithelial to mesenchymal transition-activating signaling pathways (SNAIL1/SNAIL2, basic helix–loop–helix, E47, E2-2, TWIST1/TWIST2, and ZEB (ZEB1/ZEB2) families)24 that act as E-cadherin repressors25 and, ultimately, enhance cell migration and invasiveness in various cancers including EC.26, 27, 28
After validation by RT–qPCR of miRNA selected from the previous step, we found that miR-34b-5p, miR-34c-5p, miR-34c-3p, miR-375, and miR-184 emerged as being particularly relevant to determine positive or negative LN metastatic status in women with grade 1–2 early-stage EC. Furthermore, miR-34b-5p, miR-34c-5p, miR-34c-3p, and miR-184 dominated the first principal component of the principal component analysis. This is in accordance with current literature: the miR-34 family (miR-34a, b, and c) is attracting a lot of attention since it has been found to play a key role as a tumor suppressor in several cancers.29 Indeed, it is a direct target of the tumor-suppressor gene p53 inducing apoptosis, cell cycle arrest, and senescence when upregulated. It also negatively influences the viability of cancer stem cells and inhibits metastasis formation. The miR-34 family acts on apoptosis and cell cycle through the repression of many proteins involved in the regulation of these two biologic processes. In particular, the miR-34 family members bind to the 3′-UTRs of genes such as CDK4 and CDK6 (cell cycle proteins), Bcl-2 (an apoptosis regulator), SNAIL (epithelial–mesenchymal transition) and CD44 (migration and metastasis), thus repressing their expression.29 Numerous studies have shown the dysregulation of miR-34 in various types of cancers, including hepatocellular, mesothelial and colon cancer, melanomas, leukemia, nasopharyngeal cancer,30 prostate cancer,31 neuroblastomas,32 glioblastoma33 and breast cancer.34 Yet, little research has been conducted in EC, apart from a functional study by Li et al35 demonstrating that miR-34c acts as a tumor suppressor in HEC-1-B cells, and that E2F3 protein may be a target of miR-34c.
In the same way, miR-184 functions as a tumor suppressor in various cancers (renal carcinoma,36 glioma,37, 38 non-small-cell lung cancer,39 oral squamous cell carcinoma,40 breast cancer41) by repressing oncogenes such as TNFAIP2,37 SND1,38 CDC25A, c-MYC,39 and regulating the AKT/mTORC1 pathway.41 As suggested by various studies, miR-184 might be a useful diagnostic and therapeutic tool for malignant diseases.38, 39 However, to the best of our knowledge, ours is the first study to provide data about miR-184 expression in EC.
Finally, a recent review by Yan et al emphasized the emerging role of miR-375 in cancer and specified that miR-375 is frequently downregulated in multiple types of cancer, especially in hepatocarcinoma,42 esophagus cancer,43 osteosarcoma,44 lung cancer,45 and acts as a tumor suppressor by repressing many critical oncogenes such as PDK1, JAK2, IGF1R,46 AEG-1,42 and suppressing the PI3K/Akt pathway.44 Furthermore, miR-375 might be used as a diagnostic and prognostic biomarker in various cancers. In this setting, reduced miR-375 expression could be a predictor of poor outcome (HR: 12.8, 95% CI: 3–49) and distant metastasis (HR: 8.7, 95% CI: 2–31) in head and neck squamous cell carcinomas.47 Also, women with metastatic non-small-cell lung cancer had lower plasma miR-375 expression than those with non-metastatic non-small-cell lung cancer (P<0.05).45 However, miR-375 has only rarely been assessed in EC. One study found miR-375 to be associated with histologic type EC (downregulated in papillary-serous carcinoma compared with carcinosarcoma) (FC=−5.1, P-value=3.8e−04).48
We focused our analysis on postoperative FFPE EC primary tumors since it is known that FFPE specimens can be used for real-time PCR-based quantitative miRNA expression studies.16 This meant we could draw on the entire specimen collection of the Pathology Department containing samples taken between 2003 and 2012, and select samples to form two groups fully respecting the matching criteria. However, to provide a diagnostic tool, this analysis should be validated on preoperative samples. Unfortunately, assessing miRNA expression in preoperative biopsy samples might be difficult due to the contamination of normal endometrial or myometrial tissue. A heterogenic sample, with distinct miRNA profiles,13 could lead to a wrong analysis. Interestingly, two recent studies have demonstrated that circulating miRNAs have a high accuracy in diagnosing endometrioid EC ((miR-15b, -27a, and -233)49 and (miR-9/miR-1228 and miR-9/miR-92a)50). The miRNA described in our study may be preoperatively assessed in blood samples of women with grade 1–2 early-stage EC and might serve as a novel, non-invasive biomarker in the future, avoiding discrepancies in histologic type and grade between preoperative biopsies and final histology.
Some limitations of the study should be underlined. Because of the use of FFPE specimens with insufficient quality and RNA integrity, we could not assess the expression of exact targets for the miRNAs validated in this study. Future research is needed to resolve this problem.
Conclusion
In conclusion, this is the first study investigating the relative expression of mature miRNA genes in early-stage grade 1–grade 2 EC primary tumors according to the nodal status. This miRNA expression profile may provide a basis for further study of the miRNA function in endometrioid adenocarcinoma, and be used as a diagnostic tool for nodal status.
References
Ferlay J, Soerjomataram I, Dikshit R et al. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer 2015;136:E359–E386.
Siegel RL, Miller KD, Jemal A . Cancer statistics, 2015. CA Cancer J Clin 2015;65:5–29.
Morice P, Leary A, Creutzberg C et al. Endometrial cancer. Lancet Lond Engl 2015.
Colombo N, Preti E, Landoni F et al. Endometrial cancer: ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann Oncol 2013;24 (Suppl 6):vi33–vi38.
Chi DS, Barakat RR, Palayekar MJ et al. The incidence of pelvic lymph node metastasis by FIGO staging for patients with adequately surgically staged endometrial adenocarcinoma of endometrioid histology. Int J Gynecol Cancer 2008;18:269–273.
Ballester M, Dubernard G, Lécuru F et al. Detection rate and diagnostic accuracy of sentinel-node biopsy in early stage endometrial cancer: a prospective multicentre study (SENTI-ENDO). Lancet Oncol 2011;12:469–476.
Guntupalli SR, Zighelboim I, Kizer NT et al. Lymphovascular space invasion is an independent risk factor for nodal disease and poor outcomes in endometrioid endometrial cancer. Gynecol Oncol 2012;124:31–35.
Bendifallah S, Canlorbe G, Arsène E et al. French multicenter study evaluating the risk of lymph node metastases in early-stage endometrial cancer: contribution of a risk scoring system. Ann Surg Oncol 2015;22:2722–2728.
Bendifallah S, Canlorbe G, Raimond E et al. A clue towards improving the European Society of Medical Oncology risk group classification in apparent early stage endometrial cancer? Impact of lymphovascular space invasion. Br J Cancer 2014;110:2640–2646.
Nugent EK, Bishop EA, Mathews CA et al. Do uterine risk factors or lymph node metastasis more significantly affect recurrence in patients with endometrioid adenocarcinoma? Gynecol Oncol 2012;125:94–98.
Guo H, Ingolia NT, Weissman JS et al. Mammalian microRNAs predominantly act to decrease target mRNA levels. Nature 2010;466:835–840.
Lewis BP, Burge CB, Bartel DP . Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 2005;120:15–20.
Farazi TA, Hoell JI, Morozov P et al. MicroRNAs in human cancer. Adv Exp Med Biol 2013;774:1–20.
Plummer PN, Freeman R, Taft RJ et al. MicroRNAs regulate tumor angiogenesis modulated by endothelial progenitor cells. Cancer Res 2013;73:341–352.
Tsukamoto O, Miura K, Mishima H et al. Identification of endometrioid endometrial carcinoma-associated microRNAs in tissue and plasma. Gynecol Oncol 2014;132:715–721.
Lee H, Choi HJ, Kang CS et al. Expression of miRNAs and PTEN in endometrial specimens ranging from histologically normal to hyperplasia and endometrial adenocarcinoma. Mod Pathol 2012;25:1508–1515.
Querleu D, Planchamp F, Narducci F et al. Clinical practice guidelines for the management of patients with endometrial cancer in France: recommendations of the Institut National du Cancer and the Société Française d’Oncologie Gynécologique. Int J Gynecol 2011;21:945–950.
Briët JM, Hollema H, Reesink N et al. Lymphvascular space involvement: an independent prognostic factor in endometrial cancer. Gynecol Oncol 2005;96:799–804.
Irizarry RA, Hobbs B, Collin F et al. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostat Oxf Engl 2003;4:249–264.
Ibrahim GM, Morgan BR, Macdonald RL . Patient phenotypes associated with outcomes after aneurysmal subarachnoid hemorrhage: a principal component analysis. Stroke J Cereb Circ 2014;45:670–676.
Frost JA, Webster KE, Bryant A et al. Lymphadenectomy for the management of endometrial cancer. Cochrane Database Syst Rev 2015;9:CD007585.
Sianou A, Galyfos G, Moragianni D et al. The role of microRNAs in the pathogenesis of endometrial cancer: a systematic review. Arch Gynecol Obstet 2015;292:271–282.
Kontomanolis EN, Koukourakis MI . microRNA: the potential regulator of endometrial carcinogenesis. MicroRNA 2015;4:18–25.
Díaz-López A, Moreno-Bueno G, Cano A . Role of microRNA in epithelial to mesenchymal transition and metastasis and clinical perspectives. Cancer Manag Res 2014;6:205–216.
Peinado H, Olmeda D, Cano A . Snail, Zeb and bHLH factors in tumour progression: an alliance against the epithelial phenotype? Nat Rev Cancer 2007;7:415–428.
Montserrat N, Mozos A, Llobet D et al. Epithelial to mesenchymal transition in early stage endometrioid endometrial carcinoma. Hum Pathol 2012;43:632–643.
Liao Y, He X, Qiu H et al. Suppression of the epithelial-mesenchymal transition by SHARP1 is linked to the NOTCH1 signaling pathway in metastasis of endometrial cancer. BMC Cancer 2014;14:487.
Nieto MA . Epithelial plasticity: a common theme in embryonic and cancer cells. Science 2013;342:1234850.
Agostini M, Knight RA . miR-34: from bench to bedside. Oncotarget 2014;5:872–881.
Li Y-Q, Ren X-Y, He Q-M et al. MiR-34c suppresses tumor growth and metastasis in nasopharyngeal carcinoma by targeting MET. Cell Death Dis 2015;6:e1618.
Hagman Z, Haflidadottir BS, Ansari M et al. The tumour suppressor miR-34c targets MET in prostate cancer cells. Br J Cancer 2013;109:1271–1278.
Welch C, Chen Y, Stallings RL . microRNA-34a functions as a potential tumor suppressor by inducing apoptosis in neuroblastoma cells. Oncogene 2007;26:5017–5022.
Gao H, Zhao H, Xiang W . Expression level of human miR-34a correlates with glioma grade and prognosis. J Neurooncol 2013;113:221–228.
Yu F, Jiao Y, Zhu Y et al. MicroRNA 34c gene down-regulation via DNA methylation promotes self-renewal and epithelial-mesenchymal transition in breast tumor-initiating cells. J Biol Chem 2012;287:465–473.
Li F, Chen H, Huang Y et al. miR-34c plays a role of tumor suppressor in HEC-1-B cells by targeting E2F3 protein. Oncol Rep 2015;33:3069–3074.
Su Z, Chen D, Li Y et al. microRNA-184 functions as tumor suppressor in renal cell carcinoma. Exp Ther Med 2015;9:961–966.
Cheng Z, Wang HZ, Li X et al. microRNA-184 inhibits cell proliferation and invasion, and specifically targets TNFAIP2 in Glioma. J Exp Clin Cancer Res 2015;34:27.
Emdad L, Janjic A, Alzubi MA et al. Suppression of miR-184 in malignant gliomas upregulates SND1 and promotes tumor aggressiveness. Neuro-Oncology 2015;17:419–429.
Lin T-C, Lin P-L, Cheng Y-W et al. microRNA-184 deregulated by the microRNA-21 promotes tumor malignancy and poor outcomes in non-small cell lung cancer via targeting CDC25A and c-Myc. Ann Surg Oncol 2015.
Manikandan M, Deva Magendhra Rao AK, Rajkumar KS et al. Altered levels of miR-21, miR-125b-2*, miR-138, miR-155, miR-184, and miR-205 in oral squamous cell carcinoma and association with clinicopathological characteristics. J Oral Pathol Med 2014;44:792–800.
Phua YW, Nguyen A, Roden DL et al. microRNA profiling of the pubertal mouse mammary gland identifies miR-184 as a candidate breast tumour suppressor gene. Breast Cancer Res 2015;17:83.
He X-X, Chang Y, Meng F-Y et al. microRNA-375 targets AEG-1 in hepatocellular carcinoma and suppresses liver cancer cell growth in vitro and in vivo. Oncogene 2012;31:3357–3369.
Mayne GC, Hussey DJ, Watson DI . microRNAs and esophageal cancer—implications for pathogenesis and therapy. Curr Pharm Des 2013;19:1211–1226.
Shi Z-C, Chu X-R, Wu Y-G et al. microRNA-375 functions as a tumor suppressor in osteosarcoma by targeting PIK3CA. Tumour Biol 2015;36:8579–8584.
Yu H, Jiang L, Sun C et al. Decreased circulating miR-375: a potential biomarker for patients with non-small-cell lung cancer. Gene 2014;534:60–65.
Yan J-W, Lin J-S, He X-X . The emerging role of miR-375 in cancer. Int J Cancer 2014;135:1011–1018.
Harris T, Jimenez L, Kawachi N et al. Low-level expression of miR-375 correlates with poor outcome and metastasis while altering the invasive properties of head and neck squamous cell carcinomas. Am J Pathol 2012;180:917–928.
Ratner ES, Tuck D, Richter C et al. MicroRNA signatures differentiate uterine cancer tumor subtypes. Gynecol Oncol 2010;118:251–257.
Wang L, Chen Y-J, Xu K et al. Circulating microRNAs as a fingerprint for endometrial endometrioid adenocarcinoma. PloS One 2014;9:e110767.
Torres A, Torres K, Pesci A et al. Diagnostic and prognostic significance of miRNA signatures in tissues and plasma of endometrioid endometrial carcinoma patients. Int J Cancer 2013;132:1633–1645.
Acknowledgements
We thank Sebastien Jacques and Florent Dumont (Plateforme Genomic, INSERM U1016, Institut Cochin), Annie Cortez (Department of Pathology, Tenon University Hospital) and Michèle Oster (UMRS 938) for expert contributions.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
GC received support from Association pour la Recherche sur le Cancer (ARC). The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Canlorbe, G., Wang, Z., Laas, E. et al. Identification of microRNA expression profile related to lymph node status in women with early-stage grade 1–2 endometrial cancer. Mod Pathol 29, 391–401 (2016). https://doi.org/10.1038/modpathol.2016.30
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2016.30
This article is cited by
-
Computing microRNA-gene interaction networks in pan-cancer using miRDriver
Scientific Reports (2022)
-
Potential of blood-based biomarker approaches in endometrium and breast cancer: a case-control comparison study
Archives of Gynecology and Obstetrics (2022)
-
MiR-369-3p participates in endometrioid adenocarcinoma via the regulation of autophagy
Cancer Cell International (2019)
-
Identification of micro-RNA expression profile related to recurrence in women with ESMO low-risk endometrial cancer
Journal of Translational Medicine (2018)
-
Impact of hospital and surgeon case volume on morbidity in colorectal endometriosis management: a plea to define criteria for expert centers
Surgical Endoscopy (2018)