Abstract
In clinical routine pathology today, detection of protein in intact formalin-fixed, paraffin-embedded tissue is limited to immunohistochemistry, which is semi-quantitative. This study presents a new and reliable quantitative immunohistochemistry method, qIHC, based on a novel amplification system that enables quantification of protein directly in formalin-fixed, paraffin-embedded tissue by counting of dots. The qIHC technology can be combined with standard immunohistochemistry, and assessed using standard bright-field microscopy or image analysis. The objective was to study analytical performance of the qIHC method. qIHC was tested under requirements for an analytical quantitative test, and compared with ELISA and flow cytometry for quantitative protein measurements. Human epidermal growth factor receptor 2 (HER2) protein expression was measured in five different cell lines with HER2 expression from undetectable with immunohistochemistry to strong positive staining (IHC 3+). Repeatability, reproducibility, robustness, linearity, dynamic range, sensitivity, and quantification limits were evaluated. Reproducibility and robustness were assessed in a setup to resemble daily work in a laboratory using a commercial immunohistochemistry platform. In addition, qIHC was correlated to standard HER2 immunohistochemistry in 44 breast cancer specimens. For all evaluated parameters, qIHC performance was either comparable or better than the reference methods. Furthermore, qIHC has a lower limit of detection than both immunohistochemistry and the ELISA reference method, and demonstrated ability to measure HER2 accurately and precise within a large dynamic range. In conclusion, the results show that qIHC provides a sensitive, quantitative, accurate, and robust assay for measurement of protein expression in formalin-fixed, paraffin-embedded cell lines, and tissue.
Similar content being viewed by others
Main
In clinical routine pathology today, detection of protein in intact formalin-fixed, paraffin-embedded tissue is limited to immunohistochemistry, which is semi-quantitative. Accurate and robust measurement of protein expression from formalin-fixed, paraffin-embedded samples, without loss of morphological information, is a challenge. Nevertheless, in routine assessments the expression levels of protein biomarkers are reported and used for treatment decisions.
The human epidermal growth factor receptor 2 (HER2) is overexpressed in 10–25% of breast carcinomas. The Food and Drug Administration approved the first immunohistochemistry-based companion diagnostic HER2 test (HercepTest) in 1998 to assess protein levels in a semi-quantitative setting followed by approval of in situ hybridization testing for HER2 gene amplification. HER2 testing on either gene level or protein level determines patient eligibility for treatment with trastuzumab in breast and in gastric cancer.1, 2, 3, 4, 5, 6, 7, 8, 9, 10
In immunohistochemistry, and to some degree as for in situ hybridization, different diagnoses for the same patient have been reported when patient tissues were tested at different laboratories11, 12, 13, 14, 15 (inter-laboratory disconcordance). One issue is local differences in procedures for obtaining and preserving tissue before analysis. Within the last decade, standardizing the process from the operating room to the pathologist’s assessment has been an area of focus. The ASCO/CAP guidelines specify the pre-analytic, analytic, and post-analytic factors necessary for achieving optimal HER2 assay results by either immunohistochemistry, fluorescence, or chromogenic in situ hybridization.10 Similarly, following guidelines from Clinical and Laboratory Standards Institute and manufacturers’ protocols all contributes to improve the quality of results. Panels evaluating biomarkers have published scoring and handling recommendations1, 10, 16, 17, 18 all aiming toward standardization of diagnostic evaluation. Nevertheless, both interobserver19, 20, 21, 22, 23, 24, 25 and intraobserver variation25, 26, 27, 28, 29 have been demonstrated to have an undesired role in patient diagnosis. Immunohistochemistry relies heavily on the skill set of the trained pathologist.
A number of methods to obtain more quantitative results from formalin-fixed, paraffin-embedded tissue have seen light within the last decade, either directed at protein or gene level. Some examples are the AQUA Technology,30, 31, 32, 33, 34 HERmark,35, 36, 37, 38 Mammaprint,39, 40, 41, 42 Oncotype Dx,43, 44, 45, 46, 47 and the PAM50 Breast Cancer Intrinsic Classifier.48, 49 A review50 compared PAM50, Oncotype Dx, and the immunohistochemistry-based immunohistochemistry 4,51, 52, 53 and concluded that PAM50 demonstrated the most prognostic results in the TransATAC study.50 Unfortunately, most of these new assays providing quantitative data need to be performed at a centralized laboratory.
We have developed a new reliable quantitative immunohistochemistry method, qIHC, based on a novel amplification system that enables quantification of protein directly in formalin-fixed, paraffin-embedded tissue by counting of dots. The qIHC method addresses the need for a robust and quantitative assay, which can be combined with the morphological information gained from immunohistochemistry. The assay is based on the iCARD chemistry, which has been described in Lohse et al.54 It converts antibody/antigen complexes into red dots, which can subsequently be counted and quantified.
qIHC combines the ease, speed, and morphological information gained from classical immunohistochemistry assessment with a quantitative assessment of protein expression in situ. Using HER2 as a model system, this study assesses the analytical performance of the qIHC assay in both formalin-fixed, paraffin-embedded cell pellets, and patient breast cancer specimens.
The qIHC method (Figure 1) is in principle similar to classic immunohistochemistry, and like classic immunohistochemistry, the basis for the amplification is enzyme deposition (typically Horse radish peroxidase (HRP)). Step (1) The primary antibody binds the target (protein/receptor) of interest. Step (2) A secondary, HRP-labeled antibody recognizes the primary antibody and binds. These first two steps in the qIHC reaction are directly comparable to standard immunohistochemistry. However, in qIHC, only a pre-determined fraction of the secondary antibodies is labeled. The secondary-labeled antibody is mixed with non-labeled antibody to increase robustness of the assay. Finally, an amplification reaction generates a dot centered around the labeled antibody, ie, directly at the single target (Step 3–5). The number of dots can be counted and as the ratio between labeled and unlabeled secondary antibody is known, the qIHC assay allows for a direct correlation between number of dots and amount of biomarker present in the tissue.
1: Primary antibody binds antigen after antigen retrieval. 2: Primary antibody is recognized by a HRP-labeled secondary antibody. A pre-determined fraction of the secondary antibodies is labeled. 3: Enzyme substrate is added and deposited. 4: AP-labeled antibody binds substrate and cross linker, resulting in. 5: Red dot reaction.
In this study, analytical performance of the qIHC assay including parameters such as precision, repeatability, linearity, dynamic range, and sensitivity have been assessed using HER2-expressing cancer cell lines. Furthermore, compatibility with clinical specimens was evaluated as the assay was applied to anonymized breast cancer patient samples.
Materials and methods
Cells
The following cancer cell lines were obtained from the American Type Culture Collection (ATCC, USA): MDA-MB-468, MDA-MB-231, MDA-MB-175-VII, MDA-MB-453, and SK-BR-3. All cell lines were maintained in RPMI medium supplemented with 10% fetal calf serum, 1% penicillin–streptomycin, 1% sodium pyruvate, and 10 mM Glutamax-I. All media and supplements were purchased from Invitrogen, USA. All cells were kept in a 37 °C humidified incubator in 95% air, 5% CO2 according to ATCC recommendations. Cells were harvested in the exponential phase and divided into three fractions for analysis. The cell fractions were respectively (1) formalin-fixed, paraffin-embedded for immunohistochemistry and qIHC analysis, (2) lysed for ELISA or western blotting or (3) re-suspended into a 0.01 M PBS buffer pH 7.4 for flow cytometry analysis. For immunohistochemistry, the cells were embedded into formalin-fixed, paraffin-embedded multi blocks including all five cell lines in one formalin-fixed, paraffin-embedded block. Consecutive serial sections were processed as described below from the paraffin-embedded cell pellets for immunohistochemistry analysis and qIHC and stored cold until staining was performed.
Cell Lysis
For cell lysis, cells were harvested by trypsination and counted. The cell pellets were lysed according to procedure described by the manufacturer of the Cell Lysis Buffer (Cell Signaling Technology, USA). Cell lysates were stored at −18 °C until further analysis.
Tissue
The study included 44 formalin-fixed, paraffin-embedded invasive breast cancer tissue specimens with known fixation history (10% neutral buffered formalin, 18–24 h).
All specimens were bought from commercial tissue vendors. All specimens were completely anonymized prior to receipt. Specimens were not individually identifiable and it was impossible to trace the identity of the patients.
According to the Danish law on the Research Ethics Committee System and handling of biomedical research projects and communication between Dako and the Danish Committee on Biomedical Research Ethics and the Regional Ethics Committee, the tests performed at Dako on anonymous residual tissue are not subject to an approval by the Regional Ethics Committee system because such studies are considered quality control projects. Therefore, no Regional Ethics Committee approval for this work has been obtained.
Blocks were selected based on their HER2 immunohistochemistry scores (HercepTest standard categories of expression), in accordance with the HercepTest scoring system. The slides were chosen to have a balanced distribution of 0, 1+, 2+, and 3+ HER2 categories.
Formalin-Fixed, Paraffin-Embedded Processing, and Antigen Retrieval
Formalin-fixed, paraffin-embedded blocks were sectioned at a thickness of 4 μm, dried for 1 h at 65 °C, deparaffinized, rehydrated, and subjected to target retrieval in the pre-treatment module, PT Link (Dako, Denmark) at 95 °C for 40 min in Epitope Retrieval Solution (Dako) according to manufacturer’s instruction for HercepTest (Dako).
Immunohistochemistry
Immunohistochemical analysis was performed using the HercepTest kit (Dako). The sections were stained within 5 days of cutting using an Autostainer Link 48 (Dako) in strict accordance with the manufacturer’s instructions. Control cell lines for run control, supplied with the kit, were included together with a negative control sample in each run.
qIHC
The qIHC assay was stained using an Autostainer Link 48, and quantitative assessment was done by counting dots/cell using image analysis. HER2 antibody and Epitope Retrieval Solution were from HercepTest kit and used according to manufacturers’ instruction. qIHC HER2 assays were performed as previously described.54
An example of a qIHC stain and the corresponding immunohistochemistry stain is presented in Figure 2, wherein cut sections from the same tumor on two neighboring cut sections were stained with either HER2 immunohistochemistry or HER2 qIHC, respectively.
Whole-Slide Scanning and Image Analysis
Whole-slide digital image acquisition was performed using the Aperio ScanScope CS System (Aperio Technologies, USA), × 20 objective. HER2 immunohistochemistry was quantified in tissues using Aperio Membrane v9 algorithm, version 9.2. Image analysis was applied using a dot counting algorithm (Indica Labs in situ hybridization quantification tool v. 1.2, Indicalab, Corrales, NM) developed for bright-field in situ hybridization, and intended for research use only. The image algorithm identified cells based on their nuclear staining with hematoxylin. Only qIHC dots within the cell were evaluated. Other dots were disregarded as background noise, and not evaluated. A total number of cells with individual information of dots per cell was evaluated for each case. Each case was reported as a unified median of average number of dots/cell (including s.d).
Histoscore is a measure to convert classic immunohistochemistry into a more quantitative range. Histoscore or H-score is based on staining intensity and percentages of stained cell. Histoscore is based on four immunohistochemistry categories reported in percent cells: negative (0), weak (1+), moderate (2+), and strongly (3+) stained membranes. In each case, a histoscore with a potential range of 0–300 was calculated as follows: HistoScore=((1 × % weakly stained cells)+(2 × % moderately stained cells)+(3 × % strongly stained cells)).
Statistical Analysis
All statistical analyses were performed in a univariate setup, without any studies being reanalyzed, thus there was no need for P-value correction. A P-value<0.05 was considered statistically significant. For analytical performance testing, sample size calculations were performed using alpha at 0.05 and power of 0.95. SPSS 21 (IBM Corp. Released 2012. IBM SPSS Statistics for Windows, Version 21.0. Armonk, NY: IBM) or GraphPad Prism version 5.04 for Windows, (GraphPad Software, La Jolla, CA, USA) was used for statistical analysis.
Flow Cytometry
Flow cytometry analyses were performed using a monoclonal mouse anti-HER2 antibody developed internally at Dako. The specificity was characterized by epitope mapping and examined for cross reactivity to HER1, HER3 and HER4 by western blot.
Cells were harvested by trypsination and counted. Samples of 5 × 105 cells were permeabilized to enable intracellular staining of the HER2 membrane protein according to instructions for use (Dako, Denmark). Subsequently, the cells were labeled according to the procedure described in QIFIKIT (Dako) for quantitative analysis of indirect immunofluorescence staining by flow cytometry. Mouse IgG1 (Dako) was used as negative control antibody. Flow cytometry analysis was performed on a FacsCalibur cytometer (BD Biosciences, San Jose, CA). FITC fluorescence intensity of labeled cells was determined on a FL1 detector (530/30 band filter). FluoroSpheres (Dako) was used for calibration of the instrument setting prior to analysis.
ELISA HER2 protein levels were measured by sandwich ELISA in accordance with the recommended procedure by the manufacturer of c-erbB2/c-neu Rapid Format ELISA kit (Merck Millipore, Germany).
The total protein content in the individual cell lysate was determined using the BCA method (BCA Protein Assay Reagent, Pierce, Thermo Scientific, Rockford, IL, USA) according to the BSA standard. The total protein content was used to normalize the HER2 values for each sample.
Cell lysates were tested and measured in duplicate serial dilutions.
Results
The ASCO/CAP guideline for HER2 testing10, 55 covers standardized setup for analytic processing procedures for HER2 testing using immunohistochemistry or fluorescence in situ hybridization. All Food and Drug Administration-approved HER2 immunohistochemistry diagnostic assays include control cell lines based on immortalized cancer cell lines with different HER2 expression for internal run controls. As a result the accuracy of the tests can only be estimated and not fully determined, although the precision of the assay can be measured.56 There are currently no HER2 immunohistochemistry assays for direct HER2 quantification.
In our study, the recommended principles and practices57, 58, 59 for validating a quantitative test were followed including testing for accuracy, reproducibility, repeatability, robustness, and precision using cancer cell lines. The HER2 expression in the cell lines ranged from undetectable with current HER2 immunohistochemistry (~1000 receptors/cell) to strongly positive (immunohistochemistry 3+, >5.5 × 106 receptors/cell). Using cell lines for analytical validation instead of patient samples provides the advantage that the HER2 expression in cell lines can be quantified using other linear and quantitative methods. Hence, we were able to correlate qIHC analysis on tissue sections and sections of paraffin-embedded cell pellets with ELISA measurements of HER2 protein in cell lysates and flow cytometric quantification of HER2 in living cells. Comparison of the results from the different tests and methods is used for demonstrating the quantitative nature of the qIHC assay.
Accuracy
Closeness of agreement between the test result and an accepted reference value was tested by assessing multiple independent cell lines. HER2 protein levels for all cell lines have previously been characterized and reported with established quantitative methods, such as western blot,60, 61 ELISA,31, 36, 60 or flow cytometry.36, 37, 60, 61 Variance in HER2 protein levels reported in the literature suggests local laboratory variation, most likely due to difference in cell handling, HER2 antibody, or methodology setup. In order to reduce the cell handling parameter, measurements of HER2 protein levels were performed on five different cell lines but the same cell culture using HER2 qIHC in parallel with ELISA and flow cytometry (Table 1).
The cell lines were selected on basis of the span in HER2 protein level, and cover a dynamic range of 103 receptors/cell. The resulting data demonstrated overall concordance, and all methods demonstrated ability to measure and rank the five cell lines according to expected HER2 profile.
Precision
To evaluate the variability of repeated measurements’ precision, studies were based on International Standard ISO 5725 as well as independent Clinical and Laboratory Standards Institute guidelines EP05-A3.62 The same batch of reagents was used for all tests. In addition, the ability to separate the five different cell lines by qIHC was evaluated. For precision testing, three staining runs, including the same formalin-fixed, paraffin-embedded five cell line multi block, were tested in nine replicates, resulting in a total of 135 data points. Data for the individual cell lines were pooled between the individual test runs, thus adding run-to-run variation on top of the expected within-run variation. Combined data of five different cell lines were tested 45 times (five runs, nine replicates in each), and separation between the individual cell lines were tested by multiple comparisons using Kruskal–Wallis test with Dunn's multiple comparison which compare inter and intraspecific variability. qIHC measurements of all five cell lines were statistical significantly different (P<0.0001–0.0033, with an overall P<0.0001), and are well separated (Figure 3).
Sensitivity
Sensitivity is often measured by diluting the analyte until it cannot be measured. This is not possible for immunohistochemistry assays because no reference analyte for tissue exists. Instead, the analytical sensitivity of HER2 qIHC was tested by the ability to obtain measureable results with HER2 qIHC on cell lines with low HER2 expression. The cell lines, MDA-MB-468 and MDA-MB-231, cannot be measured by immunohistochemistry and are both immunohistochemistry negative. Furthermore, although MDA-MB-231 is quantifiable with ELISA, MDA-MB-468 is not as demonstrated in this study and in an earlier study.60 To determine the background level, control experiments without HER2 antibody were run. Nine formalin-fixed, paraffin-embedded cell multi blocks were stained with and without primary HER2 antibody and assessed using image analysis. The qIHC value of MDA-MB-468 is well separated from the MDA-MB-231 value (unpaired t-test P<0.0001) and both are above observed background level (Figure 4).
Limit of detection is the lowest amount of analyte that is statistically distinguishable from background or a negative control. Limit of quantification on the other hand is the lowest and highest concentrations of analyte that can be quantitatively determined with suitable precision and accuracy.63 In order to claim limit of detection, a signal to noise level above three must be obtained, whereas limit of quantification requires a signal to noise level above 10. The signal to noise levels verify that MDA-MB-468 is detectable, however at the limit of the system, whereas MDA-MB-231 represents limit of quantification (Table 2 and Figure 4). The measurements on the cell lines have a coefficient of variation ranging from 4 to 15%, which compares well to immunoassays used for blood testing, for example.
Reproducibility and Robustness
To be able to trust a quantitative method, the method should be reproducible and not be affected by day-to-day variation. In the clinical pathology laboratory, the same formalin-fixed, paraffin-embedded block should return the same result if the specimen needs to be reassessed at a later date. We combined reproducibility (testing for consistent results when performing the same HER2 qIHC procedure on the same instrument) with robustness testing to mimic the process in a clinical pathology laboratory. Guidelines point toward 20 separate runs in order to evaluate day-to-day variability, but in this study a smaller initial assessment of run variation was assessed. Sections from the same formalin-fixed, paraffin-embedded block containing the SK-BR-3 cell line were cut and processed on five different days. The sectioning was done by different trained operators. Each of the five days, 12 sections were cut from the formalin-fixed, paraffin-embedded block (nine for HER2 qIHC testing, and three serving as negative controls). The slides were stained with qIHC method the same day as the sections were cut. Antibody retrieval and qIHC staining were performed on the same instrument each time, but four different trained technicians performed the staining (setup according to Figure 5). The overall % CV for all five days was 6.21 %, and the within run variation from the five different test days ranged from 3.94% to 8.48% (Figure 6 and Table 3).
Combined reproducibility and robustness testing of HER2 qIHC. Twelve cut sections from one FFPE cell block were cut on 5 different days by different trained operators. Nine slides were tested with HER2 qIHC, three slides were tested without HER2 antibody (negative control). Four different technicians tested the slides with HER2 qIHC. Antibody retrieval and qIHC were performed on the same instruments.
Repeatability
To further assess within-run reproducibility, we examined a small formalin-fixed, paraffin-embedded cell block containing cell pellets of all five individual cell lines embedded. Each staining run was comprised of nine HER2 qIHC stainings, as well as three negative control stains. Staining 1 was plotted against staining 2–5, resulting in a linear plot, demonstrating high repeatability without any significant day-to-day variation (Figure 6).
All five runs were compared using two-way ANOVA and Turkey post test for multiple comparisons, resulting in all P-values of>0.999, thus no significant difference between each run. Pearson correlation demonstrated statistical significant correlation and thus linearity (Pearson’s r: 0.9995–0.9998, R2: 0.9991–0.999, P: 0.0002–0.0005).
Linearity and Correlation to Reference Quantitative Analytical Methods
The ability of the test to return values that are directly proportional to the concentration of the analyte in the sample is of great importance for quantitative methods. The standard approach for determining linearity is using a dilution, as suggested by Clinical and Laboratory Standards Institute guideline EP6-A,64 however it is not possible to dilute a formalin-fixed, paraffin-embedded cell pellet, or tissue section, thus a setup using other quantitative methods was needed in order to be able to demonstrate linearity. Two established reference methods were used, and if both reference methods have linear correlation, qIHC must correlate to both methods in a linear fashion to demonstrate linearity. The dynamic range and linearity of the assay was investigated by correlating the qIHC on repetitive runs on all five cell lines in nine replicates to results of an in-house developed flow cytometry assay as well as a commercial available ELISA kit for HER2 measurement. See Figure 7 for linearity plots.
Linearity of HER2 protein measurements using ELISA and flow cytometry resulted in statistical significant correlation for all methods (P<0.0001). I: qIHC correlation to flow cytometry: Pearson’s r=0.9813, P<0.0001. II: qIHC correlation to ELISA Pearson’s r: 0.9918, P<0.0001. III: ELISA correlation to flow cytometry: Pearson’s r=0.9967, P<0.0001.
All tests demonstrated statistical significant correlation with P<0.0001 (Pearson’s r: qIHC vs flow cytometry: r=0.9813, qIHC vs ELISA: r=0.9918 and ELISA vs flow cytometry: r=0.9967). No overlap was observed between the different cell lines, and a broad dynamic range could be demonstrated ranging from 4400 receptors/cell up to 565 000 receptors/cell.
HER2 Breast Tumor Concordance
In order to evaluate qIHC on tissue specimens, consecutive formalin-fixed, paraffin-embedded sections were cut from 44 breast cancer resection specimens and tested in triplicates with either HercepTest or HER2 qIHC, using the same HER2 antibody and antigen retrieval. The HER2 qIHC antibody used is the same as for HercepTest. Several tissue sections stained by HER2 qIHC assay was examined thoroughly by trained observers to verify that dots were located in relevant tissue areas (see Figure 2), ie, the same areas as HercepTest. Control stainings without HER2 antibody were also performed, and only a few single dots evenly spread over tissue could be observed.
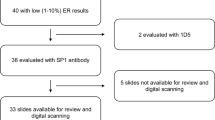
All samples were stained on the same instrument. The 44 breast cancer specimens were selected on basis of their HER2 immunohistochemistry score (HER2 immunohistochemistry distribution Figure 8). For examples of immunohistochemistry and qIHC-stained specimens, respectively, please see Figure 9. Slides were analyzed using histoscore algorithm (H-score), which is a semi-quantitative indexation of the immunohistochemistry HER2-staining intensities.
A one-way ANOVA with Turkeys post test for multiple comparisons demonstrated that all four categories could be separated from each other (adjusted P-values: 0.0003–<0.0001).
To ensure a correct correlation without any tissue variation, the same tumor regions were evaluated on all six slides (three slides of HER2 immunohistochemistry and three slides of HER2 qIHC). Correlating qIHC with traditional immunohistochemistry resulted in a statically significant correlation (Figure 10, Pearson’s r: 0.9606, P: <0.0001). By analyzing the plot, it is evident that while HER2 qIHC can quantify protein expression also in the high range, traditional immunohistochemistry reaches a saturation level (3+ for all cells). Immunohistochemistry is recognized as having a limited dynamic range. This feature does not have any clinical impact for HercepTest since it is designed to robustly separate 0, 1+ (HER2 negative) from 2+ (HER2 equivocal, which needs to be tested with fluorescence in situ hybridization), and 3+ (HER2 positive, which is eligible for trastuzumab treatment). Nevertheless, the ability to separate both high and low levels of HER2 protein present in tumors with qIHC is evidence for an enhanced dynamic range on tissue compared with immunohistochemistry.
Discussion
IHC-based methods are appealing from a practical perspective. Factors such as ease of use, turnaround time, cost, and morphological information are reasons for the widespread use of IHC as a routine test. However, traditional IHC is semi-quantitative at best, which has been evaluated in different studies.65, 66, 67, 68, 69, 70, 71 Assessment of qIHC-stained tissue sections is performed using bright-field microscopy displaying intact and assessable tissue morphology. The concept of dot counting circumvents potential issues with linearity of DAB intensity, which has been discussed in literature.72, 73 Another potential benefit relates to robustness. Small changes in the reagents, lab-to-lab variation etc, can have a significant effect on a classic IHC stain in the form of lower or higher staining intensity. This is not the case for qIHC, here the size of the dot will change in response to the variation. Most importantly, the dot will still be there and will be counted as a dot. Furthermore, if an image analysis algorithm is used to count the dots on digitally scanned slides, the risk of interpretation bias is removed. However, like immunohistochemistry, accurate results of qIHC are dependent upon antibodies with high specificity and standardization of pre-analytical factors.
At the present time, no existing guideline includes analytical validation of quantitative protein measurements in tumor tissue. Internal or external standards do not exist; instead correlation has been made to other methods using similar references. In this study, HER2 levels in cells harvested in the same passage were analyzed with three different techniques. Flow cytometry applied on living cells and ELISA on cell lysates were used as reference methods and compared with the results of qIHC HER2 analysis on formalin-fixed cells embedded in paraffin. Keeping the analyte as similar as possible allows for a correlation of the obtained results across different techniques. It must be stated, however, that a variation in protein levels can be expected as the methods measure HER2 in different sample types (ie, living cells for flow cytometry, lysed cells for ELISA and formalin-fixed, paraffin-embedded cells for IHC and qIHC).
A practical feature of qIHC is the ability to tune the assay for the biomarker of interest, ie, adjust the ratio between labeled and unlabeled secondary antibody. If the biomarker is a ‘low expressor’, such as HER2 in MDA-MB-468, it is possible to amplify a larger percentage of receptors, whereas for a highly expressed biomarker, such as HER2 in SK-BR-3, an assay where a smaller fraction of the antigens are converted into red dots can be used. In this way, a simple titration enables adjustment of the assay to optimize the performance in concordance with clinical need. For this study, the assay was titrated to show dynamic range and the same titration was used for all experiments.
qIHC was tested under requirements for an analytical quantitative test, and compared with established methods for quantitative protein measurements. Reproducibility and robustness were assessed in a setup to resemble daily work in a laboratory using a commercial IHC platform. For all evaluated parameters, qIHC performance was either comparable to or better than the reference methods. It was thereby demonstrated by means of a defined analytical validation and verification process, based on current standards and guidelines, that the qIHC test is a quantitative, robust, and precise test that can be performed reproducibly and accurately.
The lower end of the dynamic range is limited by the ability to separate a data point from a blank sample. In the high end of the qIHC assay, dots overlap to an extent that the algorithm cannot separate them as individual and the readout at the maximum detection level will no longer be within its linear range. In this study, HER2 could be measured using qIHC even in IHC-negative cell lines, and at the same time it was possible to separate higher levels of HER2 protein in the tumors, which is traditionally defined as 3+ in IHC. None of this is possible with IHC, concluding that qIHC has a substantially broader dynamic range than IHC. The ability to cover the dynamic range tested in this study is an advantage compared with, for example, previously reported results with the AQUA technique, a fluorescence technique, which applies two different assays depending on the HER2 concentration.31 Furthermore, qIHC has a lower limit of detection than both IHC and the ELISA reference method. The high sensitivity and larger dynamic range compared with IHC allows for a better separation with greater precision of protein expression. Being able to differentiate even low levels of HER2 could open up applications of new HER2 targeted therapies, which are targeted to the patient population without benefit of trastuzumab or other high HER2 levels targeted therapies. Two clinical studies, NSABP B-47 (NCT01275677) and NeuVax (NCT015170036), are currently in progress, which target low levels (IHC 1+ and 2+) of HER2 in breast cancer. Using a method such as qIHC could potentially allow accurate assessment of small differences in low levels of HER2 in the HER2-negative patient population. With HER2 qIHC, it is possible to measure the low levels of HER2 expressed in the cell lines MDA-MB-468, which are not measurable with ELISA, and MDA-MB-231, which is the de facto IHC 0 control cell line in Food and Drug Administration-approved HER2 IHC products. We have shown that the two cell lines can be measured and quantified, and the results are easily separated.
To demonstrate the utility of qIHC in a clinical setting, a successful comparison with HER2 IHC was done by performing a study on a small cohort of 44 breast cancer patients. The aim was to show that qIHC could give a result correlating to HER2 IHC scores used in clinics today, no analytical parameters were challenged. Despite the fact that HER2 IHC is designed for optimal separation and sensitivity in the equivocal range, we found a good correlation between qIHC and IHC. In the HER2-overexpressed range, traditional IHC was no longer able to differentiate the HER2 levels (no clinical impact). This was not a challenge for qIHC with its broader dynamic range.
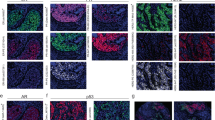
Other biomarkers including membrane, cytoplasm, and nuclear markers have been tested with qIHC in our laboratory. In a similar way as for HER2, all those biomarkers followed the same staining pattern as their corresponding IHC stain (some examples in Figure 11).
In conclusion, the results show that qIHC provides a sensitive, quantitative, accurate, and robust assay for measurement of protein expression in formalin-fixed, paraffin-embedded cell lines and tissue. It is potentially a valuable tool for patient selection in clinical investigations. The use of a more sensitive and quantitative test could provide better and more precise diagnosis and thus improved patient care.
References
Ruschoff J, Hanna W, Bilous M, et al. HER2 testing in gastric cancer: a practical approach. Mod Pathol 2012; 25: 637–650.
Moelans CB, de Weger RA, Van der Wall E, et al. Current technologies for HER2 testing in breast cancer. Crit Rev Oncol Hematol 2011; 80: 380–392.
Albarello L, Pecciarini L, Doglioni C . HER2 testing in gastric cancer. Adv Anat Pathol 2011; 18: 53–59.
Provenzano E, Johnson N . Overview of recommendations of HER2 testing in breast cancer. Diagn Histopathol 2009; 15: 478–484.
Freudenberg JA, Wang Q, Katsumata M, et al. The role of HER2 in early breast cancer metastasis and the origins of resistance to HER2-targeted therapies. Exp Mol Pathol 2009; 87: 1–11.
Cuadros M, Villegas R . Systematic review of HER2 breast cancer testing. Appl Immunohistochem Mol Morphol 2009; 17: 1–7.
Barron JJ, Cziraky MJ, Weisman T, et al. HER2 testing and subsequent trastuzumab treatment for breast cancer in a managed care environment. Oncologist 2009; 14: 760–768.
van de Vijver M, Bilous M, Hanna W, et al. Chromogenic in situ hybridisation for the assessment of HER2 status in breast cancer: an international validation ring study. Breast Cancer Res 2007; 9: R68.
Schnitt SJ, Jacobs TW . Current status of HER2 testing: caught between a rock and a hard place. Am J Clin Pathol 2001; 116: 806–810.
Wolff AC, Hammond ME, Schwartz JN, et al. American Society of Clinical Oncology/College of American Pathologists guideline recommendations for human epidermal growth factor receptor 2 testing in breast cancer. J Clin Oncol 2007; 25: 118–145.
Roche PC, Suman VJ, Jenkins RB, et al. Concordance between local and central laboratory HER2 testing in the breast intergroup trial N9831. J Natl Cancer Inst 2002; 94: 855–857.
Reddy JC, Reimann JD, Anderson SM, et al. Concordance between central and local laboratory HER2 testing from a community-based clinical study. Clin Breast Cancer 2006; 7: 153–157.
Press MF, Sauter G, Bernstein L, et al. Diagnostic evaluation of HER-2 as a molecular target: an assessment of accuracy and reproducibility of laboratory testing in large, prospective, randomized clinical trials. Clin Cancer Res 2005; 11: 6598–6607.
Perez EA, Suman VJ, Davidson NE, et al. HER2 testing by local, central, and reference laboratories in specimens from the North Central Cancer Treatment Group N9831 intergroup adjuvant trial. J Clin Oncol 2006; 24: 3032–3038.
Wludarski SC, Lopes LF, Berto ESTR, et al. HER2 testing in breast carcinoma: very low concordance rate between reference and local laboratories in Brazil. Appl Immunohistochem Mol Morphol 2011; 19: 112–118.
Bast RC, Bates S, Bredt AB, et al. Clinical practice guidelines for the use of tumor markers in breast and colorectal cancer. J Clin Oncol 1996; 14: 2843–2877.
Dowsett M, Nielsen TO, A'Hern R, et al. Assessment of Ki67 in breast cancer: recommendations from the International Ki67 in Breast Cancer working group. J Natl Cancer Inst 2011; 103: 1656–1664.
Harris L, Fritsche H, Mennel R, et al. American Society of Clinical Oncology 2007 update of recommendations for the use of tumor markers in breast cancer. J Clin Oncol 2007 25: 5287–5312.
Tendal B, Higgins JP, Juni P, et al. Disagreements in meta-analyses using outcomes measured on continuous or rating scales: observer agreement study. BMJ 2009; 339: b3128.
Mudduwa L, Liyanage T . Immunohistochemical assessment of hormone receptor status of breast carcinoma: interobserver variation of the quick score. Indian J Med Sci 2009; 63: 21–27.
Kirkegaard T, Edwards J, Tovey S, et al. Observer variation in immunohistochemical analysis of protein expression, time for a change? Histopathology 2006; 48: 787–794.
Meijer SL, Wesseling J, Smit VT, et al. HER2 gene amplification in patients with breast cancer with equivocal IHC results. J Clin Pathol 2011; 64: 1069–1072.
Bueno-de-Mesquita JM, Nuyten DS, Wesseling J, et al. The impact of inter-observer variation in pathological assessment of node-negative breast cancer on clinical risk assessment and patient selection for adjuvant systemic treatment. Ann Oncol 2010; 21: 40–47.
Umemura S, Osamura RY, Akiyama F, et al. What causes discrepancies in HER2 testing for breast cancer? A Japanese ring study in conjunction with the global standard. Am J Clin Pathol 2008; 130: 883–891.
Gavrielides MA, Gallas BD, Lenz P, et al. Observer variability in the interpretation of HER2/neu immunohistochemical expression with unaided and computer-aided digital microscopy. Arch Pathol Lab Med 2011; 135: 233–242.
Varga Z, Diebold J, Dommann-Scherrer C, et al. How reliable is Ki-67 immunohistochemistry in grade 2 breast carcinomas? A QA study of the swiss working group of breast- and gynecopathologists. PLoS One 2012; 7: e37379.
Mandrekar JN . Measures of interrater agreement. J Thorac Oncol 2011; 6: 6–7.
Voros A, Csorgo E, Nyari T, et al. An intra- and interobserver reproducibility analysis of the Ki-67 proliferation marker assessment on core biopsies of breast cancer patients and its potential clinical implications. Pathobiology 2013; 80: 111–118.
Fonyad L, Krenacs T, Nagy P, et al. Validation of diagnostic accuracy using digital slides in routine histopathology. Diagn Pathol 2012; 7: 35.
Camp RL, Dolled-Filhart M, King BL, et al. Quantitative analysis of breast cancer tissue microarrays shows that both high and normal levels of HER2 expression are associated with poor outcome. Cancer Res 2003; 63: 1445–1448.
McCabe A, Dolled-Filhart M, Camp RL, et al. Automated quantitative analysis (AQUA) of in situ protein expression, antibody concentration, and prognosis. J Natl Cancer Inst 2005; 97: 1808–1815.
Giltnane JM, Molinaro A, Cheng H, et al. Comparison of quantitative immunofluorescence with conventional methods for HER2/neu testing with respect to response to trastuzumab therapy in metastatic breast cancer. Arch Pathol Lab Med 2008; 132: 1635–1647.
Gustavson MD, Bourke-Martin B, Reilly D, et al. Standardization of HER2 immunohistochemistry in breast cancer by automated quantitative analysis. Arch Pathol Lab Med 2009; 133: 1413–1419.
Harigopal M, Barlow WE, Tedeschi G, et al. Multiplexed assessment of the Southwest Oncology Group-directed Intergroup Breast Cancer Trial S9313 by AQUA shows that both high and low levels of HER2 are associated with poor outcome. Am J Pathol 2010; 176: 1639–1647.
Huang W, Reinholz M, Weidler J, et al. Comparison of central HER2 testing with quantitative total HER2 expression and HER2 homodimer measurements using a novel proximity-based assay. Am J Clin Pathol 2010; 134: 303–311.
Larson JS, Goodman LJ, Tan Y, et al. Analytical validation of a highly quantitative, sensitive, accurate, and reproducible assay (HERmark) for the measurement of HER2 total protein and HER2 homodimers in FFPE breast cancer tumor specimens. Patholog Res Int 2010; 2010: 814176.
DeFazio-Eli L, Strommen K, Dao-Pick T, et al. Quantitative assays for the measurement of HER1-HER2 heterodimerization and phosphorylation in cell lines and breast tumors: applications for diagnostics and targeted drug mechanism of action. Breast Cancer Res 2011; 13: R44.
Joensuu H, Sperinde J, Leinonen M, et al. Very high quantitative tumor HER2 content and outcome in early breast cancer. Ann Oncol 2011; 22: 2007–2013.
Zanconati F, Cusumano P, Tinterri C, et al. P205 the 70-gene expression profile, mammaprint, for breast cancer patients in mainly European hospitals. Breast 2011; 20: S45.
Kok M, Koornstra RH, Mook S, et al. Additional value of the 70-gene signature and levels of ER and PR for the prediction of outcome in tamoxifen-treated ER-positive breast cancer. Breast 2012; 21: 769–778.
Saghatchian M, Mook S, Pruneri G, et al. Additional prognostic value of the 70-gene signature (MammaPrint((R))) among breast cancer patients with 4-9 positive lymph nodes. Breast 2013; 22: 682–690.
Torrisi R, Garcia-Etienne CA, Losurdo A et al, Potential impact of the 70-gene signature in the choice of adjuvant systemic treatment for ER positive, HER2 negative tumors: a single institution experience. Breast 2013; 22: 419–424.
Cronin M, Sangli C, Liu ML, et al. Analytical validation of the Oncotype DX genomic diagnostic test for recurrence prognosis and therapeutic response prediction in node-negative, estrogen receptor-positive breast cancer. Clin Chem 2007; 53: 1084–1091.
Tang G, Shak S, Paik S, et al. Comparison of the prognostic and predictive utilities of the 21-gene Recurrence Score assay and Adjuvant! for women with node-negative, ER-positive breast cancer: results from NSABP B-14 and NSABP B-20. Breast Cancer Res Treat 2011; 127: 133–142.
Ignatiadis M, Sotiriou C . Breast cancer: should we assess HER2 status by ocotype DX? Nat Rev Clin Oncol 2012; 9: 12–14.
Markopoulos C, Xepapadakis G, Venizelos V, et al. Clinical experience of using Oncotype DX as an additional treatment decision tool in early breast cancer - a retrospective analysis from 5 Greek institutions. Eur J Surg Oncol 2012; 38: 413–419.
Ingoldsby H, Webber M, Wall D, et al. Prediction of Oncotype DX and TAILORx risk categories using histopathological and immunohistochemical markers by classification and regression tree (CART) analysis. Breast 2013; 22: 879–886.
Nielsen TO, Parker JS, Leung S, et al. A comparison of PAM50 intrinsic subtyping with immunohistochemistry and clinical prognostic factors in tamoxifen-treated estrogen receptor-positive breast cancer. Clin Cancer Res 2010; 16: 5222–5232.
Bastien RR, Rodriguez-Lescure A, Ebbert MT, et al. PAM50 breast cancer subtyping by RT-qPCR and concordance with standard clinical molecular markers. BMC Med Genomics 2012; 5: 44.
Dowsett M, Sestak I, Lopez-Knowles et al. Comparison of PAM50 risk of recurrence score with oncotype DX and IHC4 for predicting risk of distant recurrence after endocrine therapy. J Clin Oncol 2013; 31: 2783–2790.
Cuzick J, Dowsett M, Pineda S, et al. Prognostic value of a combined estrogen receptor, progesterone receptor, Ki-67, and human epidermal growth factor receptor 2 immunohistochemical score and comparison with the Genomic Health recurrence score in early breastcancer. J Clin Oncol 2011; 29: 4273–4278.
Dowsett M, Salter J, Zabaglo L, et al. Predictive algorithms for adjuvant therapy: TransATAC. Steroids 2011; 76: 777–780.
Barton S, Zabaglo L, A'Hern R, et al. Assessment of the contribution of the IHC4+C score to decision making in clinical practice in early breast cancer. Br J Cancer 2012; 106: 1760–1765.
Lohse J, Petersen KH, Woller NC, et al. Improved catalyzed reporter deposition, iCARD. Bioconjug Chem 2014; 25: 1036–1042.
Wolff AC, Hammond ME, Hicks DG, et al. Recommendations for human epidermal growth factor receptor 2 testing in breast cancer: American Society of Clinical Oncology/College of American Pathologists clinical practice guideline update. J Clin Oncol 2013; 31: 3997–4013.
Williams PM, Lively TG, Jessup JM, et al. Bridging the gap: moving predictive and prognostic assays from research to clinical use. Clin Cancer Res 2012; 18: 1531–1539.
Jennings L, Van Deerlin VM, Gulley ML, College of American Pathologists Molecular Pathology Resource C. Recommended principles and practices for validating clinical molecular pathology tests. Arch Pathol Lab Med 2009; 133: 743–755.
Guideline T . Q2B validation of analytical procedures: methodology. Fed Regist 1997; 62: 27463–27467.
Peters FT, Drummer OH, Musshoff F . Validation of new methods. Forensic Sci Int 2007; 165: 216–224.
Shi Y, Huang W, Tan Y, et al. A novel proximity assay for the detection of proteins and protein complexes: quantitation of HER1 and HER2 total protein expression and homodimerization in formalin-fixed, paraffin-embedded cell lines and breast cancer tissue. Diagn Mol Pathol 2009; 18: 11–21.
Rait AS, Pirollo KF, Rait V, et al. Inhibitory effects of the combination of HER-2 antisense oligonucleotide and chemotherapeutic agents used for the treatment of human breast cancer. Cancer Gene Ther 2001; 8: 728–739.
CLSI Evaluation of Precision Performance of Quantitative Measurement Methods; Approved Guideline, 2nd edn. EP5-A3 ed. Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2014.
CLSI Evaluation of detection capability for clinical laboratory measurement procedures: Approved guideline . EP17-A2 ed. Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2012.
CLSI Evaluation of the Linearity of Quantitative Measurement Procedures: A Statistical Approach; Approved Guideline. EP6-A ed. Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2003.
Ellis CM, Dyson MJ, Stephenson TJ et al, HER2 amplification status in breast cancer: a comparison between immunohistochemical staining and fluorescence in situ hybridisation using manual and automated quantitative image analysis scoring techniques. J Clin Pathol 2005; 58: 710–714.
Hall BH, Ianosi-Irimie M, Javidian P, et al. Computer-assisted assessment of the human epidermal growth factor receptor 2 immunohistochemical assay in imaged histologic sections using a membrane isolation algorithm and quantitative analysis of positive controls. BMC Med Imaging 2008; 8: 11.
Aitken SJ, Thomas JS, Langdon SP, et al. Quantitative analysis of changes in ER, PR and HER2 expression in primary breast cancer and paired nodal metastases. Ann Oncol 2010; 21: 1254–1261.
Lloyd MC, Allam-Nandyala P, Purohit CN, et al. Using image analysis as a tool for assessment of prognostic and predictive biomarkers for breast cancer: How reliable is it? J Pathol Inform 2010; 1: 29.
Laurinaviciene A, Dasevicius D, Ostapenko V et al, Membrane connectivity estimated by digital image analysis of HER2 immunohistochemistry is concordant with visual scoring and fluorescence in situ hybridization results: algorithm evaluation on breast cancer tissue microarrays. Diagn Pathol 2011; 6: 87.
Di Cataldo S, Ficarra E, Macii E . Computer-aided techniques for chromogenic immunohistochemistry: status and directions. Comput Biol Med 2012; 42: 1012–1025.
Bechmann T, Olsen DA, Jakobsen EH et al, Quantitative detection of HER2 protein concentration in breast cancer tissue does not increase the number of patients eligible for adjuvant HER2-targeted therapy. Oncol Rep 2013; 29: 1475–1482.
van der Loos CM . Multiple immunoenzyme staining: methods and visualizations for the observation with spectral imaging. J Histochem Cytochem 2008; 56: 313–328.
Rimm DL . What brown cannot do for you. Nat Biotechnol 2006; 24: 914–916.
Acknowledgements
We would like to thank for the laboratory assistance from Marcussen, M, Bruun, A, Maarbjerg, AM, and Thuran, N.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Jensen, K., Krusenstjerna-Hafstrøm, R., Lohse, J. et al. A novel quantitative immunohistochemistry method for precise protein measurements directly in formalin-fixed, paraffin-embedded specimens: analytical performance measuring HER2. Mod Pathol 30, 180–193 (2017). https://doi.org/10.1038/modpathol.2016.176
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2016.176
This article is cited by
-
Quantitative performance assessment of Ultivue multiplex panels in formalin-fixed, paraffin-embedded human and murine tumor specimens
Scientific Reports (2024)
-
Spatially resolved multimarker evaluation of CD274 (PD-L1)/PDCD1 (PD-1) immune checkpoint expression and macrophage polarisation in colorectal cancer
British Journal of Cancer (2023)
-
Overexpression of microRNA-205-5p promotes cholangiocarcinoma growth by reducing expression of homeodomain-interacting protein kinase 3
Scientific Reports (2023)
-
High-plex spatial transcriptomic profiling reveals distinct immune components and the HLA class I/DNMT3A/CD8 modulatory axis in mismatch repair-deficient endometrial cancer
Cellular Oncology (2023)
-
β-Elemene alleviates cisplatin resistance in oral squamous cell carcinoma cell via inhibiting JAK2/STAT3 pathway in vitro and in vivo
Cancer Cell International (2022)