Abstract
MYC dysregulation, including MYC gene rearrangement and Myc protein overexpression, is of increasing clinical importance in diffuse large B-cell lymphoma (DLBCL). However, the roles of MYC and the relative importance of rearrangement vs overexpression remain to be refined. Gaining knowledge about the tumor biology associated with MYC dysregulation is important to understand the roles of MYC and MYC-associated biology in lymphomagenesis. In this study, we determined MYC rearrangement status (n=344) and Myc expression (n=535) in a well-characterized DLBCL cohort, individually assessed the clinical and pathobiological features of patients with MYC rearrangement and Myc protein overexpression, and analyzed the prognosis and gene expression profiling signatures associated with these MYC abnormalities in germinal center B-cell-like and activated B-cell-like DLBCL. Our results showed that the prognostic importance of MYC rearrangement vs Myc overexpression is significantly different in germinal center B-cell-like vs activated B-cell-like DLBCL. In germinal center B-cell-like DLBCL, MYC-rearranged germinal center B-cell-like DLBCL patients with Myc overexpression significantly contributed to the clinical, biological, and prognostic characteristics of the overall Myc-overexpressing germinal center B-cell-like DLBCL group. In contrast, in activated B-cell-like DLBCL, the occurrence, clinical and biological features, and prognosis of Myc overexpression were independent of MYC rearrangement. High Myc levels and Myc-independent mechanisms, either tumor cell intrinsic or related to tumor microenvironment, conferred significantly worse survival to MYC-rearranged germinal center B-cell-like DLBCL patients, even among MychighBcl-2high DLBCL patients. This study provides new insight into the tumor biology and prognostic effects associated with MYC dysregulation and suggest that detection of both MYC translocations and evaluation of Myc and Bcl-2 expression is necessary to predict the prognosis of DLBCL patients.
Similar content being viewed by others
Main
MYC is a proto-oncogene that encodes the Myc protein, which is critical for cell proliferation, growth, metabolism, differentiation, apoptosis, and immune responses. In mouse models, Myc inactivation induces sustained tumor regression via both tumor cell-intrinsic and host-dependent mechanisms.1 The selective small-molecule bromodomain inhibitor JQ1 has a potent antiproliferative effect that was associated with the effective downregulation of MYC and Myc target genes2 rendering this agent to have great therapeutic potential.3
The chromosomal rearrangement or translocation involving MYC and other genes (most commonly the immunoglobulin heavy-chain gene (IGH) locus) leads to Myc overexpression and occurs in ~10% of diffuse large B-cell lymphomas (DLBCLs).4, 5, 6, 7, 8 DLBCL is the most common type of non-Hodgkin's lymphoma, and among MYC-rearranged aggressive lymphomas, DLBCL is the entity that clinicians most commonly encounter.9, 10 Several studies have reported that MYC translocations independently predicted significantly poor survival in DLBCL patients.11, 12, 13, 14, 15, 16 However, other studies found inconsistent results17, 18, 19 or limitations of its prognostic significance.20, 21
The clinical significance of Myc overexpression in DLBCL has also been the source of much attention and controversy. Several groups including ours have found that DLBCL with high Myc protein expression detected by immunohistochemistry had inferior survival.5, 13, 22, 23, 24 Furthermore, the poor prognosis associated with Myc overexpression was contributed by cases with Myc/Bcl-2 coexpression—‘double-positive lymphoma’ (DPL)—which account for 18–44% of DLBCLs.13, 20, 23, 24, 25, 26 However, one study showed that the prognostic value of double-positive lymphoma was lost in younger DLBCL patients with poor prognosis.27 Inconsistent results have been reported regarding whether the prognostic significance of Myc or Myc/Bcl2 protein expression depends on MYC or MYC/BCL2 gene rearrangement status or not.8, 13, 20 Other issues include whether Myc/Bcl2 immunohistochemistry is robust and reproducible,28 and that immunohistochemistry cutoff values have varied among different study groups,13, 20, 23, 24, 25, 26 which may affect the specificity of this combined biomarker for poorer prognosis.
Possible molecular mechanisms underlying the inconsistent clinical results may include presence or absence of other genetic abnormalities and oncogenic pathways,29, 30, 31 as well as another aspect of Myc function: promoting apoptosis.32 Moreover, researchers have recently reported findings that Myc is a universal amplifier of 10–15% in human genome, suggesting that Myc function is nonspecific and that the consequences of MYC activities are affected by pre-existing molecular programs in the tumor cells.33, 34 Therefore, tumor biology associated with MYC/Myc (designated MYC herein) abnormalities may have important roles in the observed adverse prognosis.
Taken together, both MYC gene rearrangement and Myc protein overexpression have been correlated with significantly adverse prognosis in DLBCL. However, how much of these two biomarkers overlap and differ, how much their associated tumor biology affects the prognostic effects, and whether the MYC functional role is molecular context-dependent are not very clear. In this study, we compared the occurrence and clinicopathologic features of patients stratified by MYC rearrangement and Myc expression status, and analyzed the differential prognosis and gene expression profiling associated with these MYC abnormalities in a well-characterized DLBCL cohort to assess the utility of these two genetic and protein biomarkers and explore the prognostic determinants. This study is important for achieving the goal of precision medicine in DLBCL.
Patients and methods
Patients
The study cohort consisted of 539 R-CHOP-treated patients with de novo DLBCL from the International DLBCL Rituximab-CHOP Consortium Program, including 466 cases from the training set of a previous study,24 and additional 73 cases with either Myc immunohistochemistry or MYC gene rearrangement status determined. The diagnostic criteria, review process, and eligibility and exclusion criteria have been described previously.35, 36 The cell-of-origin classification as either the germinal center B-cell-like (GCB) or activated B-cell-like (ABC) subtype was determined using gene expression profiling and/or immunohistochemistry for CD10, BCL6, GCET-1, FOXP1, and MUM1 using the Visco-Young and/or Choi algorithms as described previously.24, 35, 36 Totally, 276 cases were classified as GCB, 259 cases were classified as ABC, and 4 cases were unclassifiable. All patients underwent standard R-CHOP or R-CHOP-like therapy, and the median follow-up time was 45 months (range, 30–176.1 months). This study was conducted in accordance with the Declaration of Helsinki and was approved as being of minimal to no risk or as exempt by the institutional review boards of all participating centers, including The University of Texas MD Anderson Cancer Center.
Fluorescence In Situ Hybridization, Immunohistochemistry, and Gene Sequencing
MYC translocation was detected by fluorescence in situ hybridization using two probes (a locus-specific identifier IGH/MYC/CEP8 tri-color dual-fusion probe and a locus-specific identifier MYC dual-color break-apart probe) (n=344). Myc expression was assessed by immunohistochemistry using tissue microarray sections and a monoclonal anti-(c)MYC antibody, clone Y69 (Epitomics, Burlingame, CA, USA) (n=535). The experimental techniques and scoring processes have been described previously.15, 21, 24
Evaluation of other biomarker expression by immunohistochemistry was also performed on tissue microarray sections using corresponding antibodies: p53 (DO-7; Dako, Carpinteria, CA, USA), MDM2 (IF2; Calbiochem, Billerica, MA, USA), Bcl-2 (Clone-124; Dako), Ki-67 (MIB-1; Dako), pAKT (726E11; CST), Bcl-6 (PG-B6p; DAKO), FOXP1 (EPR4113; Abcam), MUM1/IRF4 (Dako), CD10 (56C6; Vantana), CD30 (clone BerH2; Dako), BLIMP-1 (EPR16655; Epitomics), NF-κB subunits (Dako), CXCR4 (Abcam, Cambridge, MA, USA), and survivin (EP2880Y; Epitomics). BCL6 and BCL2 translocations and TP53 mutations were detected as described previously.15, 21, 24, 35, 36, 37, 38, 39, 40, 41, 42
Gene Expression Profiling
Gene expression profiling for 457 patients was performed using the Affymetrix GeneChip Human Genome HG-U133 Plus 2.0 Array as described previously.21, 24, 35, 36, 37 The CEL files have been deposited in the National Center for Biotechnology Information Gene Expression Omnibus repository (GSE31312). Microarray data were normalized for further supervised clustering analysis. Multiple t-tests were used to identify differentially expressed genes and the P-values obtained were corrected for the false discovery rate using the β-uniform mixture method.
Statistical Analysis
The clinical and pathobiological features of DLBCL patients at the time of presentation were compared using the χ2 test. The mean expression levels of biomarkers between DLBCL groups with or without MYC abnormalities were also compared by unpaired t-test. Overall survival (OS) was calculated from the time of diagnosis to last follow-up or death from any cause. Progression-free survival (PFS) was calculated from the time of diagnosis to disease progression, disease relapse, or death from any cause. Patients who were alive and disease progression-free at last follow-up were censored. Survival analysis was performed using the Kaplan–Meier method with the Prism 5 program (GraphPad Software, San Diego, CA, USA), and differences in survival were compared using the log-rank (Mantel–Cox) test. Multivariate survival analysis was performed using a Cox proportional hazards regression model with the SPSS software program (version 19.0; IBM Corporation, Armonk, NY, USA). All differences with P≤0.05 were considered statistically significant.
Results
Myc Expression in Germinal Center B-Cell-Like and Activated-B-Cell-Like Subtypes of DLBCL
We observed variable levels of nuclear Myc expression in DLBCL (n=535) (Figures 1a–c). The mean expression level of Myc in ABC-DLBCL was significantly higher compared with that in GCB-DLBCL (Figure 1d). Among the cases successfully tested for MYC rearrangement status (n=344), 27 (16.3%) of 166 GCB-DLBCL cases and 13 (7.3%) of 177 ABC-DLBCL cases had MYC rearrangement/translocation. Most but not all MYC-rearranged cases (MYC-R+) had high MYC-mRNA and Myc expression, with an average percentage of Mychigh nuclei in the tumor samples of 70.5% (Supplementary Figure S1A). We thus set the cutoff for Myc overexpression (Mychigh) at ≥70%, so that Myc expression levels in MYC-non-rearranged (MYC-R−) Mychigh patients comparably ‘matched’ those in MYC-R+/Mychigh cases (Supplementary Figure S1C) in comparisons of clinical and biological significance of Myc overexpression vs MYC translocation.
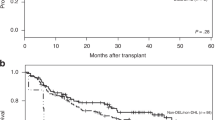
(a–c) Histograms of Myc expression according to immunohistochemistry staining in all diffuse large B-cell lymphoma (DLBCL) patients, germinal center B-cell-like (GCB) DLBCL patients, and activated B-cell-like (ABC) DLBCL patients. (d) The mean Myc protein levels of ABC-DLBCL were significantly higher compared with those of GCB-DLBCL. (e and f) MYC rearrangement-positive (MYC-R+) DLBCL without high Myc immunohistochemistry scores (either GCB or ABC subtype) had significantly lower MYC-mRNA levels. (g) MYC-R+ DLBCL patients without high Myc immunohistochemistry scores had significantly better overall survival compared with MYC-R+/Mychigh DLBCL. Mychigh, high Myc protein expression; Myclow, low Myc protein expression.
Using this cutoff, we found that 175 (32.7%) of 535 DLBCL patients, including 76 (27.9%) of 272 GCB-DLBCL patients and 98 (37.8%) of 259 ABC-DLBCL patients, were Mychigh (Figures 2a and b). Nineteen (73%) of the 26 MYC-R+ GCB patients (one MYC-R+ GCB case had no expression data available) and 7 (54%) of the 13 MYC-R+ ABC patients were Mychigh, who had significantly higher MYC transcripts and significantly worse survival compared with the MYC-R+/Myclow cases (Figures 1e–g and Supplementary Figures S1D). Compared with MYC-R− patients, MYC-R+/Mychigh patients had significantly worse survival in GCB-DLBCL (P=0.0001 for OS and P<0.0001 for PFS) but not in ABC-DLBCL (P=0.56 for OS and P=0.14 for PFS) (Supplementary Figures S1E and F). Comparison of MYC expression levels between the MYC-R+ GCB, MYC-R− GCB, MYC-R+ ABC, and MYC-R− ABC Mychigh groups revealed that MYC-R+/Mychigh GCB-DLBCL had significantly higher Myc protein expression levels compared with all other subsets, and that among both the GCB and ABC subtypes, MYC-R+/Mychigh cases had a significantly higher level of MYC-mRNA levels compared with MYC-R−/Mychigh cases (Figures 2c and d).
Occurrence of MYC rearrangement (MYC-R) and overexpression (Mychigh) and MYC expression levels in diffuse large B-cell lymphoma (DLBCL). (a and b) Schematic diagrams showing the frequencies of MYC translocation (MYC-R+) and Myc overexpression (Mychigh) and their overlaps in germinal center B-cell-like (GCB) and activated B-cell-like (ABC) DLBCL in the current study cohort. (c and d) Comparisons of Myc protein and MYC-mRNA expression levels in DLBCL patients with or without MYC dysregulation. Among Mychigh groups, MYC-R+ GCB-DLBCL had highest Myc protein levels; MYC-R+ DLBCL (both GCB and ABC subtypes) had significantly higher MYC-mRNA levels compared with MYC-R− DLBCL. Note: Each dot represents one patient in the study cohort. MYC-R+, MYC rearrangement-positive; MYC-R−, MYC rearrangement-negative; Mychigh, high Myc protein expression/Myc overexpressing; Myclow, low Myc protein expression; MYC-RN/A, MYC rearrangement status not available.
Clinical Parameters Associated with Myc Overexpression and MYC Rearrangement
We analyzed the clinical parameters associated with Myc overexpression (Table 1) and MYC rearrangement (Supplementary Table 1). We further compared MYC-R+/Mychigh and MYC-R−/Mychigh cases (Supplementary Table S2). Results of significant characteristics are summarized in Supplementary Table S3. Largely, both Myc overexpression and MYC rearrangement could identify a subgroup of patients with adverse clinical features in GCB-DLBCL, whereas patients with these abnormalities did not show such distinct characteristics among ABC-DLBCLs.
For example, in GCB-DLBCL, Mychigh patients compared with Myclow patients more often had stage III/IV disease, ≥2 extranodal sites, ECOG performance status ≥2, tumor size ≥5 cm, International Prognostic Index >2, bone marrow involvement at clinical presentation, and less likely to have complete response. In ABC-DLBCL, Mychigh patients compared with Myclow patients had a higher proportion of women, and did not show significant association with other clinical parameters. The GCB and ABC subtypes of Mychigh patients differed significantly only in their frequencies of patients with age ≥60 years (Table 1). Similarly, in GCB-DLBCL, MYC-R+ patients were enriched in patients with adverse clinical features, whereas in ABC-DLBCL, MYC rearrangement was only associated with primary extranodal origin. The GCB subtype compared with the ABC subtype of MYC-R+ patients showed trends toward having a higher proportion of patients with bone marrow involvement and non-complete treatment response (Supplementary Table S1).
In both the GCB- and ABC-DLBCL groups, MYC-R+/Mychigh patients appeared to have similar clinical features with MYC-R−/Mychigh patients (except complete response rate), yet may have elevated serum lactate dehydrogenase levels and higher frequency of extranodal origin (Supplementary Table S2).
Molecular Biomarkers Associated with Myc Overexpression and MYC Rearrangement
Mychigh DLBCL compared with Myclow DLBCL, in addition to having higher Myc levels and significantly higher frequencies of MYC translocation and ABC subtype (Table 1), had higher expression levels of p53, MDM2, Bcl-2 (Mychigh ABC-DLBCL only), Bcl-6, FOXP1, IRF4/MUM1 (Mychigh ABC-DLBCL only), Ki-67, pAKT, CXCR4, and CD10 (Mychigh GCB-DLBCL only), but lower expression levels of BLIMP-1 (in ABC-DLBCL only) and nuclear expression of the NF-κB subunits c-Rel and RelB (in ABC-DLBCL only, P=0.019, figure not shown) (Figure 3). At the mRNA level, only MDM2 (in GCB-DLBCL only), FOXP1 (in ABC-DLBCL only), IRF4 (in ABC-DLBCL only), and MME/CD10 transcript levels showed corresponding correlations consistent with those at the protein level (Figures 4a–d). PRDM1 mRNA levels were significantly lower in Mychigh GCB-DLBCL compared with that in Myclow GCB-DLBCL (P=0.0005). The GCB and ABC subtypes of Mychigh patients differed in frequencies of MYC translocation, BCL2 translocation, TP53 mutation (all higher in the GCB subtype), and expression levels of Bcl-2, p50, c-Rel (all higher in the ABC subtype), pAKT (higher in the GCB subtype), and cell-of-origin-related biomarkers (Bcl-6, BLIMP-1, GCET-1, CD10, FOXP1, MUM1) (Table 1, Figure 3, and Supplementary Table S3).
Expression of pathobiological markers in germinal center B-cell-like (GCB) and activated B-cell-like (ABC) diffuse large B-cell lymphoma with or without Myc overexpression. In both GCB- and ABC-DLBCL, Myc overexpression was associated with significantly higher levels of p53 (a), MDM2 (b), Ki-67 (d), Bcl-6 (f), and FOXP1 (g), and a significantly lower level of c-Rel (k). In GCB-DLBCL only, Myc overexpression was associated with significantly higher levels of CD10 (e), pAKT (j), and CXCR4 (l). In ABC-DLBCL only, Myc overexpression was associated with significantly higher levels of Bcl-2 (c) and MUM1 (h) levels and a significantly lower level of BLIMP-1 (i). Mychigh, high Myc protein expression; Myclow, low Myc protein expression.
(a–d) Myc overexpression was correlated with significantly higher levels of MDM2, IRF4, and MME/CD10 mRNA expression in germinal center B-cell-like (GCB) diffuse large B-cell lymphoma and significantly higher levels of FOXP1 and IRF4 mRNA expression in activated B-cell-like (ABC) diffuse large B-cell lymphoma (DLBCL). (e) The MYC-R+/Mychigh compared with MYC-R−/Mychigh group had significantly lower levels of MUM1 expression. (f–j) The MDM2, CD30, CD10, and FOXP1 levels in Mychigh GCB-DLBCL patients with MYC translocation were significantly different from those of Mychigh GCB-DLBCL patients without MYC translocation. (k and l) The Bcl-6 and pAKT levels of Mychigh ABC-DLBCL patients with MYC translocation were significantly lower compared with those of Mychigh ABC-DLBCL patients without MYC translocation. Mychigh, high Myc protein expression; Myclow, low Myc protein expression; MYC-R+, MYC rearrangement-positive; MYC-R−, MYC rearrangement-negative; MYC-RN/A, MYC rearrangement status not available.
Some of the pathobiological associations with Mychigh DLBCL were shared by MYC-R+ GCB-DLBCL patients (compared with MYC-R− GCB-DLBCL; Supplementary Tables S1 and S3). On the other hand, in addition to the difference in Myc activation mechanisms (MYC translocation or not), MYC-R+/Mychigh compared with MYC-R−/Mychigh patients had higher frequencies of GCB subtype and BCL2 translocation, and lower frequencies of BCL6 translocation, MUM1, CD30, and p52 expression (Supplementary Table S2 and S3). Among GCB-DLBCL patients, MYC-R+/Mychigh compared with MYC-R−/Mychigh patients had significantly higher expression levels of MDM2, CD10, and FOXP1 but a lower expression level of CD30; among ABC-DLBCL patients, MYC-R+/Mychigh compared with MYC-R−/Mychigh was associated with lower expression levels of Bcl-6 and pAKT (Figures 4f–l).
Prognostic Effect of Myc Overexpression with or without MYC Translocations and Associated Gene Expression Profiling Signatures
We combined the survival analysis with gene expression profiling comparisons to assess the role of MYC abnormalities in DLBCL and to study potential mechanisms. As in earlier studies,13, 22, 23, 24 in the present study, high expression levels of Myc was associated with significantly worse OS and PFS in both GCB- and ABC-DLBCL (Supplementary Figure S2). Similar prognostic impact was also shown by high MYC-mRNA levels in GCB- and ABC-DLBCL (Supplementary Figures S3A–D). Gene expression profiling analysis showed that Mychigh GCB- and ABC-DLBCL had different gene expression profiling signatures (Table 2, Supplementary Figures S2D and F, and Supplementary Table S4A). The significantly differentially expressed genes between these two groups included CDCA7L, IGF2BP3, and RUVBL2, which are known to have roles in the oncogenic transformation by MYC or to interact with MYC.
MYC rearrangement did not show prognostic significance in the Myclow cases [for OS: P=0.25 (OS of MYC-R+/Myclow patients was slightly better than MYC-R−/Myclow DLBCL); for PFS: P=0.71]. Dividing the Mychigh cases with unfavorable prognosis into MYC-R+/Mychigh and MYC-R−/Mychigh two types and comparing their prognosis and gene expression profiling features, we further found that MYC-R+/Mychigh GCB-DLBCL and MYC-R−/Mychigh ABC-DLBCL were the main contributors to the overall worse prognosis and distinct gene expression profiling signatures of the Mychigh GCB-DLBCL and Mychigh ABC-DLBCL groups, respectively.
In the GCB-DLBCL group, MYC-R+/Mychigh patients had significantly worse survival than both Myclow and MYC-R−/Mychigh patients did. Although MYC-R−/Mychigh GCB-DLBCL showed trends towards worse survival compared with the overall Myclow GCB-DLBCL (P=0.40 for OS, Figure 5a; P=0.48 for PFS; Supplementary Figure S4A), and the MYC-R−/Myclow GCB-DLBCL (for OS: P=0.32; Supplementary Figure S4C; for PFS: P=0.21), the differences were not significant. Biologically, only MYC-R+/Mychigh (but not MYC-R−/Mychigh) GCB-DLBCL compared with Myclow GCB-DLBCL showed a distinct gene expression profiling signature (false discovery rate <0.01) (Figure 5b,Supplementary Table S4B, and Table 3), and this signature mostly overlapped the Mychigh gene expression profiling signature identified in the overall GCB-DLBCL group (Table 2 and Supplementary Figure S2D) involving cell proliferation, gene expression, metabolism, apoptosis, microenvironment and immune response, and microRNA genes. In contrast, the MYC-R−/Mychigh GCB-DLBCL patients compared with Myclow GCB-DLBCL (Supplementary Figures S4D, Table 3, and Supplementary Table S4B) or MYC-R+/Mychigh GCB-DLBCL (Figure 5c, Table 3, and Supplementary Table S4B) only showed a few differentially expressed genes below false discovery rate threshold of 0.30 (not including MYC), most of which are involved in cell proliferation, gene expression, ribosome biogenesis, and metabolism, suggesting the presence of heterogeneity and post-transcriptional regulation of MYC as a cause of Myc overexpression within the MYC-R−/Mychigh GCB-DLBCL group. The comparison between MYC-R+ GCB-DLBCL and MYC-R− GCB-DLBCL overall is shown in Supplementary Figure S4E.
Combined prognostic and biologic analysis of germinal-center-B-cell-like (GCB) and activated-B-cell-like (ABC) diffuse large B-cell lymphoma (DLBCL) with or without MYC translocation (MYC-R) and/or Myc overexpression (Mychigh) in the current study cohort. (a) GCB-DLBCL patients with both MYC translocation and Myc overexpression (MYC-R+/Mychigh) had significantly worse overall survival compared with GCB-DLBCL patients with low Myc expression (Myclow) and Mychigh patients without MYC translocation (MYC-R−/Mychigh). The MYC-R−/Mychigh group did not have significant poorer survival compared with the Myclow group. (b) Genes significantly differentially expressed between MYC-R+/Mychigh GCB-DLBCL patients and Myclow GCB-DLBCL patients (false discovery rate <0.01, fold change >2.38). (c) Genes significantly differentially expressed between MYC-R+/Mychigh and MYC-R−/Mychigh GCB-DLBCL patients (false discovery rate <0.30). (d) Only MYC-R−/Mychigh but not MYC-R+/Mychigh ABC-DLBCL patients had significantly OS compared with Myclow ABC-DLBCL patients. (e) Genes significantly differentially expressed between MYC-R−/Mychigh ABC-DLBCL and Myclow ABC-DLBCL (false discovery rate <0.01, fold change >1.57). (f) Genes significantly differentially expressed between ABC and GCB subtypes of MYC-R+ DLBCL. (g) MYC-R+/Mychigh GCB-DLBCL showed trend towards worse progression-free survival compared with MYC-R−/Mychigh ABC-DLBCL patients with a borderline P-value. (h) Genes significantly differentially expressed between MYC-R+ and MYC-R−DLBCL (false discovery rate <0.01, fold change >1.66). (i) Genes significantly differentially expressed between MYC-R+/Mychigh and MYC-R−/Mychigh DLBCL (false discovery rate <0.01, fold change >2.05). ABC, activated-B-cell-like; GCB, germinal-center-B-cell-like; Mychigh, high Myc protein expression; Myclow, low Myc protein expression; MYC-R+: MYC rearrangement-positive; MYC-R−, MYC rearrangement-negative.
In contrast, in ABC-DLBCL, the prognostic significance (Figure 5d) and gene expression profiling features of Myc overexpression did not depend on MYC translocation. MYC-R−/Mychigh ABC-DLBCL had significantly poorer survival compared with the overall Myclow or MYC-R−/Myclow ABC-DLBCL (Figure 5d and Supplementary Figures S4G–I) and distinct gene expression profiling signatures (false discovery rate <0.01; Figure 5e and Supplementary Table S4C). The genes in the MYC-R−/Mychigh ABC-DLBCL gene expression profiling signature, overlapped with differentially expressed genes between the overall Mychigh vs Myclow ABC-DLBCL patients, included typical Myc targets mainly related to cell proliferation, the cell cycle, gene expression, ribosome biogenesis, metabolism (Table 4), and cooperating oncogenes such as RUVBL2, as well as IGF2BP3 involved in post-transcriptional regulation of MYC, and HINT1 modulating p53 levels and the p53 pathway. Different from the MYC-R−/Mychigh ABC-DLBCL cases, MYC-R+/Mychigh ABC-DLBCL compared with overall Myclow ABC-DLBCL only showed nonsignificant trends towards worse OS and PFS (Figure 5d and Supplementary Figures S4G–I) and only a few differentially expressed genes (false discovery rate <0.30, Supplementary Figure S4F). No genes were found differentially expressed between MYC-R+/Mychigh ABC-DLBCL and MYC-R−/Mychigh ABC-DLBCL, or between MYC-R+/Mychigh ABC-DLBCL and the overall MYC-R− ABC-DLBCL group. This is in contrast with the distinct gene expression profiling feature of MYC-R+/Mychigh GCB-DLBCL shown in Figure 5b and Supplementary Figures S4E. Comparison between overall MYC-R+/Mychigh ABC-DLBCL and MYC-R+/Mychigh GCB-DLBCL indicated their different and potentially heterogeneous tumor biology (Figure 5f) (between MYC-R+/Mychigh and MYC-R+/Myclow GCB-DLBCL, or between MYC-R+/Mychigh and MYC-R+/Myclow ABC-DLBCL, we did not find significant differentially expressed genes below false discovery rate thresholds of 0.05–0.50). MYC-R+ ABC-DLBCL appears to have decreased B-cell receptor signaling compared with MYC-R+ GCB-DLBCL (false discovery rate <0.30; Table 4 and Supplementary Table S4C). However, the ‘loss’ of the gene expression profiling signature may also be due to the small case number.
Comparing between GCB and ABC subtypes of Mychigh patients, the overall GCB and ABC subtypes of Mychigh patients showed no significant difference in survival (ABC subtype showed nonsignificant trends towards poorer survival; Supplementary Figures S3E and F). However, MYC-R+/Mychigh GCB-DLBCL showed unfavorable trends compared with Mychigh ABC-DLBCL (either MYC-R− or MYC-R+). The P-value for the difference in PFS between the MYC-R+/Mychigh GCB-DLBCL and MYC-R−/Mychigh ABC-DLBCL patients was 0.058 (Figure 5g). MYC-R+/Mychigh GCB-DLBCL compared with MYC-R−/Mychigh ABC-DLBCL or Myclow ABC-DLBCL showed distinct gene expression profiling signatures overlapping with the one comparing with MYC-R− GCB-DLBCL. Comparisons between MYC-R+ DLBCL and MYC-R− DLBCL overall (regardless of Mychigh or Myclow, GCB or ABC), and between MYC-R+/Mychigh DLBCL overall and MYC-R−/Mychigh DLBCL overall (regardless of GCB or ABC) are shown in Figures 5h and i (Table 5 and Supplementary Table S4D). These analyses suggest the distinctive biology of MYC-R+/Mychigh GCB-DLBCL and cell of origin during lymphomagenesis may have a role in defining its biological feature.
Concurrent Evaluation of Myc/Bcl-2 Overexpression and MYC Translocations
We examined whether the prognostic value of Myc overexpression is contributed by or depends on the molecular marker associations with Mychigh DLBCL as shown in Figure 3. As shown previously, the prognostic significance of Mychigh and Bcl-2high in DLBCL significantly depend on each other.13, 20, 23, 24, 25, 26 In addition, to a certain extent, the prognostic significance of Mychigh showed dependence on CXCR4,40 FOXP1, and MUM1 overexpression, which are also associated with Bcl-2 overexpression (the association of MUM1 was only in ABC but not in GCB), high Ki-67 (for OS but not for PFS), and low BLIMP-1 expression (Supplementary Figure S5).
The lack of prognostic significance of Myc overexpression without MYC translocation in GCB-DLBCL (Figure 5a) could be attributable to the favorable prognosis of Mychigh/Bcl-2low GCB-DLBCL patients (Figures 6a and b and Supplementary Figures S6A and B). In contrast, in the ABC-DLBCL group, most MYC-R−/Mychigh patients (64%) also had Bcl-2 overexpression, which contributed to this group’s worse survival. Concurrent Bcl-2 overexpression also had significant prognostic impact in MYC-R+/Mychigh GCB-DLBCL (Figures 6c and d), but not in overall MYC-R+ ABC-DLBCL, MYC-R+/Mychigh ABC-DLBCL, or MYC-R+/Myclow cases (Supplementary Figures S6C and D); however, we could not distinguish whether the synergy in GCB is with Bcl-2 protein or with BCL2 translocation. Compared with Myclow, MYC-R−/Mychigh GCB-DLBCL, or overall MYC-R− GCB-DLBCL, MYC-R+/Mychigh GCB-DLBCL showed nonsignificant trends toward higher Bcl-2 levels (P=0.34, 0.27, and P=0.17, respectively; figures not shown; Supplementary Tables S1 and 2). The gene expression profiling analysis revealed no significant differentially expressed genes between MYC-R+ GCB-DLBCL with Bcl-2 expression and MYC-R+ GCB-DLBCL without Bcl-2 expression. Correspondingly, patients with MYC/BCL2 double-hit lymphoma and those with single MYC rearrangements did not have significant differentially expressed genes, which is consistent with an earlier study.16 The survival of MYC-R+ Mychigh/Bcl-2high GCB-DLBCL patients was markedly poorer compared with that of MYC-R− Mychigh/Bcl-2high GCB-DLBCL patients but this difference was not significant (Figures 6e and f). However, this difference was significant when we used cutoffs of ≥40% or >50% for Myc overexpression.
Prognostic analysis in Mychigh germinal-center-B-cell-like (GCB) diffuse large B-cell lymphoma (DLBCL) and in MychighBcl-2high DLBCL patients. (a and b) Among MYC-non-rearranged (MYC-R−) Mychigh GCB-DLBCL patients, those with Bcl-2 overexpression had significantly worse overall survival and progression-free survival compared with those who did not have Bcl-2 overexpression. (c and d) Among patients with MYC-R+/Mychigh GCB-DLBCL, those with Bcl-2 overexpression had significantly poorer progression-free survival compared with those who did not have Bcl-2 overexpression. The P-value for overall survival was not significant. (e and f) Among patients with MychighBcl-2high GCB-DLBCL, those with MYC rearrangement had poorer overall and progression-free survival compared with those without MYC rearrangement did, but these differences were not statistically significant. (g and h) Among MychighBcl-2high DLBCL patients, GCB-DLBCL patients with MYC rearrangement had significantly poorer overall and progression-free survival than MychighBcl-2high patients without MYC rearrangement did. ABC, activated-B-cell-like; Bcl-2high, high Bcl-2 protein expression; Bcl-2low, low Bcl-2 protein expression; GCB, germinal-center-B-cell-like; Mychigh, high Myc protein expression; MYC-R+, MYC rearrangement-positive; MYC-R−, MYC rearrangement-negative.
Although concurrent evaluation of Myc and Bcl-2 expression improves the specificity of Myc biomarker in DLBCL, MYC rearrangement continues to demonstrate clinical value. In our cohort, there was no significant difference in survival between patients with the GCB or ABC subtypes of MYC-R− Mychigh/Bcl-2high (ie, double-positive lymphoma). Among overall double-positive lymphoma patients, patients with MYC-R+ GCB double-positive lymphoma patients had significantly poorer survival compared with those double-positive lymphoma patients without MYC rearrangement (Figures 6g and h). The survival of MYC-R+ double-positive lymphoma patients with the GCB or ABC subtypes appeared to be different, although P-values were not significant and the case numbers were small.
Discussion
Previously, we reported MYC translocation and Myc overexpression as adverse prognostic biomarkers individually.15, 24 In this study, we analyzed the occurrence of MYC translocation and Myc overexpression in GCB- and ABC-DLBCL (Figures 2a and b), and compared with the clinical features and tumor biology associated with these two overlapping biomarkers which have not been done by previous studies, and examined the dependence/independence between their indicated prognoses. To reduce the difference in Myc expression levels as a causing factor for the differential prognostic effect and tumor biology between MYC translocation and Myc overexpression activated by other mechanisms, the cutoff for Mychigh was set at ≥70% in this study, which is the optimal cutoff for predicting MYC translocation according to previous studies.6, 7, 31 Using this cutoff, the frequency of Myc overexpression was 32.7% in overall DLBCL patients (close to the frequencies by other independent studies13, 22, 23), 73% in MYC-R+ GCB-DLBCL, and 54% in MYC-R+ ABC-DLBCL (lower than the 93% in Green et al6 and 85% in Horn et al10). Tables 6–7 summarize the results of published MYC studies including ours. The current study shows that MYC translocation and Myc overexpression in DLBCL only partially overlaps and evaluation of both is critical for stratifying patients and predicting treatment outcomes. MYC-rearranged DLBCL without Myc protein overexpression did not show significantly worse survival (Figure 1g). However, evaluation of Myc overexpression alone is also insufficient for predicting poorer prognosis in MYC-R− cases, especially in MYC-R− GCB-DLBCL (Figure 5a). This low specificity for Mychigh as an adverse prognostic factor can be improved by concurrent evaluation of Bcl-2 expression (Supplementary Figures S5 and S6), which is overexpressed mainly in ABC-DLBCL and associated with Myc overexpression. However, the survival of MYC-rearranged MychighBcl-2high GCB double-positive lymphoma patients remains significantly worse than other double-positive lymphoma patients (Figures 6g and h).
The biological investigation (in this regard, ≥70% is a better cutoff compared with ≥40% for Mychigh in our cohort) revealed that MYC activation was associated with significantly increased or decreased expression of genes and proteins involved in cell proliferation (e.g., pAKT, Ki-67), apoptosis (p53, Bcl-2, FAS, BCL2A1, PEG10, HINT1, TRAF1), differentiation (PRDM1, BLIMP-1, BACH2 (which represses PRDM1)43), noncoding RNAs (eg, LINC00152, GAS5, SNHG1, NAPSB) and microRNAs, microenvironment, and immune responses, as well as cell-of-origin markers (Figure 3 and Tables 3, 4, 5). Corresponding to the differences in prognostic effect between various Mychigh subtypes, only MYC-R+/Mychigh GCB-DLBCL and MYC-R−/Mychigh ABC-DLBCL, but not MYC-R−/Mychigh GCB-DLBCL or MYC-R+/Mychigh ABC-DLBCL, demonstrated distinct gene expression profiling feature compared with the Myclow subgroup (Figures 5b and e and Supplementary Figures S4D and F). Remarkably, MYC-R+/Mychigh GCB-DLBCL had a characteristic gene expression profiling in DLBCL. Myc activation was associated with gene expression profiling signatures suggesting decreased immune responses and a number of microRNAs overlapped with the molecular Burkitt lymphoma signature,29, 30 including MIR17HG, which was markedly upregulated, and MIR21 (ref. 44) and MIR155HG, which were significantly downregulated. Other studies have shown these microRNAs to be regulated by the Myc, Bcl-6, STAT3, and NF-κB pathways, and the MIR17HG locus was frequently amplified in Burkitt lymphoma.45, 46, 47, 48 Downregulation of MIR155HG expression may contribute to the pathogenesis of MYC translocation, as miR-155 suppresses activation-induced cytidine deaminase, which mediates MYC/IGH translocation.49 These microRNA signatures may also be implicated in defining the gene expression profiling features of MYC-R+/Mychigh GCB-DLBCL. In lymphoma cells, ectopic expression of miR-155 is associated with downregulation of IGJ, FAS, SMAD3/5, and BACH1, as well as HLA genes.45 IGJ, FAS, and genes involved in BMP/SMAD pathways (such as SMAD1, BMP3, and BMP7), as well as SMARCA4 (miR-21 target gene50), were upregulated in MYC-R+ GCB-DLBCL (Tables 3 and 5). miR-155 target gene BCL2 showed increased Bcl-2 protein expression in ABC-DLBCL (Figure 3c). Phosphorylated SMAD proteins have roles in BMP-induced cell growth inhibition (this inhibition can be overcame via MYC translocations),51, 52 immunoregulation,53 and ionizing radiation-induced double-strand break signaling.54 In addition, in MYC-R+/Mychigh GCB-DLBCL, antiapoptotic PEG10 and ZNF385B, which modulates p53 activity resulting in cell-cycle arrest over apoptosis, were significantly upregulated.
One possible reason for the differential prognoses and gene expression profiling signatures among GCB or ABC subtype of Mychigh and MYC-R+ cases is the difference in Myc protein levels. For example, MYC-R+/Mychigh GCB-DLBCL had highest Myc expression levels (Figure 2c) and significant or nonsignificant trends towards worse prognosis compared with all other three Mychigh groups. MYC-R+ cases with low Myc protein expression had good prognosis even though having an MYC rearrangement. Different MYC translocation partners, and/or breakpoints outside of MYC gene leaving MYC repressor element intact during rearrangement,4, 17 may cause the low MYC-mRNAs in these MYC-R+/Myclow DLBCL cases (Figures 1e–g). Compared with MYC-R+/Mychigh GCB-DLBCL, ABC subtype of MYC-R+/Mychigh DLBCL had similar MYC-mRNA but significantly lower Myc protein levels (Figure 2d) and trends of better survival. MYC-R−/Mychigh GCB-DLBCL compared with Myclow GCB-DLBCL had significantly higher levels of MYC-mRNA and Myc protein, but the false discovery rate for MYC upregulation in gene expression profiling analysis was high (>0.45), which may suggest either Mychigh or Myclow group are heterogeneous and the molecular mechanisms inducing Myc in this Mychigh group include post-transcriptional regulation. A previous Myc study in immature and mature GCB cells during germinal cell formation also demonstrated the lack of correlation between Myc protein and mRNA levels.55 The lack of distinct gene expression profiling signatures and better survival of MYC-R−/Mychigh GCB-DLBCL and MYC-R+/Mychigh ABC-DLBCL compared with MYC-R+/Mychigh GCB-DLBCL may indicate lower Myc activities corresponding to intracellular Myc protein levels; however, small case numbers and/or heterogeneity among these two Mychigh DLBCL groups could also be possible causes.
However, the Myc immunohistochemistry levels were similar between MYC-R−/Mychigh GCB-DLBCL and MYC-R−/Mychigh ABC-DLBCL (although GCB type had slightly lower level of MYC-mRNA) but their prognosis showed differences. MYC-R+/Mychigh ABC-DLBCL had similar Myc protein level to that of MYC-R−/Mychigh ABC-DLBCL but only the latter showed typical Myc gene expression profiling signatures (Figure 2c). Therefore, Myc-associated molecular mechanisms in GCB or ABC subtype of MYC-R+/− Mychigh cases impacted the prognostic and biological effect of Myc. GCB and ABC subtypes of Mychigh DLBCL had difference in frequencies of TP53 mutation, MYC translocation, BCL2 translocation, and Bcl-2 expression, as well as cell-of-origin biomarkers (Table 1 and Figure 3). MYC-R+/Mychigh and MYC-R−/Mychigh DLBCL had significantly different levels of MDM2, the cell-of-origin markers, and CD30 (in GCB-DLBCL) and pAKT (in ABC-DCLBL) (Figure 4), as well as significantly different gene expression profiling signatures at the mRNA level (Figure 5i). Recently, studies have posited a ‘c-Myc function rule,’ in which c-Myc is a ‘universal amplifier’ of active (expressed) genes in lymphocytes.33, 34, 57
In summary, both MYC rearrangement and Myc overexpression have advantage and limitations as a single biomarker in DLBCL, and their prognostic importance is significantly different in GCB- vs ABC-DLBCL. GCB subtype (opposite to the general association of ABC-DLBCL with poorer survival) of MYC-R+ DLBCL with Myc overexpression was associated with significantly poorer survival, likely contributed by significantly higher Myc protein levels as well as associated tumor biology. In ABC-DLBCL, Myc overexpression associated with Bcl-2 overexpression was a significantly adverse biomarker independent of MYC translocation. Our results suggest that fluorescence in situ hybridization analysis for MYC rearrangements and immunohistochemistry evaluation for Myc and Bcl-2 expression are both needed to determine the prognosis in subsets of patients.57 Insights gained into the tumor biology associated with MYC abnormalities are important for understanding the functional role of MYC in lymphomagenesis and chemoresistance, and help identify oncogenic targets for therapeutic intervention (Table 7).
References
Felsher DW . MYC inactivation elicits oncogene addiction through both tumor cell-intrinsic and host-dependent mechanisms. Genes Cancer 2010;1:597–604.
Delmore JE, Issa GC, Lemieux ME et al. BET bromodomain inhibition as a therapeutic strategy to target c-Myc. Cell 2011;146:904–917.
Chesi M, Matthews GM, Garbitt VM et al. Drug response in a genetically engineered mouse model of multiple myeloma is predictive of clinical efficacy. Blood 2012;120:376–385.
Ott G, Rosenwald A, Campo E . Understanding MYC-driven aggressive B-cell lymphomas: pathogenesis and classification. Blood 2013;122:3884–3891.
Kluk MJ, Chapuy B, Sinha P et al. Immunohistochemical detection of MYC-driven diffuse large B-cell lymphomas. PLoS One 2012;7:e33813.
Green TM, Nielsen O, de Stricker K et al. High levels of nuclear MYC protein predict the presence of MYC rearrangement in diffuse large B-cell lymphoma. Am J Surg Pathol 2012;36:612–619.
Tapia G, Lopez R, Munoz-Marmol AM et al. Immunohistochemical detection of MYC protein correlates with MYC gene status in aggressive B cell lymphomas. Histopathology 2011;59:672–678.
Wang XJ, Medeiros LJ, Lin P et al. MYC cytogenetic status correlates with expression and has prognostic significance in patients with MYC/BCL2 protein double-positive diffuse large B-cell lymphoma. Am J Surg Pathol 2015;39:1250–1258.
Cheah CY, Oki Y, Westin JR et al. A clinician's guide to double hit lymphomas. Br J Haematol 2015;168:784–795.
Horn H, Staiger AM, Vohringer M et al. Diffuse large B-cell lymphomas of immunoblastic type are a major reservoir for MYC-IGH translocations. Am J Surg Pathol 2015;39:61–66.
Barrans S, Crouch S, Smith A et al. Rearrangement of MYC is associated with poor prognosis in patients with diffuse large B-cell lymphoma treated in the era of rituximab. J Clin Oncol 2010;28:3360–3365.
Savage KJ, Johnson NA, Ben-Neriah S et al. MYC gene rearrangements are associated with a poor prognosis in diffuse large B-cell lymphoma patients treated with R-CHOP chemotherapy. Blood 2009;114:3533–3537.
Horn H, Ziepert M, Becher C et al. MYC status in concert with BCL2 and BCL6 expression predicts outcome in diffuse large B-cell lymphoma. Blood 2013;121:2253–2263.
Obermann EC, Csato M, Dirnhofer S et al. Aberrations of the MYC gene in unselected cases of diffuse large B-cell lymphoma are rare and unpredictable by morphological or immunohistochemical assessment. J Clin Pathol 2009;62:754–756.
Tzankov A, Xu-Monette ZY, Gerhard M et al. Rearrangements of MYC gene facilitate risk stratification in diffuse large B-cell lymphoma patients treated with rituximab-CHOP. Mod Pathol 2014;27:958–971.
Aukema SM, Kreuz M, Kohler CW et al. Biological characterization of adult MYC-translocation-positive mature B-cell lymphomas other than molecular Burkitt lymphoma. Haematologica 2014;99:726–735.
Bertrand P, Bastard C, Maingonnat C et al. Mapping of MYC breakpoints in 8q24 rearrangements involving non-immunoglobulin partners in B-cell lymphomas. Leukemia 2007;21:515–523.
van Imhoff GW, Boerma EJ, van der Holt B et al. Prognostic impact of germinal center-associated proteins and chromosomal breakpoints in poor-risk diffuse large B-cell lymphoma. J Clin Oncol 2006;24:4135–4142.
Tibiletti MG, Martin V, Bernasconi B et al. BCL2, BCL6, MYC, MALT 1, and BCL10 rearrangements in nodal diffuse large B-cell lymphomas: a multicenter evaluation of a new set of fluorescent in situ hybridization probes and correlation with clinical outcome. Hum Pathol 2009;40:645–652.
Johnson NA, Savage KJ, Ludkovski O et al. Lymphomas with concurrent BCL2 and MYC translocations: the critical factors associated with survival. Blood 2009;114:2273–2279.
Visco C, Tzankov A, Xu-Monette ZY et al. Patients with diffuse large B-cell lymphoma of germinal center origin with BCL2 translocations have poor outcome, irrespective of MYC status: a report from an International DLBCL rituximab-CHOP Consortium Program Study. Haematologica 2013;98:255–263.
Johnson NA, Slack GW, Savage KJ et al. Concurrent expression of MYC and BCL2 in diffuse large B-cell lymphoma treated with rituximab plus cyclophosphamide, doxorubicin, vincristine, and prednisone. J Clin Oncol 2012;30:3452–3459.
Perry AM, Alvarado-Bernal Y, Laurini JA et al. MYC and BCL2 protein expression predicts survival in patients with diffuse large B-cell lymphoma treated with rituximab. Br J Haematol 2014;165:382–391.
Hu S, Xu-Monette ZY, Tzankov A et al. MYC/BCL2 protein coexpression contributes to the inferior survival of activated B-cell subtype of diffuse large B-cell lymphoma and demonstrates high-risk gene expression signatures: a report from The International DLBCL Rituximab-CHOP Consortium Program. Blood 2013;121:4021–4031.
Green TM, Young KH, Visco C et al. Immunohistochemical double-hit score is a strong predictor of outcome in patients with diffuse large B-cell lymphoma treated with rituximab plus cyclophosphamide, doxorubicin, vincristine, and prednisone. J Clin Oncol 2012;30:3460–3467.
Valera A, Lopez-Guillermo A, Cardesa-Salzmann T et al. MYC protein expression and genetic alterations have prognostic impact in patients with diffuse large B-cell lymphoma treated with immunochemotherapy. Haematologica 2013;98:1554–1562.
Horn H, Ziepert M, Wartenberg M et al. Different biological risk factors in young poor-prognosis and elderly patients with diffuse large B-cell lymphoma. Leukemia 2015;29:1564–1570.
Pfreundschuh M . Growing importance of MYC/BCL2 immunohistochemistry in diffuse large B-cell lymphomas. J Clin Oncol 2012;30:3433–3435.
Hummel M, Bentink S, Berger H et al. A biologic definition of Burkitt's lymphoma from transcriptional and genomic profiling. N Engl J Med 2006;354:2419–2430.
Dave SS, Fu K, Wright GW et al. Molecular diagnosis of Burkitt's lymphoma. N Engl J Med 2006;354:2431–2442.
Thieblemont C, Briere J . MYC, BCL2, BCL6 in DLBCL: impact for clinics in the future? Blood 2013;121:2165–2166.
Albihn A, Johnsen JI, Henriksson MA . MYC in oncogenesis and as a target for cancer therapies. Adv Cancer Res 2010;107:163–224.
Lin CY, Loven J, Rahl PB et al. Transcriptional amplification in tumor cells with elevated c-Myc. Cell 2012;151:56–67.
Nie Z, Hu G, Wei G et al. c-Myc is a universal amplifier of expressed genes in lymphocytes and embryonic stem cells. Cell 2012;151:68–79.
Visco C, Li Y, Xu-Monette ZY et al. Comprehensive gene expression profiling and immunohistochemical studies support application of immunophenotypic algorithm for molecular subtype classification in diffuse large B-cell lymphoma: a report from the International DLBCL Rituximab-CHOP Consortium Program Study. Leukemia 2012;26:2103–2113.
Xu-Monette ZY, Wu L, Visco C et al. Mutational profile and prognostic significance of TP53 in diffuse large B-cell lymphoma patients treated with R-CHOP: report from an International DLBCL Rituximab-CHOP Consortium Program Study. Blood 2012;120:3986–3996.
Xu-Monette ZY, Moller MB, Tzankov A et al. MDM2 phenotypic and genotypic profiling, respective to TP53 genetic status, in diffuse large B-cell lymphoma patients treated with rituximab-CHOP immunochemotherapy: a report from the International DLBCL Rituximab-CHOP Consortium Program. Blood 2013;122:2630–2640.
Hu S, Xu-Monette ZY, Balasubramanyam A et al. CD30 expression defines a novel subgroup of diffuse large B-cell lymphoma with favorable prognosis and distinct gene expression signature: a report from the International DLBCL Rituximab-CHOP Consortium Program Study. Blood 2013;121:2715–2724.
Ok CY, Xu-Monette ZY, Li L et al. Evaluation of NF-kappaB subunit expression and signaling pathway activation demonstrates that p52 expression confers better outcome in germinal center B-cell-like diffuse large B-cell lymphoma in association with CD30 and BCL2 functions. Mod Pathol 2015;28:1202–1213.
Chen J, Xu-Monette ZY, Deng L et al. Dysregulated CXCR4 expression promotes lymphoma cell survival and independently predicts disease progression in germinal center B-cell-like diffuse large B-cell lymphoma. Oncotarget 2015;6:5597–5614.
Li L, Xu-Monette ZY, Ok CY et al. Prognostic Impact of c-Rel nuclear expression and REL amplification and crosstalk between c-Rel and the p53 pathway in diffuse large B-cell lymphoma. Oncotarget; advance online publication, 30 June 2015 [e-pub ahead of print].
Liu Z, Xu-Monette ZY, Cao X et al. Prognostic and biological significance of survivin expression in patients with diffuse large B-cell lymphoma treated with rituximab-CHOP therapy. Mod Pathol; advance online publication, 7 August 2015 [e-pub ahead of print].
Huang C, Geng H, Boss I et al. Cooperative transcriptional repression by BCL6 and BACH2 in germinal center B-cell differentiation. Blood 2014;123:1012–1020.
Lu TX, Hartner J, Lim EJ et al. MicroRNA-21 limits in vivo immune response-mediated activation of the IL-12/IFN-gamma pathway, Th1 polarization, and the severity of delayed-type hypersensitivity. J Immunol 2011;187:3362–3373.
Iqbal J, Shen Y, Huang X et al. Global microRNA expression profiling uncovers molecular markers for classification and prognosis in aggressive B-cell lymphoma. Blood 2015;125:1137–1145.
Di Lisio L, Sanchez-Beato M, Gomez-Lopez G et al. MicroRNA signatures in B-cell lymphomas. Blood Cancer J 2012;2:e57.
Basso K, Schneider C, Shen Q et al. BCL6 positively regulates AID and germinal center gene expression via repression of miR-155. J Exp Med 2012;209:2455–2465.
Testoni M, Zucca E, Young KH et al. Genetic lesions in diffuse large B-cell lymphomas. Ann Oncol 2015;1069–1080.
Dorsett Y, McBride KM, Jankovic M et al. MicroRNA-155 suppresses activation-induced cytidine deaminase-mediated Myc-Igh translocation. Immunity 2008;28:630–638.
Schramedei K, Morbt N, Pfeifer G et al. MicroRNA-21 targets tumor suppressor genes ANP32A and SMARCA4. Oncogene 2011;30:2975–2985.
Holien T, Vatsveen TK, Hella H et al. Bone morphogenetic proteins induce apoptosis in multiple myeloma cells by Smad-dependent repression of MYC. Leukemia 2012;26:1073–1080.
Huse K, Bakkebo M, Walchli S et al. Role of Smad proteins in resistance to BMP-induced growth inhibition in B-cell lymphoma. PLoS One 2012;7:e46117.
Malhotra N, Kang J . SMAD regulatory networks construct a balanced immune system. Immunology 2013;139:1–10.
Wang M, Saha J, Hada M et al. Novel Smad proteins localize to IR-induced double-strand breaks: interplay between TGFbeta and ATM pathways. Nucleic Acids Res 2013;41:933–942.
Calado DP, Sasaki Y, Godinho SA et al. The cell-cycle regulator c-Myc is essential for the formation and maintenance of germinal centers. Nat Immunol 2012;13:1092–1100.
Akyurek N, Uner A, Benekli M et al. Prognostic significance of MYC, BCL2, and BCL6 rearrangements in patients with diffuse large B-cell lymphoma treated with cyclophosphamide, doxorubicin, vincristine, and prednisone plus rituximab. Cancer 2012;118:4173–4183.
Cai Q, Medeiros LJ, Young KH . MYC-driven aggressive B-cell lymphomas: biology, entity, differential diagnosis and clinical management. Oncotarget 2015 [e-pub ahead of print].
Acknowledgements
This study was supported by the National Cancer Institute/National Institutes of Health (R01CA138688, R01CA187415 and 1RC1CA146299 to YL and KHY). ZYXM is the recipient of the Harold C and Mary L Daily Endowment Fellowships and Shannon Timmins Fellowship for Leukemia Research Award. GM is supported by a grant from the Michael and Susan Dell Foundation. KHY is supported by The University of Texas MD Anderson Cancer Center Lymphoma Moonshot Program, Institutional Research and Development Fund, an Institutional Research Grant Award, an MD Anderson Cancer Center Lymphoma Specialized Programs on Research Excellence (SPORE) Research Development Program Award, an MD Anderson Cancer Center Myeloma SPORE Research Development Program Award, a Gundersen Lutheran Medical Foundation Award, and partially supported by the National Cancer Institute/National Institutes of Health (P50CA136411 and P50CA142509), and by MD Anderson’s Cancer Center Support Grant CA016672.
Author contributions
Conception and design: ZYX-M, KHY; research performance: ZYX-M, KHY; provision of study materials, key reagents and technology: ZYX-M, BSD, XW, MT, GCM, AT, YX, LZ, CV, KD, LY, AC, AO, YZ, GB, KLR, EDH, WWLC, JHvK, MP, AJMF, MBM, BMP, XZ, JNW, MAP, TJM, RNM, YL, LJM, KHY; collection and assembly of data under approved IRB and MTA: ZYX-M, BSD, XW, AT, YX, CV, KD, AC, AO, YZ, GB, KLR, EDH, WWLC, JHvK, MP, AJMF, MBM, BMP, XZ, JNW, MAP, KHY; data analysis and interpretation: ZYX-M, KHY; manuscript writing: ZYX-M, LJM, KHY; final approval of manuscrip: all authors.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
KHY also receives research support from Roche Molecular System, Gilead Sciences Pharmaceutical, Seattle Genetics, Dai Sanyo Pharmaceutical, Adaptive Biotechnology, and HTG Molecular Diagnostics.
Additional information
Supplementary Information accompanies the paper on Modern Pathology website
Supplementary information
Rights and permissions
About this article
Cite this article
Xu-Monette, Z., Dabaja, B., Wang, X. et al. Clinical features, tumor biology, and prognosis associated with MYC rearrangement and Myc overexpression in diffuse large B-cell lymphoma patients treated with rituximab-CHOP. Mod Pathol 28, 1555–1573 (2015). https://doi.org/10.1038/modpathol.2015.118
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2015.118
This article is cited by
-
A practical approach to FISH testing for MYC rearrangements and brief review of MYC in aggressive B-cell lymphomas
Journal of Hematopathology (2020)
-
Mucosal intralymphatic spread in a relapsed diffuse large B cell lymphoma
Journal of Hematopathology (2020)
-
PD-1/PD-L1 expression and interaction by automated quantitative immunofluorescent analysis show adverse prognostic impact in patients with diffuse large B-cell lymphoma having T-cell infiltration: a study from the International DLBCL Consortium Program
Modern Pathology (2019)
-
Prognostic significances of overexpression MYC and/or BCL2 in R-CHOP-treated diffuse large B-cell lymphoma: A Systematic review and meta-analysis
Scientific Reports (2018)
-
NF-κB p50 activation associated with immune dysregulation confers poorer survival for diffuse large B-cell lymphoma patients with wild-type p53
Modern Pathology (2017)