Abstract
Long non-coding RNAs (lncRNAs) are a class of newly identified non-coding RNA molecules that are emerging as key regulators of tumor initiation and development. Colorectal cancer (CRC) remains a major health problem worldwide, and there remains a need to further refine the current screening approaches as well as provide tailored diagnostic and therapeutic approaches. Multiple dysregulated lncRNAs participate in tumorigenesis through a variety of molecular mechanisms, and various regulatory factors frequently contribute to the aberrant expression of lncRNAs in CRC, thereby allowing malignant transformation. Additionally, the association of dysregulated lncRNAs with specific developmental stages and clinical outcomes indicates their potential as strong diagnostic and prognostic predictors as well as therapeutic targets. Here we provide a brief overview of the known functions of CRC-associated lncRNAs, describe some potential molecular mechanisms that underlie changes in lncRNA expression in CRC, and attempt to uncover their clinical and therapeutic potential.
Similar content being viewed by others
Main
High-throughput genome-scale studies have demonstrated that more than 93% of the DNA sequences in the human genome are actively transcribed.1 However, only approximately 5–10% of the sequences are stably transcribed into mRNA or non-coding RNA (ncRNA). Genome tiling arrays have revealed that the amount of non-coding sequence is at least four times larger than the amount of coding sequence, which indicates that only 1% of the human genome is composed of protein-coding genes and the remaining 4–9% is transcribed into ncRNAs.2 Therefore, ncRNAs constitute a very large proportion of the total RNA molecules.
The function and clinical significance of short regulatory ncRNAs, such as microRNAs (miRNAs) and small interfering RNAs (siRNAs), were elucidated first,3 and the regulatory roles of miRNAs have been broadly recognized in almost all physiological and pathological processes in the body, including carcinogenesis.4 For example, we previously reported that MIR95 promotes cell proliferation and targets sorting nexin 1 in human colorectal carcinoma;5 moreover, in colorectal cancer (CRC) patients, the plasma levels of MIR29a and MIR92a are significantly upregulated and the plasma levels of MIR601 and MIR760 are significantly downregulated; thus the levels of these miRNAs have good diagnostic value for CRC screening.6, 7
According to their transcript size, ncRNAs are grouped into two major classes: (i) small ncRNAs with transcripts <200 nucleotides (nt; eg, aforementioned siRNAs and miRNAs, Piwi-interacting RNAs, and some retrotransposon-derived RNAs) and (ii) long non-coding RNAs (lncRNAs), this class includes five broad categories: sense, antisense, bidirectional, intronic, and intergenic, based on the proximity between neighboring transcripts.8 For a time, lncRNAs was commonly defined as a protein-coding transcripts that is >200 nt.9 However, this definition is arbitrary and limits in distinguishing lncRNAs from small regulatory ncRNAs, without taking its distinct structure or function characteristics into account. For example, lncRNAs bear many signatures of mRNAs, including 5′ capping, frequently transcribed by RNA polymerase II, poly-adenylation and splicing, but have little or no open reading frame (ORF).10 And moreover, a significant proportion of lncRNAs may possess coding and non-coding activities.11 Combining with these characters, Mercer et al2 proposed a definition that describes lncRNAs as ‘RNA molecules that may function as either primary or spliced transcripts and do not fit into known classes of small RNAs or into classes of structural RNAs’. This undated definition overcomes the restriction of ORF and length that was arbitrarily set. However, so far there still a lack of standard nomenclature that makes comprehensive comments on the general features of lncRNAs, which to some extent demonstrate how little we currently know about this newly discovered class of ncRNA transcripts.
According to the LNCipedia database (www.lncipedia.org), 32 183 human lncRNAs have been annotated.12 Emerging studies have revealed that particular lncRNAs are involved in diverse physiological and pathological processes, such as cell growth, apoptosis, stem cell pluripotency, and development, by acting as transcriptional, post-transcriptional, or epigenetic regulators. These studies have revealed the functional potential of these molecules;9, 10, 13, 14 however, the majority of lncRNAs have not yet been functionally characterized.Notably, observations of a few known lncRNAs have suggested that their dysregulation is linked to tumor pathogenesis, and these molecules perform essential regulatory functions by acting on cellular proliferation, apoptosis, or metastasis by participating in a variety of key signaling pathways.11, 15, 16, 17
Here, we summarize the recent research concerning alterations in and roles of lncRNAs in cancer pathogenesis, with a specific focus on CRC. Additionally, we discuss the significant and almost wholly untapped potential of lncRNAs as biomarkers for the early diagnosis of CRC, as indicators of CRC prognosis prediction, or even as targets in CRC treatment strategies.
LncRNAs in cancer
Cancer is the result of a wide array of cellular transformation processes that occur due to oncogene activation and tumor-suppressor gene defects and lead to uncontrolled cell proliferation and escape from apoptosis. Using a range of techniques, such as microarray,18 RNA sequencing,19 and real-time PCR,20 many studies have confirmed that lncRNA expression levels differ between normal and tumor tissues and vary among tumor types.21, 22, 23 Although some of these abnormal expression patterns may be secondary effects from the cancer transformation, several factors suggest that lncRNAs could influence tumorigenesis. These factors include the tremendous complexity and diversity of the lncRNA landscape, the finding that lncRNAs are often located at crucial sites (eg, regions of single-nucleotide polymorphisms (SNPs), amplifications, or at common breakpoints), the presence of sequence motifs and other elements that result in specific structures, their regulation, and their functional relationships with other nucleic acids and proteins.11, 16 In fact, previous studies have shown that particular deregulated lncRNAs are important regulators of tumor formation and progression.17, 24 For example, the antisense intergenic RNA HOX Antisense Intergenic RNA (HOTAIR) is highly expressed in multiple types of primary somatic tumors, such as hepatocellular carcinoma,25, 26 pancreatic cancer,27 and gastric cancer,28 as well as some metastatic tumors, including metastatic breast tumors29 and melanoma.30 Moreover, high HOTAIR level is associated with poor patient survival rate20, 27 and tumor recurrence.25 It was discovered that HOTAIR represses transcription across 40 kb of the HOXD locus by acting as a scaffold of histone modification complexes; it binds with polycomb chromatin remodeling complexes in trans, which lead to alterations of cells’ epigenetic state and subsequent gene expression.31 Moreover, HOTAIR can also interact with the LSD1/CoREST/REST complex, which coordinates targeting of LSD1 to chromatin for coupled histone K4 demethylation.32 Given its important role in the epigenetic regulation of gene expression by mediating the modulation of chromatin structures, it is not surprising that HOTAIR performs vital functions in increasing cancer invasiveness and metastasis25, 26, 29 and shows immense clinical relevance. Particularly, lncRNA expression levels may be extremely useful for monitoring tumors, and we consider lncRNAs to be novel cancer diagnostic tools and a source of future therapies.33
The involvement of lncRNAs in CRC
CRC remains a major health problem and represents the third most common cause of cancer-related death worldwide.34 Most cases of CRC usually progress from benign polyps to malignant adenocarcinomas and distant metastases; therefore, screening and early diagnosis, as well as early treatment, help to extend the long-term survival of patients.35 Given that the initiation, progression, and metastasis of CRC involve multiple genetic and epigenetic alterations, which permit the adaptations characteristic of malignant tumors, previous detection and treatment technologies based on genomic and proteomic analyses have got encouraging achievements to ease the disease burden.36, 37, 38, 39 However, only a limited number of CRC-associated gene and protein candidates have been clinically validated and demonstrated clinical utility. Additionally, the sensitivity and specificity of the current biomarkers used for the detection of precancerous lesions and early stages of tumor formation is weak, and their ability to predict the chemotherapeutic response is poor.36 Consequently, there is still a need for further research regarding the complex regulatory networks in CRC to refine the existing programs and to provide tailored diagnostic and therapeutic approaches.
Research conducted during the past 30 years has uncovered several critical important genes and pathways involved in the pathogenesis of many kinds of tumors including CRC, such as the WNT, RAS-MAPK, PI3K, TGF-β, P53, and DNA mismatch-repair pathways. More recently, with the aid of high-throughput techniques, a variety of systematic cancer genomics projects, such as The Cancer Genome Atlas Project (TCGA) (http://cancergenome.nih.gov/), are being used to investigate different molecular pathways and the genomic, transcriptomic, proteomic, and epigenomic alterations in each specific cancers, including CRC.40, 41, 42 This disease-specific focus has identified novel oncogenic drivers, and the genes contributing to functional change,43 importantly, revealed that different molecular features contribute to individual differences that occurred in clinicopathological characteristics, disease behavior, prognosis, and response to treatments, which thus helped to establish definitions of molecular subtypes and identified new biomarkers on the basis of omic alterations.44, 45, 46
It is now well known that some CRC cases are linked to some factors, such as environment,47 inflammation,48, 49 immunity,50 and epigenetic alterations51, 52 rather than heritable genetic changes. An interesting thing is that these factors can influence each other, for instance, epigenetic aberrations induced by environmental factors contribute to cancer processes;53 interaction of drug and molecular characteristics can influence lncRNAs and clinical outcome;46, 54, 55 and epigenetic factors such as lncRNAs can also coordinate cellular responses to environment in turn.56
The significance of lncRNAs in human CRC was realized in 2001 when Tanaka et al57 determined that a loss of imprinting of long QT intronic transcript 1 (LIT1/KCNQ1OT1) was frequently observed in CRC patients, suggesting a link between lncRNAs and CRC. Following this research, several studies focused on the aberrant expression of lncRNAs during colorectal carcinogenesis, and an accumulating number of studies indicated that specific lncRNAs had potential biological and clinical relevance in CRC (Table 1). According to these data, understanding the pathophysiological roles of lncRNAs in CRC undoubtedly represents an important aspect of current and future research, as these molecules may be the hallmark features of CRC. Furthermore, the detection and identification of potentially functional lncRNAs in CRC is an emerging avenue of lncRNA research, which will be necessary before the application of lncRNAs in cancer diagnosis and therapy (Figures 1 and 2).
Role of lncRNAs in Tumorigenesis and Progression
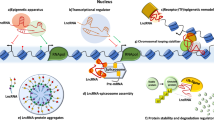
LncRNAs may be involved in carcinogenesis and the progression of tumors through a variety of mechanisms. Generally, some lncRNAs function as oncogenes or tumor suppressors by participating in pivotal signaling pathways, while others have roles in malignant transformation by interacting with other regulatory molecules, such as DNA, RNA, and proteins.17
The expression level of the lncRNA colon cancer-associated transcript 2 (CCAT2) in microsatellite-stable (MSS) CRC patients was higher than that in microsatellite instable-high (MSI-H) CRC patients, and cancer cells transduced with CCAT2-containing retrovirus exhibited numerical and structural chromosomal changes, illustrating that CCAT2 can contribute to the MSS phenotype by inducing chromosomal instability.58 Additionally, CCAT2 itself has a role in CRC pathogenesis through its direct involvement in the regulatory network. CCAT2 participates in loop formation between the genomic locus rs6983267 and the MYC promoter and works with the enhancer element to activate the transcription of the MYC oncogene. Moreover, CCAT2 enhances WNT activity by binding to TCF7L2, a pivotal transcription factor in the WNT signaling pathway, and facilitates MYC function, thereby enhancing cancer cell invasion and metastasis.58 Intriguingly, CCAT2 levels can then be reduced by WNT signaling under TCF7L2 knockdown conditions, suggesting that a feedback loop may exist between CCAT2 and WNT signaling.58 An attractive therapeutic strategy might involve attenuating CCAT2 activity and blocking the function of the downstream oncogenes in the WNT signaling pathway.
Data obtained from functional manipulations of lncRNAs have confirmed that lncRNAs have roles in tumor development by participating in a ‘competitive endogenous RNA’ network and by acting as endogenous miRNA ‘sponges’ or decoys.59, 60, 61, 62 Previous investigations revealed that an effector lncRNA downstream of TP53—LOC285194, also referred to as tumor-suppressor candidate 7—is significantly deregulated in CRC63 and can suppress tumor cell growth both in vitro and in vivo.64, 65 Exon 4 of LOC285194, which is responsible for tumor cell growth inhibition, contains an active region that harbors two MIR211 binding sites. Moreover, in vivo studies confirmed that LOC285194 can bind to and downregulate MIR211.64 Thus LOC285194 may modulate the cancer phenotype through its function as an endogenous miRNA sponge. Additionally, a reciprocal repression feedback loop exists between LOC285194 and MIR211, suggesting that miRNAs can also modulate lncRNAs.64 In this scenario, lncRNAs and miRNAs may interact as components of a new genomic regulatory network, thereby greatly expanding the functional genetic information in the human genome.
Regulation of lncRNA Expression
The expression of lncRNAs can be modulated by multiple regulatory factors, which naturally raises the question of how particular lncRNAs become deregulated in tumors. Although the underlying mechanism is generally unknown, it has been proposed that the expression of certain lncRNAs in CRC can be initiated or modulated by genetic (eg, SNPs in BA318C17.1,66 sequence mutation of MALAT-167), epigenetic (eg, MIR211-dependent regulation of LOC285194, histone deacetylation-induced hypoxia-mediated downregulation of lncRNA-LET/NPTN-IT168), and transcriptional (eg, TCF7L2-dependent regulation of CCAT2,58 TP53-dependent regulation of PVT169) regulatory factors and may allow for the exact execution of malignant transformation to some extent.58, 67, 68 Intriguingly, in addition to being affected by MIR211,64 LOC285194 was also found to be regulated by genetic deletion (copy number variation) and TP53,64, 65 suggesting that the expression of specific lncRNAs may be regulated by several regulatory mechanisms.
Most variations in regulatory sequences lead to relatively subtle phenotypic changes.70 SNPs are the most abundant DNA variations in the human genome and contribute to human phenotypic differences. The rs6983267 SNP region, located upstream of the MYC oncogene on chromosome 8q24, has been found to be consistently associated with an increased risk of CRC.71 The G allele of rs6983267, in addition to being related to a significantly increased risk of CRC,72 results in more CCAT2 transcripts than the T allele, which further affects the regulation of MYC by CCAT2; thus different rs6983267 alleles can affect CCAT2 expression and function.58 Studies examining other cancer types have shown that SNPs in the key regulatory regions of an lncRNA can alter its structural motifs73 and may consequently affect its expression and function in malignancies,74 additionally contributing to cancer risk.75, 76 SNPs in the cancer-associated lncRNAs could potentially have pivotal roles in colorectal tumorigenesis and progression. Nevertheless, the means by which these alleles affect the lncRNAs and ultimately participate in colon carcinogenesis remains to be investigated.
LncRNAs in CRC may also harbor structural variations. Calin and colleagues recently identified a class of lncRNAs termed non-coding transcribed ultraconserved regions (T-UCRs); in humans, this class of lncRNAs shares 100% sequence conservation with both mouse and rat. In CRC patients, these loci frequently contain DNA sequence variations, including somatic variations, germline variations, and SNPs, compared with the general population; four T-UCRs were shown to harbor sequence variations in 35 CRC patients, whereas none of these mutations existed in a cohort of 175 cancer-free control patients.77 Nevertheless, because no studies verified that these T-UCRs were deregulated or involved in CRC predisposition, it is difficult to determine whether the sequence mutations in these T-UCRs have specific impacts on colon carcinogenesis. Therefore, an examination of mutated lncRNAs in CRC will be important in future investigations.
Clinical applications of lncRNAs in CRC
Given the versatile biological and pathological roles played by lncRNAs, it is not surprising that these molecules are associated with cancer progression. Therefore, their expression levels may be an indicator of the intrinsic characteristics of the tumor. As molecules that enable epigenetic regulation, aberrant expression of lncRNAs occurs before the phenotypic changes observed during the progression to carcinoma. Additionally, due to their tissue specificity, lncRNAs could potentially be more sensitive for diagnosis and more specific prognostic predictors than the current DNA, protein-coding RNA, or protein biomarkers (Figure 2).
Tumor Diagnosis
Tumor-derived DNA and RNA can be released and circulated in the peripheral circulation of cancer patients, allowing for non-invasive gene expression profiling by body fluid analysis. Indeed, there is a wealth of information indicating a correlation between tumor-associated changes in genomic, epigenetic, or transcriptional patterns and alterations in the levels of cell-free circulating nucleic acids (cfCNAs).78 Our recent studies and others have suggested that cancer-specific DNA, mRNA, and miRNAs are stable and detectable in the peripheral blood of cancer patients, suggesting that cfCNAs may be promising biomarkers for cancer diagnosis and monitoring.6, 7, 79, 80, 81 Notably, similar to other nucleic acids, lncRNAs can also be present in peripheral blood components, such as serum, plasma, and peripheral blood mononuclear cells.82, 83, 84 The peripheral lncRNA profiles may potentially contribute to the characterization of novel biomarkers for CRC patients. Indeed, some lncRNAs appear to have strong diagnostic potential as blood biomarkers for colorectal neoplasia. For example, the expression of colorectal neoplasia differentially expressed-h in the plasma had a sensitivity of 87% and a specificity of 93% for detecting CRC.82 It is likely that circulating lncRNAs may represent new, relatively non-invasive molecular markers of tumor activity. However, no lncRNA-based diagnostics have been developed for use in CRC as of yet.
Because most cases of CRC usually undergo sequential progression, and polypectomy can prevent colorectal adenomas from developing into malignant lesions, biomarkers for the clinical screening of premalignant conditions could result in early detection and early treatment of CRC, leading to the prevention of advanced disease. Recently, based on its unique expression characteristics in CRC, the lncRNA CCAT1 has emerged as a potential biomarker for screening precancerous lesions. CCAT1 is markedly overexpressed in CRC84 and is also upregulated in precancerous tissues, including benign inflammatory colonic tissues and adenomatous polyps.85 CCAT1 can be detected in CRC tissue and in the blood and stool samples of CRC patients.86 Thus a CCAT1-based blood/stool assay may be explored for the screening and early detection of CRC.
Tumor Prognosis
Several studies have documented a link between the dysregulation of lncRNAs and the pathogenesis/prognosis of CRC (Table 1); thus lncRNA signatures may be utilized as a valuable tool for the prediction of disease outcomes in conjunction with the prevailing mRNA and miRNA expression signatures. The expression level of uc.73a is lower in CRC tissues than in the corresponding noncancerous tissues;87 moreover, patients with low uc.73a expression have a relatively poor overall survival. Similarly, the expression level of PCAT-1 in 108 CRC patients can successfully distinguish between short-term and long-term survivors.88 Our group also found that the downregulation of LOC285194 is correlated with several clinicopathological factors, such as tumor size and metastasis, in patients suffering from CRC. Indeed, in a multivariate analysis with other clinicopathological risk factors, LOC285194 served as a significant and independent predictor of disease-specific survival.63 These lncRNAs may be exploited as novel prognostic tools to aid in CRC patient assessment and management in oncology clinics.
Particularly, some lncRNAs in CRC tissue samples exhibit differential expression patterns during specific tumor stages or in specific locations. One such lncRNA, HOTAIR, was highly expressed in cancerous tissues compared with the corresponding noncancerous tissues in stage IV CRC patients, and the levels of HOTAIR were correlated with liver metastasis.20 HULC is an lncRNA that was identified in human HCC tissues; intriguingly, HULC expression was found to be high in CRC liver metastatic nodules but not in primary CRC tissues or the corresponding normal tissues.89 These results suggest that individual lncRNAs or lncRNA panels could potentially be utilized as tailored biomarkers for monitoring CRC metastasis. With the aid of the appropriate pharmaceutical vehicles, such as lncRNA-specific peptide nucleic acid (PNA)-based molecular beacons,90 it may be possible to differentiate malignant lesions from benign tissues and to identify metastatic lesions during surgery using real-time in vivo imaging in the near future.
Tumor Treatment
The expression levels of lncRNAs may aid in tumor diagnosis and prognosis; additionally, lncRNAs may be interesting therapeutic targets for CRC. The finding that certain pathologically dysregulated lncRNAs have crucial roles in the modulation of tumor behavior by acting as oncogenes (onco-lncRNAs) or tumor suppressors (TS-lncRNAs) suggests the potential for the development of lncRNA-oriented therapies (Figure 2).
SiRNAs can easily be designed to inhibit the function of specific lncRNAs, and the regulation of specific lncRNAs that are tightly linked to key carcinogenesis processes may have promising anticancer effects. In the case of CRC, the administration of specific siRNAs targeting the oncogenic HOTAIR, CCAT2, and uc.73a transcripts was shown to decrease the invasion or proliferation of colon cancer cells.20, 58, 62 Although the results presented in these studies are only preliminary in vitro data, these reports suggest that RNAi-based therapy strategies are amenable to further development, particularly for cellular therapies.
In contrast to blocking onco-lncRNAs, restoring the expression of TS-lncRNAs in cancer patients could also achieve anticancer effects based on their intrinsic tumor-suppressive functions. By implanting colon cancer cells into nude mice after transfection with LOC285194 expression vectors, Liu et al64 successfully demonstrated the tumor-suppressive effects of LOC285194 in vivo; both the tumor growth rate and the final tumor weight were significantly decreased when LOC285194 was overexpressed.
Given that lncRNAs can have structural interactions with other molecules, therapeutic effects can also be achieved by disrupting their structures or functional motifs or by modulating their transcriptional activity. For example, one study reported that targeting the interactions between Xist and chromatin remodeling complexes resulted in the activation of PRC2-regulated genes.91 However, the practical feasibility of these novel strategies has not been studied in detail for CRC.
The acquisition of drug resistance is one of the main obstacles encountered in cancer chemotherapy, and some publications have suggested that lncRNAs may be critical regulators of drug sensitivity or resistance.92, 93 Cancer upregulated drug resistant (CUDR) is an lncRNA that exhibits elevated expression in multiple cancers, including CRC.94, 95 Tsang et al94 provided evidence that increasing CUDR gene levels in vitro in human squamous carcinoma cells resulted in increased resistance to doxorubicin and etoposide and decreased levels of drug-induced apoptosis via the downregulation of caspase-3. The question of whether similar mechanisms operate in CRC should be examined further, but this observation suggests that lncRNA manipulation-based gene therapy may be beneficial in overcoming drug resistance and in designing personalized therapeutics for cancer patients.
Additionally, to reduce the treatment-related side effects during transgene-mediated treatment, the use of certain tumor-specific lncRNAs as therapeutic agents in CRC is currently being explored in preclinical models. For example, H19 is highly expressed in cancer tissues and is only marginally or not at all expressed in the corresponding normal tissues. This characteristic was utilized to refine the regional therapeutic methods, and a chain plasmid vector driven by specific H19 regulatory sequences was developed. This plasmid harbors diphtheria toxin-A and was administered intra-arterially to animals with colon cancer liver metastases. The plasmid was selectively distributed to the tumor region and successfully delayed tumor growth.96
Conclusions and future perspectives
There is no doubt that the dysregulation of lncRNAs affects various cancer-related signaling pathways and has a significant role in tumor development. Currently, a variety of large-scale genomic studies, such as TCGA, are being used to investigate the abnormal expression profiles of lncRNAs in tumor;97 however, the molecular nature of most lncRNAs remains poorly characterized. Due to their complicated structural characteristics, tissue-specific expression, and specific temporal and spatial expression patterns, the detailed functions and mechanisms of one lncRNA may be significantly different in different tumors.24 Therefore, to attain a thorough understanding of the role of lncRNAs, besides to elucidate the expression patterns of lncRNAs in CRC, further structural, functional, and mechanistic characterizations should also be performed.
Nevertheless, while fulfilling critical roles as transcriptional, post-transcriptional, or epigenetic regulators, lncRNAs themselves are extensively modulated by multiple inherited genetic and epigenetic alterations in addition to structural variations and transcriptional regulators; all of these factors contribute to their dysregulation in malignancies and are consequently implicated in the etiology and pathophysiology of cancer. For this reason, it is of great interest to investigate the association between these lncRNA regulatory mechanisms and their subsequent effects on lncRNA structure, expression, and function in CRC and other cancers.
Moving forward, the systematic identification and annotation of tissue-specific lncRNA signatures and their expression patterns in tumors hold great promise for the development of accurate, non-invasive biomarkers that can be utilized in early CRC detection and prognosis prediction. Although several studies have already uncovered some candidate lncRNAs and demonstrated their association with clinicopathological characteristics, there are considerable limitations to the translation of lncRNAs into clinical practice. First, one of the prerequisites of an ideal cancer marker is stability, and the stability of circulating lncRNAs remains largely unknown. Second, although numerous studies have demonstrated the presence of lncRNAs in the circulation, the levels of lncRNA transcripts and their post-transcriptional modifications are unstable and variable or are difficult to detect during different disease stages. Third, there is no simple standard assay or universal endogenous control/control set for the quantification of circulating lncRNAs. Finally, it is unknown whether these lncRNAs are secreted from the cells of tissues or whether they are present due to hematocyte contamination. It is difficult to determine the origins of circulating lncRNAs that have been isolated and quantified. Future approaches should focus on overcoming these obstacles. Even if these problems are successfully solved, the lncRNA biomarkers remain needs for further analytic and clinical validation, and demonstration of clinical utility before it should be considered for use in general practice to become generally useful.98
Finally, investigations of the molecular mechanism(s) by which lncRNAs act and in vivo animal studies exploring lncRNA intervention demonstrate the potential for the use of lncRNAs in novel anticancer therapies. Like other important ncRNAs, such as miRNAs, lncRNAs are also natural nucleic acids that regulate many gene networks involved in cancer cell transformation; thus the rationale for using lncRNAs in CRC therapies is clear. However, several common challenges in RNA therapeutics, such as the lack of reliable delivery methods, limited effective vector types, lack of optimal dosage regimes, and side effects, have made lncRNA-based therapy difficult to achieve. Moreover, due to the large size and extensive secondary structures of lncRNAs, it is difficult to interfere with their expression using conventional RNAi technologies, such as siRNA and antisense oligonucleotides, and it is also difficult to use them directly as therapeutic agents Because these technical bottlenecks cannot be ignored, novel effective and stable strategies for genome editing—as well as effective gene therapy delivery systems—warrant further investigation. Nevertheless, although at present it may be too optimistic or premature to affirm the potential of lncRNA-targeted treatment, the momentum in the lncRNA-based research field will undoubtedly stimulate research in the field of ideal therapeutics for CRC patients in the near future.
References
Birney E, Stamatoyannopoulos JA, Dutta A et al. Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature 2007;447:799–816.
Mercer TR, Dinger ME, Mattick JS . Long non-coding RNAs: insights into functions. Nat Rev Genet 2009;10:155–159.
Takizawa T, Gemma A, Ui-Tei K et al. Basic and clinical studies on functional RNA molecules for advanced medical technologies. J Nippon Med Sch 2010;77:71–79.
Shah AA, Leidinger P, Blin N et al. miRNA: small molecules as potential novel biomarkers in cancer. Curr Med Chem 2010;17:4427–4432.
Huang Z, Huang S, Wang Q et al. MicroRNA-95 promotes cell proliferation and targets sorting Nexin 1 in human colorectal carcinoma. Cancer Res 2011;71:2582–2589.
Wang Q, Huang Z, Ni S et al. Plasma miR-601 and miR-760 are novel biomarkers for the early detection of colorectal cancer. PLoS One 2012;7:e44398.
Huang Z, Huang D, Ni S et al. Plasma microRNAs are promising novel biomarkers for early detection of colorectal cancer. Int J Cancer 2010;127:118–126.
Frith MC, Pheasant M, Mattick JS . The amazing complexity of the human transcriptome. Eur J Hum Genet 2005;13:894–897.
Ponting CP, Oliver PL, Reik W . Evolution and functions of long noncoding RNAs. Cell 2009;136:629–641.
Li X, Wu Z, Fu X et al. Long noncoding RNAs: insights from biological features and functions to diseases. Med Res Rev 2013;33:517–553.
Spizzo R, Almeida MI, Colombatti A et al. Long non-coding RNAs and cancer: a new frontier of translational research? Oncogene 2012;31:4577–4587.
Volders PJ, Helsens K, Wang X et al. LNCipedia: a database for annotated human lncRNA transcript sequences and structures. Nucleic Acids Res 2013;41:D246–D251.
Rinn JL, Chang HY . Genome regulation by long noncoding RNAs. Annu Rev Biochem 2012;81:145–166.
Chen LL, Carmichael GG . Decoding the function of nuclear long non-coding RNAs. Curr Opin Cell Biol 2010;22:357–364.
Prensner JR, Chinnaiyan AM . The emergence of lncRNAs in cancer biology. Cancer Discov 2011;1:391–407.
Huarte M, Rinn JL . Large non-coding RNAs: missing links in cancer? Hum Mol Genet 2010;19:R152–R161.
Gibb EA, Brown CJ, Lam WL . The functional role of long non-coding RNA in human carcinomas. Mol Cancer 2011;10:38.
Michelhaugh SK, Lipovich L, Blythe J et al. Mining Affymetrix microarray data for long non-coding RNAs: altered expression in the nucleus accumbens of heroin abusers. J Neurochem 2011;116:459–466.
Atkinson SR, Marguerat S, Bahler J . Exploring long non-coding RNAs through sequencing. Semin Cell Dev Biol 2012;23:200–205.
Kogo R, Shimamura T, Mimori K et al. Long noncoding RNA HOTAIR regulates polycomb-dependent chromatin modification and is associated with poor prognosis in colorectal cancers. Cancer Res 2011;71:6320–6326.
Yu G, Yao W, Wang J et al. LncRNAs expression signatures of renal clear cell carcinoma revealed by microarray. PLoS One 2012;7:e42377.
Gibb EA, Becker-Santos DD, Enfield KS et al. Aberrant expression of long noncoding RNAs in cervical intraepithelial neoplasia. Int J Gynecol Cancer 2012;22:1557–1563.
Brunner AL, Beck AH, Edris B et al. Transcriptional profiling of long non-coding RNAs and novel transcribed regions across a diverse panel of archived human cancers. Genome Biol 2012;13:R75.
Hauptman N, Glavac D . Long non-coding RNA in cancer. Int J Mol Sci 2013;14:4655–4669.
Yang Z, Zhou L, Wu LM et al. Overexpression of long non-coding RNA HOTAIR predicts tumor recurrence in hepatocellular carcinoma patients following liver transplantation. Ann Surg Oncol 2011;18:1243–1250.
Geng YJ, Xie SL, Li Q et al. Large intervening non-coding RNA HOTAIR is associated with hepatocellular carcinoma progression. J Int Med Res 2011;39:2119–2128.
Kim K, Jutooru I, Chadalapaka G et al. HOTAIR is a negative prognostic factor and exhibits pro-oncogenic activity in pancreatic cancer. Oncogene 2013;32:1616–1625.
Xu ZY, Yu QM, Du YA et al. Knockdown of long non-coding RNA HOTAIR suppresses tumor invasion and reverses epithelial-mesenchymal transition in gastric cancer. Int J Biol Sci 2013;9:587–597.
Gupta RA, Shah N, Wang KC et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature 2010;464:1071–1076.
Tang L, Zhang W, Su B et al. Long noncoding RNA HOTAIR is associated with motility, invasion, and metastatic potential of metastatic melanoma. Biomed Res Int 2013;2013:251098.
Rinn JL, Kertesz M, Wang JK et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 2007;129:1311–1323.
Tsai MC, Manor O, Wan Y et al. Long noncoding RNA as modular scaffold of histone modification complexes. Science 2010;329:689–693.
Qi P, Du X . The long non-coding RNAs, a new cancer diagnostic and therapeutic gold mine. Mod Pathol 2013;26:155–165.
Jemal A, Bray F, Center MM et al. Global cancer statistics. CA Cancer J Clin 2011;61:69–90.
Gellad ZF, Provenzale D . Colorectal cancer: national and international perspective on the burden of disease and public health impact. Gastroenterology 2010;138:2177–2190.
Schweiger MR, Hussong M, Rohr C et al. Genomics and epigenomics of colorectal cancer. Wiley Interdiscip Rev Syst Biol Med 2013;5:205–219.
McHugh SM, O'Donnell J, Gillen P . Genomic and oncoproteomic advances in detection and treatment of colorectal cancer. World J Surg Oncol 2009;7:36.
Jimenez CR, Knol JC, Meijer GA et al. Proteomics of colorectal cancer: overview of discovery studies and identification of commonly identified cancer-associated proteins and candidate colorectal cancer serum markers. J Proteomics 2010;73:1873–1895.
Goel A, Boland CR . Epigenetics of colorectal cancer. Gastroenterology 2012;143:e1.
Creighton CJ, Morgan M, Gunaratne PH et al. Comprehensive molecular characterization of clear cell renal cell carcinoma. Nature 2013;499:43–49.
Koboldt DC, Fulton RS, McLellan MD et al. Comprehensive molecular portraits of human breast tumours. Nature 2012;490:61–70.
Muzny DM, Bainbridge MN, Chang K et al. Comprehensive molecular characterization of human colon and rectal cancer. Nature 2012;487:330–337.
Colussi D, Brandi G, Bazzoli F et al. Molecular pathways involved in colorectal cancer: implications for disease behavior and prevention. Int J Mol Sci 2013;14:16365–16385.
Yamauchi M, Morikawa T, Kuchiba A et al. Assessment of colorectal cancer molecular features along bowel subsites challenges the conception of distinct dichotomy of proximal versus distal colorectum. Gut 2012;61:847–854.
Rosty C, Young JP, Walsh MD et al. PIK3CA activating mutation in colorectal carcinoma: associations with molecular features and survival. PLoS One 2013;8:e65479.
Liao X, Lochhead P, Nishihara R et al. Aspirin use, tumor PIK3CA mutation, and colorectal-cancer survival. N Engl J Med 2012;367:1596–1606.
Ogino S, Lochhead P, Chan AT et al. Molecular pathological epidemiology of epigenetics: emerging integrative science to analyze environment, host, and disease. Mod Pathol 2013;26:465–484.
Wang D, Dubois RN . The role of COX-2 in intestinal inflammation and colorectal cancer. Oncogene 2010;29:781–788.
Grivennikov SI, Greten FR, Karin M . Immunity, inflammation, and cancer. Cell 2010;140:883–899.
Galon J, Mlecnik B, Bindea G et al. Towards the introduction of the Immunoscore in the classification of malignant tumors. J Pathol 2013;232:199–209.
Bardhan K, Liu K . Epigenetics and colorectal cancer pathogenesis. Cancers (Basel) 2013;5:676–713.
Lao VV, Grady WM . Epigenetics and colorectal cancer. Nat Rev Gastroenterol Hepatol 2011;8:686–700.
Stein RA . Epigenetics-the link between infectious diseases and cancer. JAMA 2011;305:1484–1485.
Supiot S, Gouraud W, Campion L et al. Early dynamic transcriptomic changes during preoperative radiotherapy in patients with rectal cancer: a feasibility study. World J Gastroenterol 2013;19:3249–3254.
Hashemi J, Worrall C, Vasilcanu D et al. Molecular characterization of acquired tolerance of tumor cells to picropodophyllin (PPP). PLoS One 2011;6:e14757.
Lakhotia SC . Long non-coding RNAs coordinate cellular responses to stress. Wiley Interdiscip Rev RNA 2012;3:779–796.
Tanaka K, Shiota G, Meguro M et al. Loss of imprinting of long QT intronic transcript 1 in colorectal cancer. Oncology 2001;60:268–273.
Ling H, Spizzo R, Atlasi Y et al. CCAT2, a novel non-coding RNA mapping to 8q24, underlies metastatic progression and chromosomal instability in colon cancer. Genome Res 2013;23:1446–1461.
Wang J, Liu X, Wu H et al. CREB up-regulates long non-coding RNA, HULC expression through interaction with microRNA-372 in liver cancer. Nucleic Acids Res 2010;38:5366–5383.
Poliseno L, Salmena L, Zhang J et al. A coding-independent function of gene and pseudogene mRNAs regulates tumour biology. Nature 2010;465:1033–1038.
Cesana M, Cacchiarelli D, Legnini I et al. A long noncoding RNA controls muscle differentiation by functioning as a competing endogenous RNA. Cell 2011;147:358–369.
Calin GA, Liu CG, Ferracin M et al. Ultraconserved regions encoding ncRNAs are altered in human leukemias and carcinomas. Cancer Cell 2007;12:215–229.
Qi P, Xu MD, Ni SJ et al. Low expression of LOC285194 is associated with poor prognosis in colorectal cancer. J Transl Med 2013;11:122.
Liu Q, Huang J, Zhou N et al. LncRNA loc285194 is a p53-regulated tumor suppressor. Nucleic Acids Res 2013;41:4976–4987.
Pasic I, Shlien A, Durbin AD et al. Recurrent focal copy-number changes and loss of heterozygosity implicate two noncoding RNAs and one tumor suppressor gene at chromosome 3q13.31 in osteosarcoma. Cancer Res 2010;70:160–171.
Davison EJ, Tarpey PS, Fiegler H et al. Deletion at chromosome band 20p12.1 in colorectal cancer revealed by high resolution array comparative genomic hybridization. Genes Chromosomes Cancer 2005;44:384–391.
Xu C, Yang M, Tian J et al. MALAT-1: a long non-coding RNA and its important 3' end functional motif in colorectal cancer metastasis. Int J Oncol 2011;39:169–175.
Yang F, Huo XS, Yuan SX et al. Repression of the long noncoding RNA-LET by histone deacetylase 3 contributes to hypoxia-mediated metastasis. Mol Cell 2013;49:1083–1096.
Barsotti AM, Beckerman R, Laptenko O et al. p53-Dependent induction of PVT1 and miR-1204. J Biol Chem 2012;287:2509–2519.
Mattick JS . The genetic signatures of noncoding RNAs. PLoS Genet 2009;5:e1000459.
Haiman CA, Le Marchand L, Yamamato J et al. A common genetic risk factor for colorectal and prostate cancer. Nat Genet 2007;39:954–956.
Tomlinson I, Webb E, Carvajal-Carmona L et al. A genome-wide association scan of tag SNPs identifies a susceptibility variant for colorectal cancer at 8q24.21. Nat Genet 2007;39:984–988.
Zhu Z, Gao X, He Y et al. An insertion/deletion polymorphism within RERT-lncRNA modulates hepatocellular carcinoma risk. Cancer Res 2012;72:6163–6172.
Jendrzejewski J, He H, Radomska HS et al. The polymorphism rs944289 predisposes to papillary thyroid carcinoma through a large intergenic noncoding RNA gene of tumor suppressor type. Proc Natl Acad Sci USA 2012;109:8646–8651.
Verhaegh GW, Verkleij L, Vermeulen SH et al. Polymorphisms in the H19 gene and the risk of bladder cancer. Eur Urol 2008;54:1118–1126.
Xue Y, Wang M, Kang M et al. Association between lncrna PCGEM1 polymorphisms and prostate cancer risk. Prostate Cancer Prostatic Dis 2013;16:S1.
Wojcik SE, Rossi S, Shimizu M et al. Non-codingRNA sequence variations in human chronic lymphocytic leukemia and colorectal cancer. Carcinogenesis 2010;31:208–215.
Schwarzenbach H, Hoon DS, Pantel K . Cell-free nucleic acids as biomarkers in cancer patients. Nat Rev Cancer 2011;11:426–437.
Shen J, Stass SA, Jiang F . MicroRNAs as potential biomarkers in human solid tumors. Cancer Lett 2013;329:125–136.
Xu Y, Xu Q, Yang L et al. Identification and validation of a blood-based 18-gene expression signature in colorectal cancer. Clin Cancer Res 2013;19:3039–3049.
Yu M . Circulating cell-free mitochondrial DNA as a novel cancer biomarker: opportunities and challenges. Mitochondrial DNA 2012;23:329–332.
Graham LD, Pedersen SK, Brown GS et al. Colorectal neoplasia differentially expressed (CRNDE), a novel gene with elevated expression in colorectal adenomas and adenocarcinomas. Genes Cancer 2011;2:829–840.
Huang X, Yuan T, Tschannen M et al. Characterization of human plasma-derived exosomal RNAs by deep sequencing. BMC Genomics 2013;14:319.
Nissan A, Stojadinovic A, Mitrani-Rosenbaum S et al. Colon cancer associated transcript-1: a novel RNA expressed in malignant and pre-malignant human tissues. Int J Cancer 2012;130:1598–1606.
Alaiyan B, Ilyayev N, Stojadinovic A et al. Differential expression of colon cancer associated transcript1 (CCAT1) along the colonic adenoma-carcinoma sequence. BMC Cancer 2013;13:196.
Cheng J, Kapranov P, Drenkow J et al. Transcriptional maps of 10 human chromosomes at 5-nucleotide resolution. Science 2005;308:1149–1154.
Sana J, Hankeova S, Svoboda M et al. Expression levels of transcribed ultraconserved regions uc.73 and uc.388 are altered in colorectal cancer. Oncology 2012;82:114–118.
Ge X, Chen Y, Liao X et al. Overexpression of long noncoding RNA PCAT-1 is a novel biomarker of poor prognosis in patients with colorectal cancer. Med Oncol 2013;30:588.
Matouk IJ, Abbasi I, Hochberg A et al. Highly upregulated in liver cancer noncoding RNA is overexpressed in hepatic colorectal metastasis. Eur J Gastroenterol Hepatol 2009;21:688–692.
Kam Y, Rubinstein A, Naik S et al. Detection of a long non-coding RNA (CCAT1) in living cells and human adenocarcinoma of colon tissues using FIT-PNA molecular beacons. Cancer Lett 2013;pii: S0304-3835:00126–2.
Sarma K, Levasseur P, Aristarkhov A et al. Locked nucleic acids (LNAs) reveal sequence requirements and kinetics of Xist RNA localization to the X chromosome. Proc Natl Acad Sci USA 2010;107:22196–22201.
Wang F, Li X, Xie X et al. a non-protein-coding RNA up-regulated in bladder carcinoma and embryo, influencing cell growth and promoting invasion. FEBS Lett 2008;582:1919–1927.
Yang Y, Li H, Hou S et al. The noncoding RNA expression profile and the effect of lncRNA AK126698 on cisplatin resistance in non-small-cell lung cancer cell. PLoS One 2013;8:e65309.
Tsang WP, Wong TW, Cheung AH et al. Induction of drug resistance and transformation in human cancer cells by the noncoding RNA CUDR. RNA 2007;13:890–898.
Wang Y, Chen W, Yang C et al. Long non-coding RNA UCA1a(CUDR) promotes proliferation and tumorigenesis of bladder cancer. Int J Oncol 2012;41:276–284.
Sorin V, Ohana P, Mizrahi A et al. Regional therapy with DTA-H19 vector suppresses growth of colon adenocarcinoma metastases in the rat liver. Int J Oncol 2011;39:1407–1412.
Zhang XQ, Sun S, Lam KF et al. A long non-coding RNA signature in glioblastoma multiforme predicts survival. Neurobiol Dis 2013;58C:123–131.
Febbo PG, Ladanyi M, Aldape KD et al. NCCN Task Force report: evaluating the clinical utility of tumor markers in oncology. J Natl Compr Canc Netw 2011;9 (Suppl 5):S1–S32 quiz S3.
Ellis BC, Molloy PL, Graham LD . CRNDE: a long non-coding RNA involved in cancer, neurobiology, and development. Front Genet 2012;3:270.
Yochum GS, Cleland R, McWeeney S et al. An antisense transcript induced by Wnt/beta-catenin signaling decreases E2F4. J Biol Chem 2007;282:871–878.
Yoshimizu T, Miroglio A, Ripoche MA et al. The H19 locus acts in vivo as a tumor suppressor. Proc Natl Acad Sci USA 2008;105:12417–12422.
Tsang WP, Ng EK, Ng SS et al. Oncofetal H19-derived miR-675 regulates tumor suppressor RB in human colorectal cancer. Carcinogenesis 2010;31:350–358.
Tian F, Tang Z, Song G et al. Loss of imprinting of IGF2 correlates with hypomethylation of the H19 differentially methylated region in the tumor tissue of colorectal cancer patients. Mol Med Rep 2012;5:1536–1540.
Matouk IJ, DeGroot N, Mezan S et al. The H19 non-coding RNA is essential for human tumor growth. PLoS One 2007;2:e845.
Lassmann S, Weis R, Makowiec F et al. Array CGH identifies distinct DNA copy number profiles of oncogenes and tumor suppressor genes in chromosomal- and microsatellite-unstable sporadic colorectal carcinomas. J Mol Med (Berl) 2007;85:293–304.
Zhang X, Zhou Y, Mehta KR et al. A pituitary-derived MEG3 isoform functions as a growth suppressor in tumor cells. J Clin Endocrinol Metab 2003;88:5119–5126.
Zhou Y, Zhong Y, Wang Y et al. Activation of p53 by MEG3 non-coding RNA. J Biol Chem 2007;282:24731–24742.
Nakano S, Murakami K, Meguro M et al. Expression profile of LIT1/KCNQ1OT1 and epigenetic status at the KvDMR1 in colorectal cancers. Cancer Sci 2006;97:1147–1154.
Acknowledgements
This study was supported by the National Clinical Key Discipline (2011–2015), Clinical Key Discipline Fund by Ministry of Health (2010–2012), Academic Award for Doctoral Candidates by Ministry of Education (2012), Mingdao Fund for Medical Graduate Student by Shanghai Medical College (2013), and Shanghai Science and Technology Development Fund (Basic Research Major Project, No. 10DJ1400500).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Xu, Md., Qi, P. & Du, X. Long non-coding RNAs in colorectal cancer: implications for pathogenesis and clinical application. Mod Pathol 27, 1310–1320 (2014). https://doi.org/10.1038/modpathol.2014.33
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2014.33
Keywords
This article is cited by
-
Identification of long non-coding RNA using single nucleotide epimutation analysis: a novel gene discovery approach
Cancer Cell International (2022)
-
RAMS11 promotes CRC through mTOR-dependent inhibition of autophagy, suppression of apoptosis, and promotion of epithelial-mesenchymal transition
Cancer Cell International (2021)
-
Long non-coding RNA HOTAIR and HOTTIP as potential biomarkers for hepatitis C virus genotype 4-induced hepatocellular carcinoma
Egyptian Journal of Medical Human Genetics (2020)
-
Genome-wide expression profiling in colorectal cancer focusing on lncRNAs in the adenoma-carcinoma transition
BMC Cancer (2019)
-
Expression of long non-coding RNA CCHE1 in colorectal carcinoma: correlations with clinicopathological features and ERK/COX-2 pathway
Molecular Biology Reports (2019)