Abstract
Inflammatory myofibroblastic tumor is a distinctive, rarely metastasizing mesenchymal neoplasm composed of fascicles of spindle cells with a prominent inflammatory infiltrate. Roughly 50% of inflammatory myofibroblastic tumors harbor ALK receptor tyrosine kinase gene rearrangements. Such tumors are usually positive for ALK by immunohistochemistry. The molecular pathogenesis of ALK-negative inflammatory myofibroblastic tumors is largely unknown. A recent study identified rearrangements of ROS1 (another tyrosine kinase receptor) in a subset of ALK-negative inflammatory myofibroblastic tumors. Immunohistochemistry for ROS1 has been shown to correlate with ROS1 rearrangement in lung adenocarcinomas. The purpose of this study was to determine whether immunohistochemistry for ROS1 could predict ROS1 rearrangement in inflammatory myofibroblastic tumor. In total, 30 inflammatory myofibroblastic tumors were evaluated, including 21 ALK-positive tumors (10 confirmed to harbor ALK rearrangements, with TPM3, CLTC, RANPB2, and FN1 fusion partners) and 9 ALK-negative tumors (including 2 known to harbor ROS1 rearrangements). Immunohistochemistry was performed on whole tissue sections following pressure cooker antigen retrieval using a rabbit anti-ROS1 monoclonal antibody. The results were scored as ‘positive’ or ‘negative,’ and the pattern of staining was recorded. Three ALK-negative inflammatory myofibroblastic tumors (including both tumors with known ROS1 rearrangements) showed immunoreactivity for ROS1, whereas all ALK-positive inflammatory myofibroblastic tumors were negative for ROS1. One ROS1-positive inflammatory myofibroblastic tumor (with YWHAE-ROS1 fusion) showed strong, diffuse cytoplasmic and nuclear staining; one case (with TFG-ROS1 fusion) showed weak, diffuse and dot-like cytoplasmic staining; and one case (fusion partner unknown) showed moderate, diffuse and dot-like cytoplasmic staining. Expression of ROS1 correlates with ROS1 gene rearrangement in inflammatory myofibroblastic tumor. These findings suggest that immunohistochemistry for ROS1 may be useful to support the diagnosis of a subset of inflammatory myofibroblastic tumors and may select some clinically aggressive cases for targeted therapy directed against ROS1.
Similar content being viewed by others
Main
Inflammatory myofibroblastic tumor is a distinctive mesenchymal neoplasm with a predilection for the lung, retroperitoneum, and abdominal cavity of children and young adults.1 This tumor type is typically composed of fascicles of myofibroblastic spindle cells and usually shows a marked inflammatory infiltrate, chiefly of lymphocytes and plasma cells. Inflammatory myofibroblastic tumor belongs to the ‘intermediate, rarely metastasizing’ managerial category in the World Health Organization classification of tumors of soft tissue and bone, based on its low (<3%) rate of metastasis.2 In general, histologic features do not correlate well with clinical behavior,3 with the exception of an aggressive epithelioid variant (epithelioid inflammatory myofibroblastic sarcoma) that arises in the mesentery and omentum and shows distinctive histologic appearances, including myxoid stroma and a predominantly neutrophilic inflammatory infiltrate.4, 5
Around 50% of inflammatory myofibroblastic tumors harbor rearrangements involving the ALK (anaplastic lymphoma kinase) gene; diverse fusion partners have been identified.6, 7, 8, 9, 10, 11, 12 Recent studies have suggested that treatment with tyrosine kinase inhibitors such as crizotinib benefit patients with aggressive inflammatory myofibroblastic tumors with ALK rearrangements.13, 14 By immunohistochemistry, positive staining for ALK correlates well with the presence of an ALK rearrangement.1, 6, 15 ALK staining is therefore both a helpful diagnostic marker and a predictive marker for targeted therapy in this tumor type, similar to the small subset of lung adenocarcinomas with ALK rearrangements.16, 17 Until recently, the molecular pathogenesis of ALK-negative inflammatory myofibroblastic tumors was unknown. Using a massively parallel (next-generation) DNA sequencing strategy, a member of our group recently discovered that a subset of ALK-negative inflammatory myofibroblastic tumors harbors rearrangements involving ROS1 (v-ros avian UR2 sarcoma virus oncogene homolog 1),18 which encodes a tyrosine kinase receptor similar to ALK. Immunohistochemistry for ROS1 has been shown to correlate with ROS1 rearrangement in lung adenocarcinoma.19, 20, 21 Expression of ROS1 in inflammatory myofibroblastic tumors has not previously been evaluated. The purpose of this study was to determine whether immunohistochemistry for ROS1 could predict ROS1 rearrangement in inflammatory myofibroblastic tumor.
Materials and methods
Tumor Samples
Archival tumor samples from 30 inflammatory myofibroblastic tumors were evaluated, including 21 ALK-positive tumors (10 previously confirmed to harbor ALK rearrangements) and 9 ALK-negative tumors (including 2 known to harbor ROS1 rearrangements).18 The gene fusion status of 10 of these tumors was previously published.5, 18 Archival samples from 60 other spindle cell neoplasms in the differential diagnosis with inflammatory myofibroblastic tumor were also evaluated, including 10 cases each of desmoid fibromatosis; myofibroblastic sarcoma; gastrointestinal stromal tumor, spindle cell type; leiomyosarcoma; follicular dendritic cell sarcoma; and dedifferentiated liposarcoma (5 morphologically low grade and 5 with the so-called ‘inflammatory MFH’ pattern).
Immunohistochemistry
Immunohistochemistry was performed on 4-μm-thick formalin-fixed paraffin-embedded whole tissue sections following pressure cooker antigen retrieval (0.001 M citrate buffer; pH 6.0), using a rabbit anti-ROS1 monoclonal antibody (1:100 dilution; 40 min incubation; clone D4D6; Cell Signaling Technology, Danvers, MA). The Envision Plus detection system (Dako, Carpinteria, CA) was used as a secondary antibody. The results were scored as ‘positive’ or ‘negative,’ and the pattern and intensity of staining were recorded. Weak nuclear staining alone was considered ‘negative.’ A lung adenocarcinoma with confirmed ROS1 rearrangement served as a positive control.
Fluorescence In Situ Hybridization (FISH)
FISH analysis was performed on interphase nuclei on 5-μm-thin sections of formalin-fixed paraffin-embedded tissue from one case that was found to be positive for ROS1 by immunohistochemistry (see Results), according to the standard protocols. ROS1 rearrangement was evaluated using the Kreatech Repeat-Free Poseidon ROS1 Break Probe kit (Kreatech Inc., Durham, NC), which contains two differentially labeled probes that flank the ROS1 gene, located at 6q22. Probes and nuclei were co-denatured simultaneously, followed by hybridization and washing, according to the manufacturer’s directions. An intact ROS1 locus is represented by two fused red/green signals. However, if a ROS1 rearrangement has occurred, one copy of the red probe will be separated from the adjacent green probe by ≥1 signal diameter, or the 5′ (green) probe signal will be lost, as previously described for lung adenocarcinomas.20
Next-Generation Sequencing
Next-generation DNA sequencing was performed on one case found to be positive for ROS1 by immunohistochemistry (with previously unknown genotype). In brief, hematoxylin and eosin-stained slides were reviewed to evaluate tumor content and viability and an area enriched for >20% tumor was circled. The entire slides from paired 5-μm-thick formalin-fixed paraffin-embedded tissue sections were scraped. Samples were digested in proteinase K overnight, and DNA was isolated according to the manufacturer’s protocol (QIAamp DNA Mini Kit, Qiagen, Gaithersburg, MD). DNA concentration was assessed using PicoGreen ds DNA detection (Life Technologies, Carlsbad, CA). The sample was subjected to targeted next-generation sequencing using a cancer genomic assay to detect sequence and copy number variations in 275 oncogenes and tumor-suppressor genes. The entire exonic sequence of the 275 target genes and selected intronic sequences of 31 genes were captured using a solution-phase Agilent SureSelect hybrid capture kit (Agilent Technologies, Inc., Santa Clara, CA) and massively parallel sequencing was performed on an Illumina HiSeq 2500 sequencer (Illumina, Inc., San Diego, CA). Mutation calls were made using Mutect and GATK software (Broad Institute, Cambridge, MA), copy number alterations were assessed using VisCap Copy (Dana Farber Cancer Institute, Boston, MA), and structural variations including rearrangements and large insertion/deletion events were detected using BreakMer (Dana Farber Cancer Institute).
Reverse Transcriptase (RT)-PCR and PCR
PCR and RT-PCR were then performed using primers that flank the breakpoints in the known YWHAE-ROS1 and TFG-ROS1 fusions.18 In brief, DNA and RNA were extracted from formalin-fixed paraffin-embedded tumor samples using the Qiagen AllPrep DNA/RNA kit according to the manufacturer’s protocol. RNA integrity was determined by Agilent BioAnalyzer. RNA and DNA from BA/F3 cells transduced with YWHAE-ROS1 or TFG-ROS1 cDNA constructs were used as positive controls, and RNA and DNA from BA/F3 parental cells were used as negative controls. RNA and DNA from BA/F3 cells were extracted using the Qiagen DNeasy or RNeasy kits according to the manufacturer’s protocols.
RT-PCR was performed using 1 μg of RNA with the Invitrogen One Step RT-PCR kit (Life Technologies) according to the manufacturer’s protocol with primers flanking breakpoints in control cells for YWHAE-ROS1 and TFG-ROS1. Primers were designed by using the UCSC Genome database to ensure exclusive target binding. Primer sequences were as follows: YWHAE: 5′-GCCACAGGAAACGACAGGAAGGAGGC-3′; ROS1: 5′-GAAGAAGGGTTCCACAGACCAGGAG-3′; TFG: 5′-GATAGTTCTGACCTTTCCTTTGCAATTC-3′; ROS1: 5′-CTCCTCTTGGGTTGGAAGAG-3′. Annealing temperatures were 65 and 62 °C, respectively. Approximately 64% of the RT-PCR product was run on a 2% agarose-TAE gel, and the gel was visualized using the AlphaImager Mini (ProteinSimple, San Jose, CA).
PCR was performed using 200 ng DNA with the Invitrogen Platinum PCR SuperMix High Fidelity kit (Life Technologies), according to the manufacturer’s protocol with the following primers flanking known breakpoints in YWHAE-ROS1 and TFG-ROS1: YWHAE: 5′-GCCACAGGAAACGACAGGAAGGAGGC-3′; ROS1: 5′-GAAGAAGGGTTCCACAGACCAGGAG-3′; TFG: 5′-GTCAGGTGAAATATCTCCGTCGAGAAC-3′; ROS1: 5′-CTCCTCTTGGGTTGGAAGAG-3′. The annealing temperature for primers flanking YWHAE-ROS1 was 61.1 °C. The annealing temperature for primers flanking TFG-ROS1 could not be determined due to the presence of background bands in the negative controls (that were the same size as the expected product). Alternatively, DNA isolated from formalin-fixed paraffin-embedded tissue samples known to harbor YWHAE-ROS1 or TFG-ROS1 fusions was used as a positive control in an attempt to identify the exact breakpoint by PCR. DNA quantity and quality were assessed by nanodrop. PCR was performed as above with the following primer sequences: Direct-YWHAE-ROS1 forward: 5′-ATCGTAACTGAATGATCTAAATCAG-3′; TFG-ROS1 forward: 5′-TATTGTGAAATAGTAAAAGGGAAAGTTGAT-3′. An eight temperature gradient (from 56 to 66 °C) was used to perform PCR for the sample with the unknown fusion partner in an attempt to identify the breakpoint. For each experiment, approximately 64% of the PCR product was run on a 1% agarose-TAE gel, and the gel was visualized using the AlphaImager Mini.
Results
The clinical features of the 30 patients with inflammatory myofibroblastic tumor are summarized in Table 1. Sixteen patients were female, and 14 patients were male, including 16 adults and 14 children. Patient age at presentation ranged from 7 months to 56 years (mean: 24 years; median: 21.5 years). Eighteen tumors arose within the abdominal cavity, including six in the wall of the gastrointestinal tract (four stomach, one duodenum, and one colon); five lung; four somatic soft tissue (three lower extremity, one upper extremity); two urinary bladder; and one pelvis.
Of the 21 ALK-positive tumors, 17 showed cytoplasmic staining, and 4 showed a nuclear membrane pattern of staining. Ten of these tumors were previously confirmed to harbor ALK gene rearrangements (by FISH, next-generation sequencing, or RT-PCR, seven with known fusion partners (four RANBP2 and one each TPM3, CLTC, and FN1).5, 18
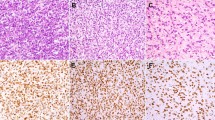
Immunohistochemistry for ROS1 was positive in three ALK-negative inflammatory myofibroblastic tumors (including both tumors with previously identified ROS1 rearrangements; Table 2; Figures 1 and 2),18 whereas all ALK-positive inflammatory myofibroblastic tumors were negative for ROS1 (Figure 3). One ROS1-positive inflammatory myofibroblastic tumor (with YWHAE-ROS1 fusion) showed strong, diffuse cytoplasmic and nuclear staining (Figure 1); one case (with TFG-ROS1 fusion) showed weak, diffuse and dot-like cytoplasmic staining (Figure 1); and one case (fusion partner unknown; see below) showed moderate, diffuse and dot-like cytoplasmic staining (Figure 2). None of the other tumor types (60 cases total) showed cytoplasmic or membranous staining for ROS1. Of note, a subset of tumors (six gastrointestinal stromal tumors, six myofibroblastic sarcomas, three leiomyosarcomas, and one follicular dendritic cell sarcoma) showed focal weak nuclear staining.
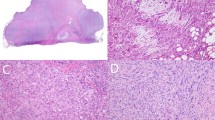
Inflammatory myofibroblastic tumors with ROS1 rearrangements. Inflammatory myofibroblastic tumor from the buttock of a 22-year-old woman composed of spindle cells with abundant pale cytoplasm (a, H&E). Note the stromal collagen and prominent neutrophils. Immunohistochemistry for ROS1 shows a cytoplasmic and nuclear staining pattern (b). This tumor harbors a YWHAE-ROS1 fusion. Inflammatory myofibroblastic tumor from the mesentery of an 8-year-old boy composed of fascicles of plump spindled and polygonal cells with amphophilic cytoplasm (c, H&E). Note the stromal lymphocytes. The tumor cells show weak diffuse and dot-like cytoplasmic reactivity for ROS1 (d). This tumor harbors a TFG-ROS1 fusion.
Inflammatory myofibroblastic tumor from the duodenum of a 56-year-old man. The tumor is grossly well circumscribed with a yellow cut surface (a, courtesy of Jiang Wang, MD, PhD, Cincinnati, OH). The tumor shows a fascicular growth pattern with numerous stromal lymphocytes and is composed of spindle cells with vesicular chromatin, prominent nucleoli, and palely eosinophilic cytoplasm (b, H&E). Immunohistochemistry for ROS1 shows diffuse and dot-like cytoplasmic staining (c). Florescence in situ hybridization showing loss of the 5′ signal in several cells and split-apart of the two signals corresponding to the 5′ and 3′ ends of ROS1 in one cell (upper left corner; d). These findings confirm the presence of a ROS1 rearrangement. The fusion partner could not be identified.
Inflammatory myofibroblastic tumor with CLTC-ALK fusion from the bladder of a 26-year-old woman. The tumor is composed of loose fascicles of spindle cells with stromal edema, lymphocytes, and plasma cells (a, H&E). Immunohistochemistry for ALK shows strong cytoplasmic staining (b). ROS1 is negative (c).
FISH, next-generation sequencing, RT-PCR, and PCR were performed on the ROS1-positive inflammatory myofibroblastic tumor with previously unknown ROS1 gene rearrangement status. FISH revealed rearrangement of ROS1 (either split signals or deletion of the 5′ region of ROS1) in 33 of 50 (66%) nuclei (Figure 2d), which exceeds the normal range (up to 6%) established in our laboratory. Isolated 3′ signals outnumbered split signals in tumor cells by a 2:1 ratio. This sample met our laboratory’s quality control standards for next-generation sequencing, with a mean target coverage of 142 (failing is <50) and 97% of targets sequenced at 30 × depth or greater. Next-generation sequencing of the tumor DNA failed to identify a ROS1 rearrangement; thus, the fusion partner could not be identified.
RT-PCR and PCR were performed using primers that flank breakpoints in known YWHAE-ROS1 and TFG-ROS1 fusions. We were unable to identify these two fusions in the sample by RT-PCR, although the RNA was severely degraded. Although BLAST searches for all primers showed exclusive target binding, bands of the target product size were present in negative control samples for PCR experiments performed with genomic DNA (data not shown). Therefore, we also performed PCR using primers that bind directly to the breakpoints in tumors harboring known YWHAE-ROS1 and TFG-ROS1 fusions. Unfortunately, the DNA from these formalin-fixed paraffin-embedded samples was also severely degraded. Nonetheless, we performed an eight temperature gradient PCR on the sample using primers that bind directly to the fusion breakpoints. However, we were unable to identify either the YWHAE-ROS1 or TFG-ROS1 fusion in the sample with this alternative method.
Discussion
Approximately 50% of inflammatory myofibroblastic tumors harbor ALK gene rearrangements and are positive for ALK by immunohistochemistry.1, 6 ALK is therefore a helpful marker to confirm the diagnosis of this tumor type. However, diagnosing ALK-negative inflammatory myofibroblastic tumors remains a significant challenge.1 When tumors arise in pediatric and adolescent patients in the lung or abdomen and show typical cytoarchitectural features (ie, fascicles of uniform spindle cells with vesicular nuclei, admixed with plasma cells and lymphocytes), the diagnosis of inflammatory myofibroblastic tumor is relatively straightforward even in the absence of ALK expression. In contrast, when such tumors arise in older patients or at unusual anatomic sites (eg, somatic soft tissues), or show notable nuclear atypia, more aggressive spindle cell sarcomas can be serious diagnostic considerations. ROS1 rearrangements were recently identified in a subset of ALK-negative inflammatory myofibroblastic tumors,18 suggesting a new diagnostic marker for this group of neoplasms.
In this study, we examined ROS1 expression by immunohistochemistry in inflammatory myofibroblastic tumor. We showed that ROS1 expression is limited to tumors with ROS1 rearrangements. Immunohistochemistry for ROS1 is consistently negative in ALK-positive inflammatory myofibroblastic tumors, as well as other spindle cell neoplasms that might be considered in the differential diagnosis, such as desmoid fibromatosis, myofibroblastic sarcoma, leiomyosarcoma, gastrointestinal stromal tumor, follicular dendritic cell sarcoma, and dedifferentiated liposarcoma. A subset of tumors showed focal, weak nuclear staining, which is a nonspecific finding that should not be misinterpreted as a positive result; cytoplasmic staining is required. The prevalence of ROS1 rearrangement in inflammatory myofibroblastic tumor is unknown. However, as three of nine ALK-negative inflammatory myofibroblastic tumors in this study were positive for ROS1, it seems that ROS1 rearrangements in this tumor type are not rare. As this was a biased cohort (the genotypes for 2 of these 9 tumors were already known), evaluation of larger series of ALK-negative inflammatory myofibroblastic tumors will be required to determine the frequency of ROS1 rearrangements in this tumor type.
The tyrosine kinase receptor genes ALK and ROS1 are remarkably similar in terms of their involvement in the pathogenesis of a wide range of tumor types of diverse lineages. ALK rearrangements are found in hematopoietic (anaplastic large cell lymphoma), mesenchymal (inflammatory myofibroblastic tumor), epithelial (lung adenocarcinoma and rare renal cell carcinomas), and melanocytic neoplasms (Spitz tumors).9, 22, 23, 24, 25, 26 ROS1 rearrangements have not yet been identified in hematopoietic tumors, but are found in these other lineages, as well as in a subset of glioblastomas.26, 27, 28, 29 This lineage promiscuity is also seen with other genes involved in translocations, such as EWSR1 and FUS.30, 31
Immunohistochemistry is a useful surrogate for molecular genetic techniques to detect an increasing number of gene fusions, not only ALK and ROS1 as discussed herein, but also ERG,32, 33, 34 WT1,35 TFE3,36 and STAT6,37, 38 among others. The introduction of such markers into clinical practice enables routine and widespread application, in some cases obviating the need for more costly molecular techniques. In this era of increasing emphasis on cost-containment, such an approach should prove valuable.
It is notable that somewhat different staining patterns were observed in the ROS1-positive inflammatory myofibroblastic tumors in this study. Two tumors (one with a TFG fusion partner) showed diffuse and dot-like cytoplasmic staining, whereas one tumor (with a YWHAE fusion partner) showed combined cytoplasmic and nuclear staining. The former finding suggested the possibility that the tumor for which the fusion partner could not be identified by next-generation sequencing might similarly harbor a TFG fusion. Both RT-PCR and PCR for TFG-ROS1 (and YWHAE-ROS1) were also negative; however, the RNA and DNA were of poor quality. We therefore cannot determine whether this tumor harbors one of these known fusion partners or whether another gene encoding a cytoplasmic protein might instead be involved. Most inflammatory myofibroblastic tumors with ALK rearrangements show a cytoplasmic pattern of staining by immunohistochemistry, although the fusion partners are diverse.1, 15 Whether the pattern of ROS1 staining correlates with the gene fusion partner remains to be determined.
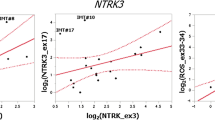
Targeted next-generation sequencing for cancer, as employed in this study, is a powerful tool for simultaneous detection of multiple tumor-related genomic alterations. By including intronic sequence coverage, this type of assay can detect more gene fusions than may be found by querying exonic sequence alone. However, because a targeted assay is designed to detect previously described alterations, it has less utility as a discovery tool than unbiased approaches such as whole-genome sequencing. The assay used in this study includes intronic coverage of ROS1 introns 30–34, where the majority of described breakpoints within ROS1-rearranged tumors have been described. Breakpoints in ROS1 occurring outside of this commonly affected region may not be detected using this assay, which may explain the lack of a detectable rearrangement in one ROS1-expressing tumor in this study.
Similar to the situation in lung adenocarcinomas,28 the identification of ROS1 rearrangements in inflammatory myofibroblastic tumors provides a new potential therapeutic option for some patients with aggressive variants of this tumor type. Although only small numbers of patients have been treated thus far, targeted therapy using crizotinib has shown clinical benefit for patients with metastatic and locally advanced ALK-positive inflammatory myofibroblastic tumors.13, 14 Similarly, a member of our group recently reported that crizotinib provided symptomatic improvement, as well as radiologic response, in a patient with advanced ROS1-rearranged inflammatory myofibroblastic tumor.18
In summary, expression of ROS1 correlates with ROS1 gene rearrangement in inflammatory myofibroblastic tumors. These findings suggest that immunohistochemistry for ROS1 may be useful to support the diagnosis of a subset of inflammatory myofibroblastic tumors and may select some clinically aggressive cases for targeted therapy directed against ROS1.
References
Gleason BC, Hornick JL . Inflammatory myofibroblastic tumours: where are we now? J Clin Pathol 2008;61:428–437.
Fletcher CDM, Bridge JA, Hogendoorn PCW, Mertens F (eds). WHO Classification of Tumours of Soft Tissue and Bone. International Agency for Research on Cancer: Lyon, 2013.
Coffin CM, Hornick JL, Fletcher CD . Inflammatory myofibroblastic tumor: comparison of clinicopathologic, histologic, and immunohistochemical features including ALK expression in atypical and aggressive cases. Am J Surg Pathol 2007;31:509–520.
Li J, Yin WH, Takeuchi K et al. Inflammatory myofibroblastic tumor with RANBP2 and ALK gene rearrangement: a report of two cases and literature review. Diagn Pathol 2013;8:147.
Marino-Enriquez A, Wang WL, Roy A et al. Epithelioid inflammatory myofibroblastic sarcoma: An aggressive intra-abdominal variant of inflammatory myofibroblastic tumor with nuclear membrane or perinuclear ALK. Am J Surg Pathol 2011;35:135–144.
Coffin CM, Patel A, Perkins S et al. ALK1 and p80 expression and chromosomal rearrangements involving 2p23 in inflammatory myofibroblastic tumor. Mod Pathol 2001;14:569–576.
Debelenko LV, Arthur DC, Pack SD et al. Identification of CARS-ALK fusion in primary and metastatic lesions of an inflammatory myofibroblastic tumor. Lab Invest 2003;83:1255–1265.
Debiec-Rychter M, Marynen P, Hagemeijer A et al. ALK-ATIC fusion in urinary bladder inflammatory myofibroblastic tumor. Genes Chromosomes Cancer 2003;38:187–190.
Griffin CA, Hawkins AL, Dvorak C et al. Recurrent involvement of 2p23 in inflammatory myofibroblastic tumors. Cancer Res 1999;59:2776–2780.
Lawrence B, Perez-Atayde A, Hibbard MK et al. TPM3-ALK and TPM4-ALK oncogenes in inflammatory myofibroblastic tumors. Am J Pathol 2000;157:377–384.
Panagopoulos I, Nilsson T, Domanski HA et al. Fusion of the SEC31L1 and ALK genes in an inflammatory myofibroblastic tumor. Int J Cancer 2006;118:1181–1186.
Patel AS, Murphy KM, Hawkins AL et al. RANBP2 and CLTC are involved in ALK rearrangements in inflammatory myofibroblastic tumors. Cancer Genet Cytogenet 2007;176:107–114.
Butrynski JE, D'Adamo DR, Hornick JL et al. Crizotinib in ALK-rearranged inflammatory myofibroblastic tumor. N Engl J Med 2010;363:1727–1733.
Mosse YP, Lim MS, Voss SD et al. Safety and activity of crizotinib for paediatric patients with refractory solid tumours or anaplastic large-cell lymphoma: a Children's Oncology Group phase 1 consortium study. Lancet Oncol 2013;14:472–480.
Cessna MH, Zhou H, Sanger WG et al. Expression of ALK1 and p80 in inflammatory myofibroblastic tumor and its mesenchymal mimics: a study of 135 cases. Mod Pathol 2002;15:931–938.
Selinger CI, Rogers TM, Russell PA et al. Testing for ALK rearrangement in lung adenocarcinoma: a multicenter comparison of immunohistochemistry and fluorescent in situ hybridization. Mod Pathol 2013;26:1545–1553.
Sholl LM, Weremowicz S, Gray SW et al. Combined use of ALK immunohistochemistry and FISH for optimal detection of ALK-rearranged lung adenocarcinomas. J Thorac Oncol 2013;8:322–328.
Lovly CM, Gupta A, Lipson D et al. Inflammatory myofibroblastic tumors harbor multiple potentially actionable kinase fusions. Cancer Discov 2014;4:889–895.
Cha YJ, Lee JS, Kim HR et al. Screening of ROS1 rearrangements in lung adenocarcinoma by immunohistochemistry and comparison with ALK rearrangements. PLoS One 2014;9:e103333.
Sholl LM, Sun H, Butaney M et al. ROS1 immunohistochemistry for detection of ROS1-rearranged lung adenocarcinomas. Am J Surg Pathol 2013;37:1441–1449.
Yoshida A, Tsuta K, Wakai S et al. Immunohistochemical detection of ROS1 is useful for identifying ROS1 rearrangements in lung cancers. Mod Pathol 2014;27:711–720.
Morris SW, Kirstein MN, Valentine MB et al. Fusion of a kinase gene, ALK, to a nucleolar protein gene, NPM, in non-Hodgkin's lymphoma. Science 1994;263:1281–1284.
Soda M, Choi YL, Enomoto M et al. Identification of the transforming EML4-ALK fusion gene in non-small-cell lung cancer. Nature 2007;448:561–566.
Marino-Enriquez A, Ou WB, Weldon CB et al. ALK rearrangement in sickle cell trait-associated renal medullary carcinoma. Genes Chromosomes Cancer 2011;50:146–153.
Smith NE, Deyrup AT, Marino-Enriquez A et al. VCL-ALK renal cell carcinoma in children with sickle-cell trait: the eighth sickle-cell nephropathy? Am J Surg Pathol 2014;38:858–863.
Wiesner T, He J, Yelensky R et al. Kinase fusions are frequent in Spitz tumours and spitzoid melanomas. Nat Commun 2014;5:3116.
Bergethon K, Shaw AT, Ou SH et al. ROS1 rearrangements define a unique molecular class of lung cancers. J Clin Oncol 2012;30:863–870.
Davies KD, Doebele RC . Molecular pathways: ROS1 fusion proteins in cancer. Clin Cancer Res 2013;19:4040–4045.
Takeuchi K, Soda M, Togashi Y et al. RET, ROS1 and ALK fusions in lung cancer. Nat Med 2012;18:378–381.
Fisher C . The diversity of soft tissue tumours with EWSR1 gene rearrangements: a review. Histopathology 2014;64:134–150.
Riggi N, Cironi L, Suva ML et al. Sarcomas: genetics, signalling, and cellular origins. Part 1: The fellowship of TET. J Pathol 2007;213:4–20.
Miettinen M, Wang ZF, Paetau A et al. ERG transcription factor as an immunohistochemical marker for vascular endothelial tumors and prostatic carcinoma. Am J Surg Pathol 2011;35:432–441.
Shah RB . Clinical applications of novel ERG immunohistochemistry in prostate cancer diagnosis and management. Adv Anat Pathol 2013;20:117–124.
Wang WL, Patel NR, Caragea M et al. Expression of ERG, an Ets family transcription factor, identifies ERG-rearranged Ewing sarcoma. Mod Pathol 2012;25:1378–1383.
Barnoud R, Sabourin JC, Pasquier D et al. Immunohistochemical expression of WT1 by desmoplastic small round cell tumor: a comparative study with other small round cell tumors. Am J Surg Pathol 2000;24:830–836.
Argani P, Lal P, Hutchinson B et al. Aberrant nuclear immunoreactivity for TFE3 in neoplasms with TFE3 gene fusions: a sensitive and specific immunohistochemical assay. Am J Surg Pathol 2003;27:750–761.
Doyle LA, Vivero M, Fletcher CD et al. Nuclear expression of STAT6 distinguishes solitary fibrous tumor from histologic mimics. Mod Pathol 2014;27:390–395.
Schweizer L, Koelsche C, Sahm F et al. Meningeal hemangiopericytoma and solitary fibrous tumors carry the NAB2-STAT6 fusion and can be diagnosed by nuclear expression of STAT6 protein. Acta Neuropathol 2013;125:651–658.
Acknowledgements
We thank Dr Jiang Wang (Cincinnati, OH) for providing a gross photograph of one of the ROS1-positive tumors. CML was supported by an American Cancer Society Institutional Research Grant (#IRG-58-009-54) as well as a for The Linds (FTL) Foundation Pilot Project award.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
This work was presented in part at the Annual Meeting of the United States and Canadian Academy of Pathology, San Diego, CA, 1–7 March 2014.
Rights and permissions
About this article
Cite this article
Hornick, J., Sholl, L., Dal Cin, P. et al. Expression of ROS1 predicts ROS1 gene rearrangement in inflammatory myofibroblastic tumors. Mod Pathol 28, 732–739 (2015). https://doi.org/10.1038/modpathol.2014.165
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2014.165
This article is cited by
-
Update from the 5th Edition of the World Health Organization Classification of Head and Neck Tumors: Soft Tissue Tumors
Head and Neck Pathology (2022)
-
Pseudosarcomatous myofibroblastic proliferations of the urinary bladder are neoplasms characterized by recurrent FN1–ALK fusions
Modern Pathology (2021)
-
The Most Common Mistake in Laryngeal Pathology and How to Avoid it
Head and Neck Pathology (2021)
-
Infantile inflammatory myofibroblastic tumors: clinicopathological and molecular characterization of 12 cases
Modern Pathology (2020)
-
Pulmonary inflammatory Myofibroblastic tumor indistinguishable from tuberculosis: a case report in a five-year-old child with hemoptysis
Journal of Cardiothoracic Surgery (2017)