Abstract
We have described a rare group of prostate adenocarcinomas that show aberrant expression of p63, a protein strongly expressed in prostatic basal cells and absent from usual-type acinar prostate cancers. The partial basal-like immunophenotype of these tumors is intriguing in light of the persistent debate surrounding the cell-of-origin for prostate cancer; however, their molecular phenotype is unknown. We collected 37 of these tumors on radical prostatectomy and biopsy and assessed subsets for a diverse panel of molecular markers. The majority of p63-expressing tumors were positive for the ΔNp63 isoform (6/7) by immunofluorescence and p63 mRNA (7/8) by chromogenic in situ hybridization. Despite p63 positivity, these tumors uniformly expressed luminal-type cytokeratin proteins such as CK18 (13/13), CK8 (8/8), and markers of androgen axis signaling commonly seen in luminal cells, including androgen receptor (10/11), NKX3.1 (8/8), and prostein (12/13). Conversely, basal cytokeratins such as CK14 and CK15 were negative in all cases (0/8) and CK5/6 was weakly and focally positive in 36% (4/11) of cases. Pluripotency markers including β-catenin, Oct4, and c-kit were negative in p63-expressing tumors (0/11). Despite nearly universal expression of androgen receptor and downstream androgen signaling targets, p63-expressing tumors lacked ERG rearrangements by fluorescence in situ hybridization (0/14) and ERG protein expression (0/37). No tumors expressed SPINK1 or showed PTEN protein loss (0/19). Surprisingly, 74% (14/19) of p63-expressing tumors expressed GSTP1 protein at least focally, and 33% (2/6) entirely lacked GSTP1 CpG island hypermethylation by bisulfite sequencing. In contrast to usual prostatic adenocarcinomas, prostate tumors with p63 expression show a mixed luminal/basal immunophenotype, uniformly lack ERG gene rearrangement, and frequently express GSTP1. These data strongly suggest that p63-expressing prostate tumors represent a molecularly distinct subclass and further study of this rare tumor type may yield important insights into the role of p63 in prostatic biology and the prostate cancer cell-of-origin.
Similar content being viewed by others
Main
We have recently described a group of prostatic adenocarcinomas that show aberrant expression of p63 protein.1, 2 In contrast to usual-type prostatic adenocarcinomas, which lack p63 expression, these unusual tumors express this benign basal cell marker in a non-basal cell distribution. p63-expressing tumors are exceedingly rare, accounting for less than 1% of all tumors in a busy consultation practice, and they coexist with usual-type prostatic adenocarcinomas in more than 80% of cases.1, 2 In some cases, p63-expressing tumors show a distinctive morphology, consisting predominantly of glands, nests, and cords with atrophic cytoplasm, hyperchromatic nuclei, and prominent nucleoli, often with a ‘basaloid’ appearance. However, in contrast to rare basal cell carcinomas of the prostate gland, these tumors express PSA and show morphologic characteristics of prostatic secretory cells.1, 3 Additionally, although p63-expressing tumors are positive for some basal cell markers typically absent in usual-type adenocarcinoma (such as BCL-2), they are negative for others, such as high molecular weight cytokeratin.1 Importantly, on limited needle biopsy specimens, the characteristic morphologic features of p63-expressing tumors may be difficult to appreciate and many cases are recognized only after immunostaining. In a recent series of radical prostatectomies, p63-expressing tumors were uniformly organ-confined, with a low proliferation rate despite showing a range of Gleason scores, raising the question of whether the traditional Gleason grading system is applicable to this rare tumor subtype.2
Taken together, these data suggest that p63-expressing prostatic carcinomas may be molecularly distinct from usual-type prostatic adenocarcinomas. To address this possibility, we collected the largest series of p63-expressing prostate carcinomas to date and systematically characterized them for expression of p63 isoforms, a large panel of luminal, basal, pluripotency, and androgen signaling markers, as well as for common molecular changes seen in usual-type prostate carcinomas, including ERG gene rearrangements, SPINK1 expression, PTEN loss, and GSTP1 methylation.
Materials and methods
Tissue Selection
A total of 37 p63-positive carcinomas were collected from the consultation files of the Johns Hopkins Pathology department from 2008 to 2012. Because of the limited tumor tissue availability in many cases, only a subset of cases were used for most expression studies described below, with the exception of ERG immunohistochemistry, where all tumors were assessed. The tumor samples were taken from radical prostatectomies (n=23), prostate needle biopsies (n=13), and one transurethral resection of the prostate. Clinicopathologic parameters of 21/23 radical prostatectomy cases were previously published.2 In 13 radical prostatectomy cases, a separate concurrent nodule of usual-type (p63-negative) acinar carcinoma was also present; however, tissue from these tumors was only available for additional studies in six cases. Tissue microarrays were manually constructed from 19 radical prostatectomy cases. In each case, a minimum of three 1.0 mm cores were punched from the p63-positive component, the usual-type acinar carcinoma component (when present), and surrounding benign prostatic tissue.
Fluorescence In Situ Hybridization (FISH)
ERG FISH to detect gene rearrangement was performed in the two TMAs constructed from a subset of 19 of the radical prostatectomy specimens using the Cymogen® 21q22 del-TECT Four Color probes (Cymogen Dx, New Windsor, NY, USA). Probe A labeled in CYMO-RED is located slightly centromeric to the ERG gene, probe B labeled in CYMO-ORANGE is slightly telomeric to the ERG gene and spans the HMGN1 gene, probe C labeled in CYMO-AQUA covers most of the DSCAM gene, and probe D labeled in CYMO-GREEN is located telomeric to the TMPRSS2 gene. In brief, a paraffin section of 5-μm thickness was baked overnight at 60 °C then de-waxed and rehydrated using xylene and graded ethanol, respectively. The slides were then incubated in 0.2 N HCL at room temperature for 10 min followed by 30 min citrate buffer (Vector Laboratories, Burlingame, CA, USA) incubation at 80 °C. After washes in 2 × SSC (Life Technologies, Carlsbad, CA, USA) and dH2O, slides were incubated in 0.01 N HCL at 37 °C for 10 min. The 21q22 del-TECT Four Color probes were added to the slides after the wash and dehydration steps and the Thermobrite Statspin (Thermo Fisher, Waltham, MA, USA) was used for overnight hybridization. Prolong® Gold anti-fade reagent with DAPI (Life Technologies) was then added to counter stain the target area before applying coverslip. These cases were scored for the presence of ERG gene rearrangement through deletion or translocation as recommended by the manufacturer. A subset of eight cases were also evaluated using a previously described custom made break apart ERG probe set, with identical results.4
Immunohistochemistry
Immunostains for NKX3.1, AR, ERG, CK14, CK15, SPINK1, PTEN, and GSTP1 were performed manually. Tissue sections were deparaffinized, rehydrated, and briefly equilibrated in water. Antigen unmasking was performed by steaming in citrate buffer (pH 6.0) for 25 min (for NKX3.1, CK15, SPINK), HTTR (Target Retrieval Solution; DAKO, Glostrup, Denmark) for 45–50 min (for AR, ERG, CK14, GSTpi), or EDTA steam for 45 min (for PTEN). Endogenous peroxidase activity was quenched by incubation with peroxidase block (DAKO) for 5 min at room temperature. Non-specific binding was blocked by incubating in 1% bovine serum albumin in Tris-HCl pH 7.5 for 20 min at room temperature. Slides were incubated with rabbit polyclonal anti-human Nkx3.1 antibody5 ( 1:1000 dilution), a rabbit monoclonal anti-human ERG antibody (Epitomics, Burlingame, CA, USA; 1:25 dilution), a rabbit polyclonal anti-human AR antibody (Santa Cruz Biotechnology, Santa Cruz, CA, USA; 1:1000 dilution), a rabbit polyclonal ΔNp63 (Biolegend, San Diego, CA, USA; 1:50), a mouse monoclonal CK14 antibody (Millipore, Billerica, MA, USA; 1:50 dilution), a rabbit monoclonal PTEN antibody (Cell Signaling, Danvers, MA, USA; 1:50 dilution), a monoclonal mouse GSTP1 antibody (Cell Signaling; 1:1000 dilution), a mouse monoclonal SPINK antibody (Abcam, Cambridge, UK; 1:150 dilution) for 45 min at room temperature, and a mouse monoclonal for CK15 (Thermo Scientific, Waltham, MA, USA; 1:600 dilution). A horseradish peroxidase-labeled polymer (PowerVision, Leica Microsystems, Bannockburn, IL, USA) was applied for 30 min at room temperature. Signal detection was performed using 3,3'-diaminobenzidine tetrahydrochloride (DAB) as the chromogen. Slides were counterstained with hematoxylin, dehydrated, and mounted.
Immunostains for p63 (Biocare Medical, Concord, CA, USA; clone 4A4), CK18 (DAKO; clone DC10), prostein (DAKO; clone 10E3), CK5/6 (Ventana,Tucson, AZ, USA; cloneD5/16 B4), β-catenin (BD Biosciences, Franklin Lakes, NJ, USA), OCT4 (Abcam; polyclonal), and c-KIT (Cellmarque, Rocklin, CA, USA) were performed on the Ventana Benchmark XT autostainer.
Immunofluorescence
Following deparaffinization and rehydration, antigen unmasking was performed by using HTTR buffer (Target Retrieval Solution; DAKO) for 45 min. Tissue sections were then blocked by protein blocking solution for 1 h at room temperature. Slides were incubated with rabbit polyclonal anti-human ΔNp63 (BioLegend; 1:50 dilution) and rat anti-human CK8 (Developmental Studies Hybridoma Bank, University of Iowa; 1: 100 dilution) for 45 min at room temperature. An anti-rabbit Alexa-fluor 488 fluorochrome-conjugated secondary antibody mixed with anti-rat secondary Alexa-fluor 594 antibody (Jackson Lab, Bar Harbor, ME, USA; 1:200 dilution) and was added to the slides for 1 h incubation. For ΔNp63 isoform staining, mouse ovarian tissue was used as a negative control as it has been previously shown to express the TAp63 isoform exclusively.6 For CK14 staining, FITC-conjugated monoclonal mouse anti-human CK14 antibody (Chemicon International, Billerica, MA, USA; 1:50 dilution) was applied to slides for 45 min at room temperature. All slides were mounted with Prolong® Gold anti-fade reagent with DAPI (Life Technologies) and cured overnight at room temperature.
Chromogenic In Situ Hybridization (CISH)
p63 mRNA expression in one of the TMAs (n=8 cases) was evaluated by using CISH assay (RNAscope® 2.0 FFPE Assay, Advanced Cell Diagnostics, Hayward, CA, USA). In brief, tissue sections were first deparaffinized and air-dried at room temperature. Per the manufacturer’s directions, a series of pretreatments was carried out as follows: Pretreat 1 (incubation at room temperature for 10 min), pre-boiled Pretreat 2 (incubation for 15 min), Pretreat 3 (incubation at 40 °C for 30 min). Target probes were then added to the tissue sections and hybridized at 40 °C for 2 h. Six steps of amplification hybridizations with various conditions were performed and colorimetric reaction (1:1 DAB mixture with equal volume of Brown-A and Brown-B, 10 min at room temperature) was used for detection. Gill’s hematoxylin was then used for counterstaining and dehydration/mounting procedures were performed.
Interpretation of Immunohistochemistry, Immunofluorescence, and CISH
Immunostains were semi-quantitatively scored in p63-positive tumor regions and separately scored in the associated usual adenocarcinoma component (where available) by a urologic pathologist (TLL). ΔNp63, ERG, AR, NKX3.1, OCT4, and β-catenin were scored as positive if >5% of cells showed nuclear staining (1 to 3+). All other immunostains were scored for cytoplasmic/membranous positivity and were considered positive if >5% of cells showed 1 to 3+ cytoplasmic/membranous staining. For cases on TMA, a case was considered positive for each marker if any of the spots from that case were positive. For p63 CISH, a case was considered positive if >30% of tumor cells showed>1 dot per cell.7
Bisulfite Sequencing
Bisulfite sequencing was carried out as described previously.8 In brief, genomic DNA was bisulfite converted using the EZ DNA methylation kit (Zymo Research, Orange, CA, USA) and amplified using primers specific to the CpG of GSTP1 (F-5′-TAGGGGTTGGGGTTGTAGTTTATA-3′, R-5′-AAAAACAAAAAAACAAATTCCTCC-3′, hg19: chr11:67 351 672–67 351 850). All PCR reactions were carried out in 40 μl volume containing 20 ng of bisulfite converted DNA, 1 × Platinum Taq buffer (Invitrogen, Carlsbad, CA, USA), 1.5 U Platinum Taq (Life Technologies), 250 μM each dNTPs, 1.5 mM MgCl2, 0.25 μg/μl bovine serum albumin, 2 μl dimethyl sulfoxide, 400 nM forward primer, and 400 nM reverse primer. Cycling conditions were 95 °C for 3 min, 38 cycles of 95 °C for 30 s, 55 °C for 30 s, and 72 °C for 30 s, followed by a 7 min extension step at 72 °C. PCR products were gel purified after electrophoresis on a 2% agarose gel, sub-cloned into pCR®2.1-TOPO® vector (Invitrogen) and analyzed by Sanger sequencing.
Results
p63 Expression
To verify that aberrant p63 expression is not due to cross-reaction of the clinically utilized p63 antibody (clone 4A4) with another nuclear antigen present in a subset of prostate cancers, we assessed p63 mRNA expression by CISH as well as p63 protein expression with an independent, isoform-specific antibody clone. The TP63 gene is comprised of 15 exons and codes for six different mRNA isoforms that share a common core DNA binding domain. Two independent promoters produce two N-terminal variants, called TA and ΔN, and alternative splicing at the 3' end generates three different C-terminal variants, termed α, β, and γ. Using a probe to the common core mRNA sequence, we found p63 mRNA expression by CISH in 87% (7/8) p63-expressing carcinomas, at levels generally equivalent to, or higher than, those seen in benign prostatic basal cells (Figure 1a). In contrast, co-existing usual-type acinar carcinomas and benign luminal cells (which are negative for p63 protein) did not express p63 mRNA using this assay (Figure 1a).
p63-expressing prostate cancers are positive for p63 mRNA by chromogenic in situ hybridization (CISH) and the ΔNp63 isoform by immunohistochemistry. (a) p63-expressing prostate carcinoma (top row) expresses p63 protein in a non-basal cell distribution (using the 4A4 antibody which detects both the ΔNp63 and TAp63 isoforms, × 400 magnification) as well as p63 mRNA by CISH (arrowheads), at levels similar to or higher than surrounding benign basal cells (arrow, × 630 magnification). The concurrent usual-type adenocarcinoma in this case (bottom row) does not express p63 protein or mRNA; however, nearby benign basal cells are positive for both (arrows). (b) Dual ΔNp63 and CK8 staining in prostatic tissues. These markers are expressed in separate compartments in benign prostatic tissue, with basal cells expressing ΔNp63 and luminal cells expressing CK8 (left panel). In p63-expressing tumors, these two markers are expressed in the same cells (middle panel). In contrast, usual-type acinar carcinomas express CK8 and are negative for ΔNp63. All images at × 400 magnification.
Previous RT-PCR studies in macrodissected or microdissected benign and malignant human prostate tissue have established that the TA, ΔN, and α, β, and γ mRNA variants are all expressed, with benign tissues showing higher levels than tumor tissues presumably because of the presence of basal cells.9 In prostate tissues, the ΔN and α variants are expressed at several-fold higher levels than the other variants, suggesting that ΔNp63α is the most common variant.9 Because the 4A4 p63 antibody detects both the TA and ΔN variants of p63, we examined whether an antibody specific to the ΔN isoform of p63 would label p63-expressing prostate tumors. Overall, 86% (6/7) of p63-expressing tumors were positive for the ΔNp63 isoform by immunofluorescence (Figure 1b). These results are consistent with recent data showing ΔNp63 isoform positivity in 96% (23/24) of p63-expressing tumors using the p40 antibody (a clinically tested antibody specific for the ΔNp63 isoform) (manuscript in process, TLL, JIE, and Peter B Illei,.10 Thus, we conclude that p63 expression in a small group of prostate tumors is not an artifact of non-specific antibody binding and these tumors express the ΔNp63 isoform similar to the benign prostate.
Luminal and Basal Cytokeratin Expression
Previous studies have demonstrated that ΔNp63 binds to conserved enhancer elements in the CK5 and CK14 promoter region and thereby directs expression of these genes in the skin.11 Despite p63 positivity, aberrantly p63-expressing tumors lack high molecular weight keratin cocktail expression (34βE12), which detects CK1, 5, 10, and 14.1 To more extensively survey luminal and basal cell marker expression in these rare tumors, we queried their expression of individual high and low molecular weight cytokeratins. We examined CK5/6 expression by immunohistochemistry in these tumors and found it to be weakly and focally positive in 36% (4/11) of cases (Figure 2). Because 34βE12 fails to mark these tumors, the limited reactivity to CK5/6 may represent CK6 expression. Similarly, CK14 (by immunofluorescence) and CK15 (by immunohistochemistry) were entirely negative in all cases, despite robust expression by benign basal cells (0/8) (data not shown).
p63-expressing prostate tumors are generally positive for low molecular weight cytokeratins and negative for high molecular weight cytokeratins. Most p63-expressing tumors diffusely and strongly express CK18 at levels similar to benign luminal cells (case 1 and 3), although some show weaker diffuse expression (case 2). Similarly, most are entirely negative for CK5/6 (cases 2 and 3), although a minority weakly and focally express CK5/6 (case 1). All images are at × 200 magnification.
In contrast to predominantly negative high molecular weight cytokeratin staining, low molecular weight cytokeratin was diffusely positive (ranging from weak to strong in intensity) in all p63-expressing tumors by immunofluorescence for CK8 (Figure 1b, 8/8) and immunohistochemistry for CK18 (Figure 2, 13/13). Interestingly, double-labeling for ΔNp63 and CK8 demonstrated co-expression of ΔNp63 and CK8, in stark contrast to the mature benign prostate (where these markers are expressed in distinct cellular compartments) and to usual-type adenocarcinomas (which do not express ΔNp63 or p63 using the 4A4 antibody) (Figure 1b).
Pluripotency/Pluripotential Markers
To assess whether the partial basal-like phenotype of aberrantly p63-expressing tumors was associated with expression of other pluripotency markers, we examined the expression of nuclear β-catenin, nuclear Oct4, and cytoplasmic c-kit. All three pluripotency/pluripotential markers were entirely negative in all p63-expressing tumors (0/11 for each marker, data not shown).
Androgen Signaling Axis and Androgen-Induced Gene Rearrangements
Benign basal cells in the prostate only weakly express androgen receptor compared with the strong expression in benign luminal cells.12 Despite a partial basal-like phenotype, p63-expressing prostate tumors expressed nuclear androgen receptor at levels comparable with surrounding benign luminal cells (10/11) (Figure 3). Consistent with this, these tumors also expressed markers of prostatic differentiation induced by active androgen signaling, such as NKX3.1 (8/8) and prostein (12/13) by immunohistochemistry (Figure 3 and data not shown). Androgen signaling was recently shown to play an important etiologic role in gene rearrangements.13 Despite an apparently intact androgen signaling axis, p63-expressing tumors uniformly lacked ERG rearrangements by fluorescence in situ hybridization (0/14). In contrast, of the three concurrent usual-type acinar carcinomas that were successfully hybridized for TMPRSS2-ERG FISH, one harbored a TMPRSS2 rearrangement, wherein the green TMPRSS2 probe was split from the ERG and flanking probes at one allele (Figure 4). This case did not express ERG protein, consistent with a TMPRSS2 rearrangement involving other ETS-family transcription factors, such as ETV1, ETV4, and ETV5, which are not on chromosome 21 (as are TMPRSS2 and ERG) but have been involved in recurrent gene fusions with TMPRSS2 in prostate cancer.14 Notably, the p63-expressing tumor from the same case did not harbor this rearrangement by FISH (Figure 4). Consistent with the apparent lack of TMPRSS2-ERG gene rearrangements in p63-expressing prostate cancers, a larger cohort of p63-expressing tumors did not express ERG protein by immunohistochemistry (0/37), despite apparently intact androgen signaling based on AR, NKX3.1, and prostein staining. The lack of ERG protein expression in p63-expressing tumors was highly statistically significant compared with unselected cohorts of usual-type adenocarcinomas at our institution where 49.3% (534/1083) are positive for ERG protein (P=0.0001 by Fisher’s exact test).
p63-expressing prostate tumors express androgen receptor (AR) and prostatic differentiation markers (NKX3.1). Two representative cases showing AR levels in tumor cells similar to those in surrounding benign luminal cells ( × 200 magnification). NKX3.1, a prostatic differentiation marker and marker of downstream androgen signaling is also expressed in p63-positive tumors ( × 400 magnification).
p63-expressing prostate tumors do not rearrange the ERG gene locus or express ERG protein. ERG break-apart FISH assay in a usual-type carcinoma occurring concurrently with a p63-expressing tumor demonstrates rearrangement involving TMPRSS2 locus (green probe), which is spatially separated from the red and orange probes (centromeric and telomeric relative to the ERG gene, respectively) ( × 630 magnification). In contrast, the p63-expressing tumor from the same patient is negative for any rearrangement at this locus. Notably, neither tumor expresses ERG protein, despite positivity in endothelial cell nuclei that provide an internal positive control (arrowheads). Lack of ERG staining in the TMPRSS2-rearranged usual-type acinar carcinoma could be consistent with a rearrangement involving another ETS-family gene ( × 400 magnification).
Recent work to define molecular subclasses of usual-type prostatic adenocarcinoma has suggested that tumors lacking TMPRSS2-ETS family gene rearrangements may be relatively enriched for other characteristic molecular changes, such as SPINK1 overexpression.15 To determine whether p63-expressing tumors also express SPINK1, we utilized a genetically validated immunohistochemistry assay to assess SPINK1 protein expression.15 None of the p63-expressing tumors expressed SPINK1 protein (0/19, data not shown), nor did the concurrent usual-type carcinomas associated with these cases (0/6). Similarly, recent work has suggested that ERG-negative usual-type adenocarcinomas less commonly harbor deletions of PTEN, the most commonly lost tumor suppressor in prostate cancer.16, 17, 18, 19 Using a genetically validated assay to assess PTEN protein expression,5 we found that p63-expressing tumors did not show PTEN protein loss (0/19, data not shown), nor did the concurrent usual-type acinar carcinomas associated with these cases (0/6). Because low Gleason score tumors (such as Gleason 3+3=6) rarely demonstrate PTEN loss,5 the lack of PTEN loss in p63-expressing tumors is consistent with their relatively indolent behavior as well as with the absence of ERG rearrangement.
GSTP1 methylation
Glutathione S-transferase π (GSTP1) is expressed at high levels in the basal cell compartment of normal prostate epithelium. In close to 95% of usual-type prostatic adenocarcinomas, the CpG island around the promoter of GSTP1 shows dense cytosine methylation resulting in transcriptional silencing of the gene.20, 21 GSTP1 hypermethylation is the most frequently observed genomic alteration in prostate cancer and can therefore be considered a hallmark of usual-type adenocarcinoma. To determine whether GSTP1 expression is lost in p63-expressing tumors, we first utilized a genetically validated immunohistochemical assay for GSTP1 protein expression. Of the concurrent usual-type adenocarcinomas associated with p63-expressing tumors, all lacked GSTP1 protein expression by immunohistochemistry (6/6, Figure 5). In contrast, 74% (14/19) of p63-expressing tumors expressed GSTP1 protein at least focally.
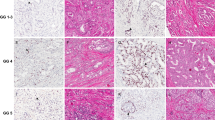
p63-expressing prostate tumors do not invariably silence GSTP1. (a) A representative p63-expressing tumor with high GSTP1 protein expression by immunohistochemistry and concomitant lack of hypermethylation at the CpG island of the GSTP1 gene by bisulfite sequencing (case A, bottom). (b) This p63-expressing tumor shows lack of GSTP1 expression, consistent with hypermethylation at the GSTP1 locus by bisulfite sequencing at levels comparable with usual-type adenocarcinoma controls (case B, bottom). (c) This p63-expressing tumor is admixed with usual-type p63-negative carcinoma and shows mixed GSTP1 protein expression. Bisulfite sequencing in this case shows hypermethylation of the GSTP1 locus at levels similar to usual-type adenocarcinomas (case C, bottom). Red bar indicates genomic location of the bisulfite sequencing amplicon; yellow bar indicates region of differential methylation between normal prostatic epithelium and usual-type adenocarcinoma; green bar indicates promoter associated CpG island. All images are at × 100 magnification.
To assess whether expression of GSTP1 protein in p63-expressing tumors was associated with a lack of promoter methylation, we surveyed GSTP1 hypermethylation in a subset of six cases using bisulfite sequencing. Of these cases, 66% (4/6) were positive for GSTP1 protein, one case showed mixed positive and negative GSTP1 staining in the p63-component and one case was negative for GSTP1 in the p63 component. Of the four cases positive for GSTP1 protein, three (75%) had a concurrent acinar component that was negative for the protein. The case with mixed GSTP1 staining in the p63 component was negative for GSTP1 in the concurrent acinar component. Bisulfite sequencing was used to assess the CpG island methylation status of GSTP1. Of the five p63-expressing cases with diffuse or partial expression of GSTP1, we found that 40% (2/5) entirely lacked GSTP1 methylation by bisulfite sequencing (Figure 5a). Similarly, the one p63-expressing case that lacked GSTP1 protein expression showed methylation of the gene similar to levels seen in usual-type adenocarcinoma (Figure 5b). Of the remaining three cases with discordant staining and methylation results, all showed positive GSTP1 staining with some degree of gene methylation. Of these cases, two could be explained by potential contamination by usual-type adenocarcinoma during macrodissection of the p63-expressing component (Figure 5c).
Discussion
Usual-type prostatic adenocarcinomas have a luminal-type immunoprofile, lacking basal markers such as p63 and high molecular weight cytokeratins, and diffusely expressing low molecular weight cytokeratins and markers of androgen axis signaling. Here, we have systematically characterized a rare group of prostatic adenocarcinomas that aberrantly express p63 and have demonstrated that they likely represent a molecularly distinct subclass of prostatic adenocarcinoma. Although these tumors are relatively rare and behave in an overwhelmingly indolent manner in limited follow-up studies, study of these tumors has the potential to expand our understanding of the function of p63 in prostatic biology and help to shed light on the persistent debate about the potential cell-of-origin of prostate cancer.
Excluding the possibility of non-specific p63 antibody reactivity in a subset of prostate tumors, we have demonstrated that these rare tumors are positive for p63 mRNA by CISH, and, much like the benign prostate, express the ΔNp63 protein. Interestingly, despite evidence that ΔNp63 may drive basal cytokeratin expression in other organs,11 p63-expressing prostate tumors show a predominantly luminal cytokeratin expression profile with only weak and focal CK5/6 positivity. Co-expression of p63 and luminal cytokeratins, such as CK8, is seen only in the embryonic urogenital sinus during early prostatic development, and is never seen in the mature prostate or in usual-type adenocarcinomas.22 Consistent with their luminal immunoprofile, p63-expressing tumors also express androgen receptor at levels comparable with benign luminal cells, as well as downstream targets of androgen receptor signaling (NKX3.1 and prostein). Yet, despite this luminal phenotype and apparently intact androgen-signaling axis, these tumors do not show TMPRSS2-ERG gene rearrangements, either by FISH or immunohistochemical staining (0/37). Given that ERG protein expression is seen in around 50% of usual-type adenocarcinomas,23 this is a highly significant result and strongly suggests that p63-expressing tumors are molecularly distinct from usual prostatic adenocarcinomas. The fact that p63-positive, ERG-rearrangement-negative tumors can be found adjacent to usual-type adenocarcinoma with ERG rearrangements suggests that p63-positive tumors are likely clonally distinct from the adjacent usual-type adenocarcinoma.
Furthermore, the differences in GSTP1 methylation status between usual-type adenocarcinoma and p63-positive tumors might indicate a more global difference in the epigenetic makeup of these tumors, which could explain many of the observed phenotypic differences.
As a master regulator of stratified epithelial development, there has been much interest in the role of p63 in organogenesis. Mice lacking both copies of the TP63 gene are born with severe defects in epithelial, craniofacial, and limb development, and die soon after birth because of lack of epidermal stratification and barrier function.24, 25 Importantly, these mice lack a developing prostate; however, they do not live long enough to allow for complete in situ development of this organ.26 If the TP63 −/− urogenital sinus is rescued by removal from the early embryo and grafted under the renal capsule of an adult male mouse, the resulting tissue lacks basal cells and differentiates towards the intestinal lineage, suggesting that p63 is essential for the maintenance of basal cells and terminal differentiation of prostatic secretory cells.27 In the skin, recent mouse models with ΔNp63-specific knockout have demonstrated that this isoform is the most critical for stratified epithelial development, consistent with the higher levels of this isoform in the prostate and other epithelial tissues.28 Along these lines, recent lineage tracing experiments of ΔNp63-expressing cells in the embryonic prostate have shown that these cells can generate all lineages (basal, luminal, and neuroendocrine) during development.22 Because other lineage tracing experiments in adult mice have shown that the prostatic lineages may each be self-sustaining,29, 30 it may be that ΔNp63-expressing cells become unipotent during later development.22
Because it is the oldest member of the p53 family, there has also been much interest in the role of p63 in epithelial tumorigenesis, which remains controversial, possibly because of tissue-specific and isoform-specific functions of the gene. p63-null mice have a variable susceptibility to tumorigenesis and unlike TP53, TP63 is rarely mutated in human tumors.31 Mutations in the coding sequence of TP63 are extremely rare in usual-type prostatic adenocarcinomas in recent whole genome sequencing efforts (www.cBioportal.org), consistent with results from older Sanger sequencing studies.9 The fact that p63 protein is not detectable in the vast majority of prostatic carcinomas, despite some level of p63 mRNA detectable by RT-PCR,9 suggests that there may be obligate posttranscriptional silencing of the gene in prostate tumors. Indeed, emerging evidence in other systems suggests that micro-RNAs may play an important role in regulating the gene.32 If suppression of p63 protein is critical for prostatic tumorigenesis, it remains possible that p63-expressing adenocarcinomas may harbor inactivating missense or nonsense mutations in the gene that have not been captured in sequencing studies because of the relative rarity of this tumor type. Studies to address this possibility are ongoing; however, it is notable that 22RV1 cells, the only prostate cancer cell line that harbors a known detrimental mutation in TP63, do not express the protein by immunohistochemistry.9 Interestingly, the relative prevalence of high level amplifications of the TP63 gene in squamous carcinomas of the cervix, lung, and head and neck may be more consistent with an oncogenic function for this gene, at least in some tumor types.31 Thus, whether p63 expression may play a role in retarding or promoting tumorigenesis in this rare subset of prostate tumors remains unclear.
Finally, it is tempting to speculate that these rare p63-expressing tumors may add an important clue as to the cell-of-origin of prostatic adenocarcinoma. Despite the fact that most prostatic adenocarcinomas display a luminal cell phenotype, tissue regeneration studies using human tissue have suggested that prostatic basal cells remain competent to generate tumor cells expressing luminal markers,33, 34 and this appears to happen in some animal models as well.35, 36 Arguing against the basal cell-of-origin theory is the fact that GSTP1 promoter methylation and ERG gene rearrangements, characteristic of approximately 95% and 50% of usual-type adenocarcinomas, respectively, are found exclusively in luminal cells in a subset of human prostatic intraepithelial neoplasia, which is widely considered to be a precursor lesion for prostatic adenocarcinomas.37, 38 Similarly, other animal models support the role of a luminal population in tumor initiation.29 The existence of p63-expressing prostate tumors that lack ERG gene rearrangements may lend some credence to the theory that human prostate tumors may originate from diverse cell populations. Although speculative, it remains possible that p63-expressing tumors are molecularly distinct from usual adenocarcinomas in part because they originate from a rare basal tumor-initiating cell population, which has undergone partial, but incomplete, differentiation towards a luminal phenotype. Additional in depth molecular studies of these rare tumors may add important evidence to this ongoing debate.
References
Osunkoya AO, Hansel DE, Sun X et al. Aberrant diffuse expression of p63 in adenocarcinoma of the prostate on needle biopsy and radical prostatectomy: Report of 21 cases. Am J Surg Pathol 2008;32:461–467.
Giannico GA, Ross HM, Lotan T et al. Aberrant expression of p63 in adenocarcinoma of the prostate: A radical prostatectomy study. Am J Surg Pathol 2013;37:1401–1406.
Ali TZ, Epstein JI . Basal cell carcinoma of the prostate: A clinicopathologic study of 29 cases. Am J Surg Pathol 2007;31:697–705.
Toubaji A, Albadine R, Meeker AK et al. Increased gene copy number of ERG on chromosome 21 but not TMPRSS2-ERG fusion predicts outcome in prostatic adenocarcinomas. Mod Pathol 2011;24:1511–1520.
Lotan TL, Gurel B, Sutcliffe S et al. PTEN protein loss by immunostaining: Analytic validation and prognostic indicator for a high risk surgical cohort of prostate cancer patients. Clin Cancer Res 2011;17:6563–6573.
Suh EK, Yang A, Kettenbach A et al. P63 protects the female germ line during meiotic arrest. Nature 2006;444:624–628.
Wang F, Flanagan J, Su N et al. RNAscope: A novel in situ RNA analysis platform for formalin-fixed, paraffin-embedded tissues. J Mol Diagn 2012;14:22–29.
Yegnasubramanian S, Lin X, Haffner MC et al. Combination of methylated-DNA precipitation and methylation-sensitive restriction enzymes (COMPARE-MS) for the rapid, sensitive and quantitative detection of DNA methylation. Nucleic Acids Res 2006;34:e19.
Parsons JK, Saria EA, Nakayama M et al. Comprehensive mutational analysis and mRNA isoform quantification of TP63 in normal and neoplastic human prostate cells. Prostate 2009;69:559–569.
Uchida K, Ross H, Epstein J et al. DeltaNp63 isoforms of p63 in aberrant diffuse p63 positive prostate cancer. Mod Path 2012;25:247.
Romano RA, Ortt K, Birkaya B et al. An active role of the DeltaN isoform of p63 in regulating basal keratin genes K5 and K14 and directing epidermal cell fate. PLoS One 2009;4:e5623.
Harper ME, Glynne-Jones E, Goddard L et al. Expression of androgen receptor and growth factors in premalignant lesions of the prostate. J Pathol 1998;186:169–177.
Haffner MC, Aryee MJ, Toubaji A et al. Androgen-induced TOP2B-mediated double-strand breaks and prostate cancer gene rearrangements. Nat Genet 2010;42:668–675.
Rubin MA, Maher CA, Chinnaiyan AM . Common gene rearrangements in prostate cancer. J Clin Oncol 2011;29:3659–3668.
Tomlins SA, Rhodes DR, Yu J et al. The role of SPINK1 in ETS rearrangement-negative prostate cancers. Cancer Cell 2008;13:519–528.
Carver BS, Tran J, Gopalan A et al. Aberrant ERG expression cooperates with loss of PTEN to promote cancer progression in the prostate. Nat Genet 2009;41:619–624.
King JC, Xu J, Wongvipat J et al. Cooperativity of TMPRSS2-ERG with PI3-kinase pathway activation in prostate oncogenesis. Nat Genet 2009;41:524–526.
Han B, Mehra R, Lonigro RJ et al. Fluorescence in situ hybridization study shows association of PTEN deletion with ERG rearrangement during prostate cancer progression. Mod Pathol 2009;22:1083–1093.
Bismar TA, Yoshimoto M, Vollmer RT et al. PTEN genomic deletion is an early event associated with ERG gene rearrangements in prostate cancer. BJU Int 2011;107:477–485.
Lee WH, Morton RA, Epstein JI et al. Cytidine methylation of regulatory sequences near the pi-class glutathione S-transferase gene accompanies human prostatic carcinogenesis. Proc Natl Acad Sci USA 1994;91:11733–11737.
Yegnasubramanian S, Kowalski J, Gonzalgo ML et al. Hypermethylation of CpG islands in primary and metastatic human prostate cancer. Cancer Res 2004;64:1975–1986.
Pignon JC, Grisanzio C, Geng Y et al. P63-expressing cells are the stem cells of developing prostate, bladder, and colorectal epithelia. Proc Natl Acad Sci USA 2013;110:8105–8110.
Pettersson A, Graff RE, Bauer SR et al. The TMPRSS2:ERG rearrangement, ERG expression, and prostate cancer outcomes: A cohort study and meta-analysis. Cancer Epidemiol Biomarkers Prev 2012;21:1497–1509.
Mills AA, Zheng B, Wang XJ et al. P63 is a P53 homologue required for limb and epidermal morphogenesis. Nature 1999;398:708–713.
Yang A, Schweitzer R, Sun D et al. P63 is essential for regenerative proliferation in limb, craniofacial and epithelial development. Nature 1999;398:714–718.
Signoretti S, Waltregny D, Dilks J et al. P63 is a prostate basal cell marker and is required for prostate development. Am J Pathol 2000;157:1769–1775.
Signoretti S, Pires MM, Lindauer M et al. P63 regulates commitment to the prostate cell lineage. Proc Natl Acad Sci USA 2005;102:11355–11360.
Romano RA, Smalley K, Magraw C et al. DeltaNp63 knockout mice reveal its indispensable role as a master regulator of epithelial development and differentiation. Development 2012;139:772–782.
Choi N, Zhang B, Zhang L et al. Adult murine prostate basal and luminal cells are self-sustained lineages that can both serve as targets for prostate cancer initiation. Cancer Cell 2012;21:253–265.
Wang ZA, Mitrofanova A, Bergren SK et al. Lineage analysis of basal epithelial cells reveals their unexpected plasticity and supports a cell-of-origin model for prostate cancer heterogeneity. Nat Cell Biol 2013;15:274–283.
Grisanzio C, Signoretti S . P63 in prostate biology and pathology. J Cell Biochem 2008;103:1354–1368.
Yi R, Poy MN, Stoffel M et al. A skin microRNA promotes differentiation by repressing 'stemness'. Nature 2008;452:225–229.
Lawson DA, Zong Y, Memarzadeh S et al. Basal epithelial stem cells are efficient targets for prostate cancer initiation. Proc Natl Acad Sci USA 2010;107:2610–2615.
Goldstein AS, Huang J, Guo C et al. Identification of a cell of origin for human prostate cancer. Science 2010;329:568–571.
Stoyanova T, Cooper AR, Drake JM et al. Prostate cancer originating in basal cells progresses to adenocarcinoma propagated by luminal-like cells. Proc Natl Acad Sci USA 2013;110:20111–20116.
Lu TL, Huang YF, You LR et al. Conditionally ablated pten in prostate basal cells promotes basal-to-luminal differentiation and causes invasive prostate cancer in mice. Am J Pathol 2013;182:975–991.
Nakayama M, Bennett CJ, Hicks JL et al. Hypermethylation of the human glutathione S-transferase-pi gene (GSTP1) CpG island is present in a subset of proliferative inflammatory atrophy lesions but not in normal or hyperplastic epithelium of the prostate: A detailed study using laser-capture microdissection. Am J Pathol 2003;163:923–933.
Mosquera JM, Perner S, Genega EM et al. Characterization of TMPRSS2-ERG fusion high-grade prostatic intraepithelial neoplasia and potential clinical implications. Clin Cancer Res 2008;14:3380–3385.
Acknowledgements
Funding for this research was provided in part by a Prostate Cancer Foundation Young Investigator Award (TLL), a David H. Koch Research Award (TLL), and the NIH/NCI Prostate SPORE P50CA58236.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Tan, HL., Haffner, M., Esopi, D. et al. Prostate adenocarcinomas aberrantly expressing p63 are molecularly distinct from usual-type prostatic adenocarcinomas. Mod Pathol 28, 446–456 (2015). https://doi.org/10.1038/modpathol.2014.115
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2014.115
This article is cited by
-
TAp63 and ΔNp63 (p40) in prostate adenocarcinomas: ΔNp63 associates with a basal-like cancer stem cell population but not with metastasis
Virchows Archiv (2021)
-
Integrated immunohistochemical and molecular analysis improves diagnosis of high-grade carcinoma in the urinary bladder of patients with prior radiation therapy for prostate cancer
Modern Pathology (2020)