Abstract
Antimicrobial resistance to clarithromycin is a growing concern in the treatment of Helicobacter pylori and is associated with three major point mutations of the 23S rRNA, A2142C, A2142G, and A2143G. The use of traditional culture-based methods for determination of clarithromycin resistance in H. pylori are time consuming and lack sensitivity. We implemented a real-time PCR with melt curve analysis to detect and characterize H. pylori in formalin-fixed, paraffin-embedded gastric biopsy specimens to assess the frequency of clarithromycin resistance mutations in our study population. One hundred and fifty-three formalin-fixed, paraffin-embedded gastric biopsies were chosen on the basis of positive immunohistochemical staining for H. pylori and an accompanying histopathological diagnosis of Helicobacter-associated gastritis. New adjacent sections were taken for immunohistochemical staining and DNA extraction with subsequent testing by PCR assay and melt curve analysis using a primer and probe combination first described by Oleastro et al.12 One hundred and forty-six samples demonstrated adequate amplification of a human DNA control target. Of these, there were 122 H. pylori immunohistochemistry-positive samples. In all, 103 out of 122 (84%) immunohistochemistry-positive samples demonstrated amplifiable H. pylori 23S rRNA gene target and 19 (16%) demonstrated no amplification of H. pylori. Twenty-two samples were negative for H. pylori by immunohistochemistry and PCR. Two were negative for H. pylori by immunohistochemistry, but were positive for H. pylori by PCR. In all, 52 out of 105 (50%) PCR-positive samples demonstrated resistance mutations, and it was determined that a heterogeneous population of mutated and unmutated organisms was present in 11 out of 52 samples. The use of PCR assays allows for a timely assessment of clarithromycin resistance status without the disadvantages of culture-based methods, and may lead to a decrease in treatment failure rates.
Similar content being viewed by others
Main
Helicobacter pylori is a Gram-negative bacillus associated with gastroesophageal reflux disease, gastric and duodenal ulcers, gastric cancer, and MALT lymphoma. H. pylori has a worldwide distribution, with a higher incidence of infection among the population in developing countries.1 It is estimated that up to 30–40% of the population of the United States harbors H. pylori within the upper gastrointestinal tract;2 however, the majority of people infected with H. pylori are asymptomatic. According to the American College of Gastroenterology, treatment is indicated for active peptic ulcer disease, a confirmed history of peptic ulcer disease that has not been previously treated for H. pylori, low-grade gastric MALT lymphoma, and following endoscopic resection of early gastric cancer. In addition, a ‘test and treat’ strategy has been recommended for uninvestigated dyspepsia in patients under 55 years of age who do not demonstrate specific ‘alarm’ features, such as anemia, vomiting, or family history of gastrointestinal malignancy.3
The ‘triple therapy’ regimen is the mainstay of pharmacological treatment for H. pylori infection in the United States, and typically consists of either amoxicillin or metronidazole used together with a proton pump inhibitor and clarithromycin.3 Although increased antimicrobial resistance has been seen with each of the agents used in treatment of H. pylori, the increase in treatment failure can be attributed in large part to resistance to clarithromycin.4, 5, 6 It has been previously demonstrated that point mutations in domain V of the 23S rRNA gene are responsible for the resistance of H. pylori to macrolides,7 with the three major point mutations being A2142G, A2143G, and A2142C.7 The most frequent of these are transitions of adenine to guanine at the 2143 or 2142 positions, which account for >80% of clarithromycin resistance seen worldwide and ∼90% of resistance seen in developed countries.8
A variety of diagnostic methods have been developed for H. pylori, including culture and histopathology, rapid urease tests, stool antigen testing, the urea breath test, serology, and more recently molecular assays.3, 9 Of these, culture is the only widely available method that also allows for antimicrobial susceptibility testing. In addition to requiring tissue obtained through endoscopy, culture has many inherent disadvantages, which include requirements of special transportation conditions with restrictive timelines to ensure optimal viability of organisms,10 the use of special media and environments for maintenance of cultures, and lengthy incubation time.11
Owing to the limitations of current laboratory methods available for the assessment of clarithromycin resistance in H. pylori, we set out to implement a method that would rapidly detect clarithromycin resistance and susceptible H. pylori without the constraints of traditional culture methods. In this report, we describe the use of real-time PCR based on methods previously described by Oleastro et al12 to detect H. pylori in formalin-fixed, paraffin-embedded gastric biopsy specimens, and to assess the frequency of clarithromycin resistance within our study population.
Materials and methods
Bacterial Culture and Sample Processing
Seven H. pylori reference strains with known mutations in the 23s rRNA gene (two A2142G and five A2143G strains) and one unmutated strain were obtained from the Centers for Disease Control and Prevention, National Campylobacter and Helicobacter Reference Laboratory. Samples were grown on Brucella agar w/5% sheep blood, Hemin, and Vitamin K (Remel, Lenexa, KS, USA) under a microaerophilic environment at 37 °C for up to 7 days. Phenotypic clarithromycin susceptibilities of the seven H. pylori strains with 23S mutations and the unmutated sample were determined using the E-test method (AB Biodisk, Solna, Sweden) with previously established guidelines of MIC ≥1 μg/ml indicating resistance.13
DNA was extracted by placing a heavy inoculum of bacteria into 250 μl of TritonX buffer solution (stock: 10 ml 2% Triton X-100, 5 ml TWEEN 20, 20 ml 0.5 M EDTA, 1 ml 1M Tris HCL (pH 8.0) and 964 ml molecular grade water) followed by placement in a heat block at 99 °C for 10 min.
Preparation of Fresh and Formalin-Fixed, Paraffin-Embedded Dilution Series
The limit of detection of the assay was evaluated using fresh isolates via a tenfold dilution series of extracted DNA from 107 to 101 genome equivalents per a final 20 μl PCR reaction volume. Genome equivalents were calculated on the basis of an H. pylori genome size of 1.64 megabases, resulting in 5.655 × 105 copies/ng purified DNA. A spectrophotometer (Nanodrop 1000, Thermo Fisher Scientific, Wilmington, DE, USA) was used to determine the concentration of DNA per μl of initial sample.
For determination of the limit of detection using formalin-fixed, paraffin-embedded specimens, mock biopsy blocks were created to serve as quantitative standards: a 0.5 McFarland dilution of the cultured strains of H. pylori was prepared and mixed with collagen at tenfold dilutions of 106–102 organisms per μl, to a final volume of 50 μl. The resulting collagen plug was formalin-fixed and paraffin-embedded according to our institutional protocol for gastric biopsy specimens. 10-μm-thick sections were prepared and mounted to glass slides, and the DNA was extracted from the sections using Pinpoint (Zymo Research, Orange, CA, USA) according to the manufacturer’s instructions.
Gastric Biopsy Selection and Processing
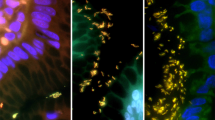
One hundred and fifty-three formalin-fixed, paraffin-embedded gastric biopsy specimens were obtained from the archives of three institutions within our hospital system with IRB approval. The gastric biopsies were each retrospectively reviewed and evaluated, and specimens were chosen on the basis of positive immunohistochemical staining for H. pylori and an accompanying histopathologic diagnosis of Helicobacter-associated gastritis. 10-μm-thick unstained sections were prepared from the gastric biopsies, and adjacent sections were submitted for immunohistochemical staining. The immunohistochemically stained glass slides were reviewed for the presence of H. pylori organisms with subsequent performance of DNA extraction from the corresponding unstained sections using Pinpoint (Zymo Research), according to the manufacturer’s instructions.
Detection of Point Mutations of the 23 rRNA Gene of H. pylori by Real-Time PCR
A 267-bp fragment of the 23S rRNA gene of H. pylori was amplified using primers HPYS and HPYA, which have been previously described.14 Amplification was detected using a 5′ LC-Red 640-labeled sensor probe with a 3′ phosphorylated end (5′-GGCAAGACGGAAAGACC-3′ nucleotides 2504–2520), and a 3′ fluorescein-labeled anchor probe (5′-TGTAGTGGAGGTGAAAATTCCTCCTACCC-3′ nucleotides 2473–2501). The sensor probe hybridized to the mutation site containing DNA region whereas the anchor probe hybridized three bases upstream, as previously described.12 The LightCycler 1.5 thermocycler (Roche Diagnostics, Indianapolis, IN, USA) was used for the PCR reactions. The glass capillaries contained a total volume of 20 μl (2 μl of DNA template, 4.0 μl of MgCl2 (25 mM), 0.5 μl of forward and reverse primers (20 μM each), 0.2 μl of sensor and anchor probes (20 μM each), 2.0 μl of FastStart DNA Master Hybridization Probes (Roche Diagnostics)). The PCR conditions consisted of an initial denaturation cycle of 95 °C for 10 min, followed by 50 amplification cycles with a temperature transition rate of 20 °C/s, with denaturation at 95 °C for 15 s, an annealing step at 57 °C for 30 s, and extension step at 72 °C for 30 s. Amplification was immediately followed by a melting program, which consisted of heating to 95 °C for 15 s, and cooling to 45 °C for 30 s with a temperature elevation rate of 20 °C/s, and an increase in the temperature to 75 °C at a rate of 0.2 °C/s with step acquisition of decreasing fluorescence. Successful amplification of a 231-bp human DNA target (p53) was used to assess the nucleic-acid quality of specimens. DNA extracted from the known samples was used in each assay run as a positive control. Melting temperature (Tm) value s.d. were determined by analysis of control samples on three separate runs. Analyses were performed on two different LightCycler instruments.
Results
Confirmation of Clarithromycin Susceptibility of Reference Strains
Each of the seven reference strains demonstrated MIC values ≥1 μg/ml when using the E-test, and were therefore phenotypically resistant to clarithromycin by current CLSI guidelines.13 The unmutated strain demonstrated an MIC value of ≤0.25 μg/ml, and was determined to be phenotypically susceptible to clarithromycin.
Detection/Confirmation of Point Mutations Conferring Resistance to Clarithromycin in the Control Strains
Melt curve analysis of DNA from the cultured reference strains produced three melt curves, with Tm of ∼61.2, 51.8, and 52.2 °C for the wild-type strain and mutant strains A2142G and A2143G, respectively (data not shown). As expected, real-time PCR with melt curve analysis accurately differentiated each of the H. pylori reference strains. The significant difference in Tm allowed for unmutated strains to be easily distinguished from the mutant strains included in the study. However, the smaller differences observed in Tm between the various mutant strains compounded by run-to-run temperature variations precluded reliable determination of specific mutations using this approach.
Detection of Point Mutations Conferring Resistance to Clarithromycin in Formalin-Fixed, Paraffin-Embedded Specimens
PCR/melt curve analysis successfully differentiated mutated and unmutated H. pylori 23S gene targets, as shown in Figure 1. Tm values of samples prepared from formalin-fixed, paraffin-embedded reference strains and from formalin-fixed, paraffin-embedded gastric biopsies demonstrated a small shift toward higher temperatures compared with fresh samples, likely because of the use of differing DNA extraction methods. Non-mutated H. pylori samples showed a Tm value of ∼62.3 °C±1.0 °C, whereas mutated H. pylori samples demonstrated Tm values of ∼53.8 °C±1.0 °C. Mutant strains A2142G and A2143G were considered indistinguishable in formalin-fixed, paraffin-embedded specimens owing to the small differences in Tms and the possibility of significant overlap in Tm ranges.
Limits of Detection
When using fresh isolates, the limit of detection was determined to be 10 genome equivalents of H. pylori. Using 10-μm-thick sections of the formalin-fixed, paraffin-embedded collagen plugs to simulate formalin-fixed, paraffin-embedded biopsy tissue, detection was good but somewhat less sensitive, with a limit of detection of <100 genome equivalents reproducibly detected (data not shown).
Assay Performance
Of the 153 total gastric biopsies studied, 7 failed to show adequate amplification of the human DNA control target and were excluded from the remainder of the study. Of the remaining 146 amplifiable samples, 122 demonstrated immunohistochemical positivity. In all, 103 out of 122 (84%) demonstrated amplifiable H. pylori 23S rRNA gene target and 19 out of 122 (16%) demonstrated no amplification of H. pylori using the PCR assay. This was likely because of formalin overfixation, which has been demonstrated to decrease the sensitivity of PCR.15 The failure of amplification in these cases may also be because of the lack of adequate numbers of organism in the sections used for PCR analysis. Alternatively, immunohistochemical methods are known to cross react with other Helicobacter species,16, 17 and these findings may represent infection with Helicobacter organisms other than H. pylori. The primer and probe combinations used in this study have been previously shown to be specific for H. pylori,12 which raises the possibility of false-positive immunohistochemistry results due to cross reactivity with an expected lack of detection by PCR.
Twenty-two samples were negative for H. pylori by immunohistochemistry and PCR; however two were negative for H. pylori by immunohistochemistry, but were positive by the PCR assay. This could have been due to either greater sensitivity of the PCR assay compared with immunohistochemical staining or due to differences in the number of organisms present in the sections used for PCR versus immunohistochemical staining.
Detection of Mutations
Of the total 105 samples demonstrating PCR positivity for H. pylori, 52 (50%) were positive for mutations associated with resistance (A2142G or A2143G) (Table 1). In addition, the PCR assay was able to demonstrate a mixture of mutated and unmutated wild-type organisms in 11 of the 52 samples.
Discussion
Several previous reports have described the identification of clarithromycin-resistant H. pylori by PCR from clinical specimens, using cultures grown from gastric biopsies12, 18, 19 and more recently from formalin-fixed, paraffin-embedded gastric biopsy tissue.20
The use of a PCR assay to detect clarithromycin-resistant H. pylori in formalin-fixed, paraffin-embedded tissues offers several potential advantages over culture-based methods. As testing can be performed from paraffin-embedded tissue, the special conditions and time constraints for transport can be avoided. The use of formalin-fixed, paraffin-embedded tissues also allows for testing decisions to be made following microscopic interpretation by a pathologist. The most appropriate sections may then be taken for PCR testing, minimizing sampling error that can occur with culture-based methods. Prior microscopic examination of tissues may also reveal that changes characteristic of H. pylori infection are not present, avoiding the need for further testing altogether. The presence of H. pylori in gastric biopsies and clarithromycin resistance status may be assessed simultaneously, guiding correct treatment at the time of diagnosis. Lastly, PCR may be used retrospectively on previously procured specimens to assess whether clarithromycin resistance may have been a factor in treatment failure.
This study suggests that the sensitivity of PCR may be less than that of immunohistochemistry-based methods for the detection of H. pylori in formalin-fixed, paraffin-embedded gastric biopsy specimens, detecting 84% of cases identified by immunohistochemistry. Careful selection of paraffin sections submitted for PCR testing as well as optimal control of formalin fixation times should serve to further mitigate any differences in sensitivity. Immunohistochemistry offers a suitable method for an initial presumed diagnosis of H. pylori infection, particularly in cases where sparse numbers of bacteria are present; however, a subsequent positive PCR assay can be used for confirmation of H. pylori infection, in addition to providing valuable information regarding clarithromycin susceptibility. Also of note, is the fact that the PCR assay was able to detect H. pylori DNA in 2 (of 24) cases demonstrating negative immunohistochemical staining. These findings suggest that utilization of a PCR assay may be helpful in some cases where suspicion for H. pylori infection remains despite negative immunohistochemical staining.
The most recent data obtained from the Helicobacter pylori Antimicrobial Resistance Monitoring Program (HARP) demonstrated an overall clarithromycin resistance rate of 12.9% for the population of the United States.21 In total, nearly half of the patients in our study (50%) were positive for mutations associated with clarithromycin resistance. Although this high percentage is striking in comparison with the HARP study, it is important to note that patients in our study population typically present for biopsy following presumed failure of initial therapy. It is known that the prevalence of antimicrobial resistant H. pylori varies significantly by region worldwide, including different regions of the United States, and has been shown to have a strong correlation to regional antimicrobial practices.22, 23 Future studies including correlation with clinical history will be helpful in elucidating the percentage of patients presenting for biopsy, following failure of therapy that includes a clarithromycin component.
In this study, the melting curves were indistinguishable between the two mutations seen (data not shown). Previous studies have suggested that the presence of the A2143G mutation confers a significantly decreased H. pylori eradication potential when using clarithromycin-containing regimens, more so than the presence of the A2142C or A2142G mutations.24 At this time, the clinical manifestations and current therapies for each are identical and there is no utility for differentiation between the strains in the clinical laboratory setting. However, differentiation of mutations may have importance in future epidemiological studies or as recommendations for antimicrobial therapies change.
The number of known positive H. pylori strains included in this study is limited, and because this was a retrospective study utilizing formalin-fixed tissues, correlation with bacterial culture was unable to be conducted using patient samples. Phenotypic findings of clarithromycin resistance using the E-test did demonstrate 100% concordance with PCR using the supplied H. pylori strains. Although we suspect that our results would be largely concordant, a prospective clinical validation would be helpful using bacterial cultures obtained from fresh endoscopy-obtained biopsy tissue, such as that obtained for rapid urease testing. Comparison with simultaneously obtained formalin-fixed, paraffin-embedded biopsy tissue would be advantageous in discerning the true sensitivity and specificity of the assay in a clinical setting.
It should be noted that no A2142C mutations were seen within the study population. Although our reference strains did not include a sample with the A2142C mutation, the melt curve has been previously shown to be readily distinguishable from unmutated strains and from either of the A to G mutations.12 This result is not surprising as A2142C mutations are significantly more rare and could likely have been absent from this particular study population.25
In summary, we have described the use of a real-time PCR assay based on the LightCycler platform for the rapid detection of H. pylori, and assessment of clarithromycin susceptibility status directly from formalin-fixed, paraffin-embedded gastric biopsy specimens utilizing a probe and primer combination first reported by Oleastro et al.12 in 2003. Failure to amplify the 23S gene was minimally problematic, and likely due to formalin overfixation or sampling error in detecting sparse numbers of organisms. Utilization of PCR primers amplifying a smaller DNA target, or the use of unfixed specimens, may yield a greater number of samples with interpretable results.
The implementation of this method into the clinical laboratory will allow for the identification of H. pylori and for assessment of clarithromycin resistance in previously fixed gastric biopsy specimens in <4-h analytic time. This method could substitute for the time-intensive culture and antimicrobial susceptibility testing process. It would also permit evaluation of formalin-fixed specimens in cases where cultures were not performed, either because they are not routinely performed by the microbiology laboratory or unsuspected H. pylori-associated gastritis is detected on histopathological examination. A rapid means of detecting clarithromycin-resistant H. pylori directly from patient specimens could potentially have a major impact on clinical management of H. pylori-associated gastritis, allowing for the timely assessment of clarithromycin resistance status leading to a decrease in treatment failure rates.
References
Everhart JE . Recent developments in the epidemiology of Helicobacter pylori. Gastroenterol Clin North Am 2000;29:559–578.
Peterson WL, Fendrick AM, Cave DR et al. Helicobacter pylori-related disease: guidelines for testing and treatment. Arch Intern Med 2000;160:1285–1291.
Chey WD, Wong BC . American College of Gastroenterology guideline on the management of Helicobacter pylori infection. Am J Gastroenterol 2007;102:1808–1825.
Vakil N, Megraud F . Eradication therapy for Helicobacter pylori. Gastroenterology 2007;133:985–1001.
Fischbach LA, Goodman KJ, Feldman M et al. Sources of variation of Helicobacter pylori treatment success in adults worldwide: a meta-analysis. Int J Epidemiol 2002;31:128–139.
Graham DY, Fischbach L . Helicobacter pylori treatment in the era of increasing antibiotic resistance. Gut 2010;59:1143–1153.
Versalovic J, Shortridge D, Kibler K et al. Mutations in 23S rRNA are associated with clarithromycin resistance in Helicobacter pylori. Antimicrob Agents Chemother 1996;40:477–480.
Megraud F . H pylori antibiotic resistance: prevalence, importance, and advances in testing. Gut 2004;53:1374–1384.
Vaira D, Gatta L, Ricci C et al. Review article: diagnosis of Helicobacter pylori infection. Aliment Pharmacol Ther 2002;16 (Suppl 1):16–23.
Winn WC, Koneman EW, (eds). Koneman’s Color Atlas and Textbook of Diagnostic Microbiology. Lippincott Williams & Wilkins: Philadelphia, 2006;407 pp.
Perez-Perez GI . Accurate diagnosis of Helicobacter pylori - culture, including transport. Gastroenterol Clin North Am 2000;29:879–884.
Oleastro M, Menard A, Santos A et al. Real-time PCR assay for rapid and accurate detection of point mutations conferring resistance to clarithromycin in Helicobacter pylori. J Clin Microbiol 2003;41:397–402.
Clinical and Laboratory Standards Institute. CLSI Document M45-A2. Methods for Antimicrobial Dilution and Disk Susceptibility Testing of Infrequently Isolated or Fastidious Bacteria; Approved Guideline. Vol. 30 2nd edn. CLSI: Wayne, PA, 2010;p 24.
Menard A, Santos A, Megraud F et al. PCR-restriction fragment length polymorphism can also detect point mutation A2142C in the 23S rRNA gene, associated with Helicobacter pylori resistance to clarithromycin. Antimicrob Agents Chemother 2002;46:1156–1157.
Inoue T, Nabeshima K, Kataoka H et al. Feasibility of archival non-buffered formalin-fixed and paraffin-embedded tissues for PCR amplification: an analysis of resected gastric carcinoma. Pathol Int 1996;46:997–1004.
Singhal AV, Sepulveda AR . Helicobacter heilmannii gastritis: a case study with review of literature. Am J Surg Pathol 2005;29:1537–1539.
Scanziani E, Simpson KW, Monestiroli S et al. Histological and immunohistochemical detection of different Helicobacter species in the gastric mucosa of cats. J Vet Diagn Invest 2001;13:3–12.
Agudo S, Perez-Perez G, Alarcon T et al. High prevalence of clarithromycin-resistant Helicobacter pylori strains and risk factors associated with resistance in Madrid, Spain. J Clin Microbiol 2010;48:3703–3707.
Lascols C, Lamarque D, Costa JM et al. Fast and accurate quantitative detection of Helicobacter pylori and identification of clarithromycin resistance mutations in H. pylori isolates from gastric biopsy specimens by real-time PCR. J Clin Microbiol 2003;41:4573–4577.
De Francesco V, Margiotta M, Zullo A et al. Claritromycin resistance and Helicobacter pylori genotypes in Italy. J Microbiol 2006;44:660–664.
Duck WM, Sobel J, Pruckler JM et al. Antimicrobial resistance incidence and risk factors among Helicobacter pylori-infected persons, United States. Emerg Infect Dis 2004;10:1088–1094.
De Francesco V, Giorgio F, Hassan C et al. Worldwide H. pylori antibiotic resistance: a systematic review. J Gastrointestin Liver Dis 2010;19:409–414.
Boyanova L, Mitov I . Geographic map and evolution of primary Helicobacter pylori resistance to antibacterial agents. Expert Rev Anti Infect Ther 2010;8:59–70.
De Francesco V, Margiotta M, Zullo A et al. Clarithromycin-resistant genotypes and eradication of Helicobacter pylori. Ann Int Med 2006;144:94–100.
van Doorn LJ, Glupczynski Y, Kusters JG et al. Accurate prediction of macrolide resistance in Helicobacter pylori by a PCR line probe assay for detection of mutations in the 23S rRNA gene: multicenter validation study. Antimicrob Agents Chemother 2001;45:1500–1504.
Acknowledgements
We thank the National Campylobacter and Helicobacter Reference Laboratory, Centers for Disease Control and Prevention, for the provision of reference strains. We also thank Scott Resnick for his assistance with the study. This study was funded by the Department of Pathology and Laboratory Medicine at NorthShore University HealthSystem, Evanston, IL, USA.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
This research was presented in part at the American Society for Microbiology 111th General Meeting, New Orleans, LA, USA 21–24 May, 2011.
Rights and permissions
About this article
Cite this article
Schmitt, B., Regner, M., Mangold, K. et al. PCR detection of clarithromycin-susceptible and -resistant Helicobacter pylori from formalin-fixed, paraffin-embedded gastric biopsies. Mod Pathol 26, 1222–1227 (2013). https://doi.org/10.1038/modpathol.2013.48
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2013.48
Keywords
This article is cited by
-
Pyrosequencing analysis for rapid and accurate detection of clarithromycin resistance-associated mutations in Iranian Helicobacter pylori isolates
BMC Research Notes (2023)
-
In situ targeted MRI detection of Helicobacter pylori with stable magnetic graphitic nanocapsules
Nature Communications (2017)
-
Helicobacter pylori Clarithromycin Resistance and Treatment Failure Are Common in the USA
Digestive Diseases and Sciences (2016)