Abstract
The objectives of this study were to determine the prognostic significance of subgrouping estrogen receptor (ER)-positive breast tumors into low- and high-risk luminal categories using Ki67 index, TP53, or progesterone receptor (PR) status. The study group comprised 540 patients with lymph node negative, invasive breast carcinoma. Luminal A subtype was defined as being ER positive, HER2 negative, and Ki67 low (<14% cells positive) and luminal B subtype as being ER positive, HER2 negative, and Ki67 high (≥14% cells positive). Luminal tumors were also subgrouped into risk categories based on the PR and TP53 status. Survival analysis was performed. Patients with luminal B tumors (n=173) had significantly worse disease-free survival compared to those with luminal A tumors (n=186) (log rank P-value=0.0164; univariate Cox regression relative risk 2.00; 95% CI, 1.12–3.58; P=0.0187). Luminal subtype remained an independent prognostic indicator on multivariate analysis including traditional prognostic factors (relative risk 2.12; 95% CI, 1.16–3.88; P=0.0151). Using TP53 status or PR negativity rather than Ki67 to classify ER-positive luminal tumors gave similar outcome results to those obtained using the proliferation index. However, it was a combination of the three markers, which proved the most powerful prognostically. Ki67 index, TP53 status, or PR negativity can be used to segregate ER-positive, HER2-negative tumors into prognostically meaningful subgroups with significantly different clinical outcomes. These biomarkers particularly in combination may potentially be used clinically to guide patient management.
Similar content being viewed by others
Main
Despite the generally favorable prognosis of estrogen receptor (ER)-positive, lymph node-negative, invasive breast carcinoma patients, a subset will experience relapse.1, 2 Prospective identification of this subgroup of early-stage ER-positive patients with a relatively poorer prognosis who may benefit from adjuvant chemotherapy is of paramount importance.
Using gene expression profiling, at least two distinct molecular subgroups (luminal A and B) have been identified within the category of hormone receptor-positive breast cancer. Luminal B tumors show lower expression of estrogen-related genes, are more often high grade with a higher Ki67 index and consistently have been shown to have poorer outcome compared with their luminal A counterparts.3, 4 Immunohistochemical panels may be used as expression surrogates to identify these intrinsic breast cancer subtypes. Despite the fact that varying criteria have been used to define each category, many studies have shown immunohistochemical-based classifiers to be prognostically significant.5, 6, 7 Luminal A cancers have been defined by a number of groups as hormone receptor-positive and HER2-negative and luminal B cancers as both hormone receptor positive and HER2 positive.8, 9, 10, 11, 12 However, this definition is flawed as ∼70% of luminal B tumors, defined by gene expression profiling, are in fact HER2 negative.13 Rather, evidence suggests that it is the proliferation rate that primarily distinguishes luminal A from luminal B cases.13, 14
Recently, Cheang et al13 proposed an immunohistochemical classifier incorporating the Ki67 index, recognizing the pivotal role of proliferation in subclassifying luminal breast cancers. However, alternative markers also warrant consideration. TP53 positivity in breast cancer is significantly associated with high histological grade and high proliferation index,12, 15 and consequently could potentially be substituted for Ki67 in an immunohistochemical-based classifier. Progesterone receptor (PR) negativity correlates with a high Oncotype DX® recurrence score in keeping with a luminal B genotype,8, 16, 17 and therefore represents another possible surrogate marker for Ki67 in segregating luminal cancers into low- and high-risk categories.
The objectives of this study were to determine whether subgrouping ER-positive, HER2-negative, node-negative breast cancers into low- and high-risk luminal categories using (i) Ki67 index, (ii) TP53 status, and (iii) PR negativity or a combination of these markers is prognostically informative.
Materials and methods
Patient Cohort and Clinical Follow-Up
The patient cohort comprises a prospectively accrued consecutive series of women with lymph node-negative, invasive breast carcinoma enrolled at eight Toronto hospitals from September 1987 to October 1996, as previously described.18 Written informed consent was obtained from all participants. Patients were followed for recurrence and death. Disease-free survival was taken as the time between diagnosis and the confirmation of non-breast recurrence. All patients were monitored for death whether or not they experienced disease recurrence. Using clinical follow-up data, patient status on 10 January 2002 determined disease-free survival times and censoring status. Follow-up data were monitored for an additional 6 months to confirm patient status at the termination date.
Approval of the study protocol was obtained from the research ethics boards of Mount Sinai Hospital (#01-0313-U) and the University Health Network (#02-0881-C). Written informed consent was received from all study participants.
Histopathology, Hormone Receptor Status, Tissue Microarray Construction, and Immunohistochemical Staining
Key pathological data were extracted from the histology reports. ER and PR status was determined biochemically at the time of surgery by ligand binding assay of frozen tissue, as this was the standard approach utilized at this time.18 While immunohistochemical determination of hormone receptor status is now the standard of practice, good correlation between the two methods has been shown (kappa=0.72).19 The cut point to define PR positivity was based on the prior validation.19 Formalin-fixed paraffin-embedded tumor blocks were available for 887 patients. Areas of invasive carcinoma were selected from an H&E-stained section of each tumor and two 0.6-mm cores of tissue were taken from the corresponding areas of the paraffin block. The selected donor cores were embedded in a recipient paraffin block and 4-μm sections were cut from this and used in a series for immunohistochemical staining for HER2, Ki67, and TP53, under the conditions described in Table 1. Ki67 was visually scored by one pathologist for the percentage of positive tumor nuclei, regardless of intensity of staining. In all, 50 nuclei were counted and therefore tissue microarray core samples with <50 tumor cells were deemed unsatisfactory, resulting in exclusion of 220 cases. The HER2 and TP53 immunohistochemical-stained sections were scored using the Allred scoring method with cut points to define positivity based on the previous validations (for HER2: >4 and for TP53: >3).20, 21 Nuclear staining was scored for TP53. Strong complete membrane staining was assessed for HER2. The raw score data were processed using a tissue microarray deconvoluter software program. As two cores from each tumor were assessed, the larger of the two values was chosen for use in statistical analysis to minimize the effect of false negatives on the array.
Definitions of Intrinsic Subgroups
The luminal group was classified as HER2 negative and ER positive, and was then further subclassified as luminal A if Ki67 is low (<14% tumor cell positivity) and luminal B if Ki67 is high (≥14% tumor cell positivity). All HER2-positive cases were excluded from analysis in this study, the rationale being that these patients are generally treated with both Trastuzumab and chemotherapy in contrast to other hormone receptor-positive cancers. The cut point chosen for Ki67 high vs Ki67 low was based on the work of Cheang et al.13 Alternatively using PR, low-risk luminal subtype was defined as PR positive and high-risk luminal subtype as PR negative; using TP53 similarly, low-risk luminal subtype was defined as TP53 negative and high-risk luminal subtype as TP53 positive. Finally, combinations of the above markers were also used to define the luminal subtypes, as described later.
Statistical Analysis
Of the 887 patients in whom tumor blocks were available for use in tissue microarray construction, complete HER2, Ki67, TP53 and biochemical ER and PR data were available in 540 cases. Univariate disease-free survival analysis according to intrinsic subgroup status was by the log-rank test with Kaplan–Meier survival curves. Univariate and multivariate disease-free survival analysis was by the Cox proportional hazards model. In addition, the luminal subgroups were assessed for association with clinical and pathological parameters, including menopausal status, age at diagnosis, tumor size, histologic grade, lymphatic invasion, and administered adjuvant therapy using the χ2 test, Fisher’s exact test, or t-test.
Results
Classification of Intrinsic Subgroups
On the basis of our definition, 359 of 540 tumors (67%) were designated as luminal cancers. Using Ki67 labelling index to stratify luminal tumors, 173 of 359 (48%) tumors were assigned to the luminal A subgroup and 186 (52%) to the luminal B subgroup. Using TP53 positivity to stratify luminal tumors, 311 of 359 (87%) tumors were assigned to the low-risk luminal subgroup and 48 (13%) to the high-risk luminal subgroup. Using PR negativity to stratify luminal tumors, 306 of 359 (85%) tumors were assigned to the low-risk luminal subgroup and 53 (15%) to the high-risk luminal subgroup.
Prognostic Relevance According to Luminal Subgroup, TP53 and PR Status
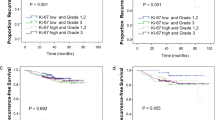
Seventy-three (66%) recurrences were observed in the tissue microarray cohort of 540 patients (median follow-up of 103 months) and fifty-two (47%) recurrences in the ER+ luminal group of 359 (median follow-up of 105 months). In comparison of the ER+ luminal subgroups, patients with luminal B tumors defined by Ki67 had a significantly worse disease-free survival than patients with luminal A tumors (log-rank P-value=0.0164) (Figure 1). Using TP53 status (log-rank P-value=0.0172) or PR negativity (log-rank P-value=0.0259) to classify the ER+ luminal tumors further refined the luminal group and gave similar disease-free survival results to those obtained using Ki67 (Figure 1). The luminal B subtype association with significantly worse disease-free survival found on univariate Cox model analysis (relative risk 2.00; 95% confidence interval, 1.12–3.58; P=0.0187) was retained on multivariate analysis adjusting for standard clinical and pathological parameters (relative risk 2.12; 95% confidence interval, 1.16–3.88; P=0.0151) (Table 2). Substitution of either TP53 or PR status for the Ki67 index in the subclassification of luminal tumors was also found to be of independent prognostic significance on multivariate analysis (Supplementary Tables S1 and S2).
To ascertain the level of agreement among the three methods for segregating luminal cases into low- and high-risk subgroups, kappa statistics were calculated that ranged from 0.06 to 0.22 indicating only slight to fair agreement. Variation in the distribution of luminal subtypes using the three different approaches and low kappa values suggest that the various methods are identifying different patients as being in the poor prognostic group. Therefore, survival analysis was repeated to investigate whether using a combination of these markers provides more accurate prognostic information. The following combinations were analyzed: high-risk luminal=Ki67 high and PR negative; high-risk luminal=Ki67 high and TP53 positive; high-risk luminal=Ki67 high and either PR negative or TP53 positive. Each time, the remaining tumors in ER+ group were defined as low-risk luminal. Using these definitions, high-risk luminal tumors were consistently found to have poorer disease-free survival compared with low-risk luminal tumors with log-rank P-values of 0.0005, 0.0283, and 0.0001, respectively (Figure 2). Furthermore, when we define high-risk luminal by Ki67 and either PR or TP53, the percentage of recurrences is highest, demonstrating that this approach to classification is the most sensitive at identifying more of the poor prognosis patients (Table 3).
Luminal Subgroups and Clinicopathologic Parameters
Correlation of the luminal intrinsic subgroups (defined using Ki67 and either TP53 or PR) with clinical parameters and tumor characteristics is presented in Table 4. High-risk luminal tumors were found to significantly correlate with higher tumor grade (P<0.0001). When the luminal subgroups defined using two markers (Ki67 and TP53 or Ki67 and PR) were compared, higher tumor grade (P<0.0001, highly significant) was again found to be associated with high-risk luminal subtype (Supplementary Tables S3 and S4).
Discussion
Gene expression profiling and immunohistochemical biomarkers have identified subsets of patients with invasive breast carcinoma that have distinct clinical outcomes. Patients with molecularly defined luminal B tumors show highest mortality after 5–8 years.22 Late relapse of the latter subtype may be triggered by discontinuation of endocrine therapy or may reflect its natural history and intrinsic aggressiveness. As such, there is a clear need to prospectively identify patients with luminal B cancers, particularly those in the lymph node-negative category, who may benefit from adjuvant chemotherapy.
Increasingly medical oncologists are utilizing molecular tests to guide therapeutic decision making with the Oncotype DX® (Genomic Health) recurrence score gaining widespread acceptance in routine practice. The recurrence score has been shown to correlate with risk of both loco-regional23 and distant recurrence in ER-positive, node-negative breast cancer.24, 25 Furthermore, this 21 multigene assay is predictive of treatment response to endocrine therapy when the recurrence score is low and to chemotherapy when the recurrence score is high.26, 27 Proliferation-related genes receive the highest weighting in the algorithm and consequently make the greatest contribution to the determination of the recurrence score. Therefore, the test essentially differentiates good prognosis luminal A cases from poorer prognosis luminal B cases.13, 28 Supportive of this conclusion is the finding that among ER-positive, luminal B tumors the rate of high-risk designation with Oncotype DX® is 93%.29
The contribution of molecular classification to clinical decision making in breast cancer is limited by its high cost and its close correlation with established prognostic markers. For example, higher Oncotype DX® recurrence scores have been shown to correlate with high grade, PR negativity, and HER2 positivity.8, 16, 17, 30 Furthermore, an immunohistochemical panel of ER, PR, HER2, and Ki67 in conjunction with standard clinicopathological parameters (IHC4 score) has been shown to provide similar prognostic information to the recurrence score in hormone receptor-positive patients treated with endocrine therapy.31 Integration of such an immunohistochemical panel into routine practice would merely require the addition of Ki67 to the pathologist’s workload. Unfortunately, Ki67 immunohistochemistry poses a number of challenges including (i) definition of a clinically relevant cut point, (ii) the impact of intratumoral heterogeneity, and (iii) the appropriateness of using the most proliferative area or the average proliferation rate.32
Given that the Ki67 proliferation index represents a continuum, where best to define a clinically relevant cut point is unclear. Although levels ranging from 10 to –20% have been most commonly used to segregate patient populations, the International Ki67 in Breast Cancer Working Group was unable to reach a consensus regarding the optimal cut point for use in clinical practice.32 Cheang et al proposed 14% as the ideal cut off point for the Ki67 index in differentiating luminal A from B cases. This threshold was calculated based on the gene-expression profiles of 357 breast cancers as determined using PAM50 and subsequently validated in an independent cohort of 4046 breast cancers. PAM50 is a 50 gene predictor of breast cancer intrinsic subtype, which has been shown to be of prognostic and predictive value independent of pathologic stage, grade, and ER status.33 On the basis of these results, they formulated the following immunohistochemical classifier: luminal A=ER and/or PR positive, HER2 negative, and Ki67 index <14%; luminal B=ER and/or PR positive, HER2 negative, and Ki67 index ≥14%. The subgroup of luminal B cases that were HER2 positive was classified separately as ‘luminal-HER2’. This classification was shown to be an independent prognostic indicator with luminal A tumors having the best outcome and luminal-HER2 the worst, regardless of lymph-node status.13 Using the same immunohistochemical definition, Hugh et al14 also demonstrated that luminal B tumors have a worse disease-free survival and overall survival than luminal A cases in a node-positive breast cancer cohort. Millar et al34 in addition to using Ki67 with a cut point of 14% also integrated TP53 into their immunohistochemical classifier, which again was shown to be prognostically informative. Our study which was confined to node-negative patients confirms the findings of these prior studies and further supports the use of 14% as a clinically relevant cut point to distinguish luminal A from luminal B tumors.
A further issue with the immunohistochemical determination of Ki67 proliferation index is the frequent occurrence of intratumoral variation in labelling. Two types of heterogeneity are described: (i) increasing staining toward the tumor edge and (ii) hot spots.32 This has led to speculation that tissue microarrays may be inappropriate for assessing the Ki67 index. Two studies have addressed this issue. A breast cancer study found all anticipated associations between the Ki67 index measured using tissue microarrays and clinicopathological parameters such as high grade and poor prognosis. In addition, expected associations with molecular markers including TP53 positivity were also demonstrated.35 A study on urothelial carcinoma similarly showed that the proliferation index can be accurately and reliably measured using a tissue microarray with a single core per tumor.36 However, there are no published studies comparing Ki67 indices on whole sections vs tissue microarrays in breast cancer.32 An additional unresolved issue is whether the area with the highest percentage of Ki67-positive cells or the average count should be utilized for scoring purposes.
When we re-classified ER-positive tumors utilizing PR negativity instead of the Ki67 index, similar survival results were obtained. In fact, ER-positive tumors that are PR negative have been shown to have a poorer prognosis than dual positive cases by many groups. This is unsurprising given that PR is regulated by, ER37 and therefore a lack of PR expression is indicative of a dysregulated ER pathway. Furthermore, many but not all studies have found PR negativity to be predictive of reduced responsiveness to endocrine therapy which may be the key factor accounting for the less favorable outcome in these patients.38
Utilization of TP53 status to segregate the luminal group was also found to be prognostically significant in our study. TP53 is a tumor-suppressor gene, which has a key role in cell-cycle checkpoint control exerting both anti-proliferative and anti-apoptotic effects.39 TP53 alterations whether detected by molecular or immunohistochemical assays have been shown to be a marker of poor prognosis and to be predictive of treatment response by some but not all investigators.7, 12, 15, 22, 38
Of note, when we substituted either PR or TP53 for Ki67 marked variation was seen in the distribution of the luminal subgroups with associated poor kappa statistics. These findings indicate that the three methods employed in this study identified different patients as being of poor prognosis, thereby limiting the clinical utility of the Ki67 classification alone. However, a combination of Ki67 with PR and/or TP53 identified a highest percentage of recurrences by picking up more of the poor prognosis patients. Furthermore, a combination of all three markers was also the best predictor of disease-free survival according to the P-value for the test of association in the multivariate model.
Our findings confirm that Ki67 can be used to segregate ER-positive, node-negative breast cancers into prognostically meaningful subgroups with the potential for guiding patient therapy. Currently, there are a number of unresolved issues with regard to measuring the proliferation index as highlighted by the International Ki67 in Breast Cancer Working Group.32 A number of potential surrogate markers including TP53 and PR may prove easier to integrate into routine practice and warrant further study. However, it is a combination of these markers, which may prove to be of greatest clinical utility.
References
Early Breast Cancer Trialists’ Collaborative Group. Effects of chemotherapy and hormonal therapy for early breast cancer on recurrence and 15-year survival: an overview of the randomised trials. Lancet 2005;365:1687–1717.
Boyages J, Taylor R, Chua B et al. A risk index for early node-negative breast cancer. Br J Surg 2006;93:564–571.
Correa Geyer F, Reis-Filho JS . Microarray-based gene expression profiling as a clinical tool for breast cancer management: are we there yet? Int J Surg Pathol 2009;17:285–302.
Wirapati P, Sotiriou C, Kunkel S et al. Meta-analysis of gene expression profiles in breast cancer: toward a unified understanding of breast cancer subtyping and prognosis signatures. Breast Cancer Res 2008;10:R65.
Abd El-Rehim DM, Ball G, Pinder SE et al. High-throughput protein expression analysis using tissue microarray technology of a large well-characterised series identifies biologically distinct classes of breast cancer confirming recent cDNA expression analyses. Int J Cancer 2005;116:340–350.
Nguyen PL, Taghian AG, Katz MS et al. Breast cancer subtype approximated by estrogen receptor, progesterone receptor, and HER-2 is associated with local and distant recurrence after breast-conserving therapy. J Clin Oncol 2008;26:2373–2378.
Mulligan AM, Pinnaduwage D, Bull S et al. Prognostic effect of basal-like breast cancers is time dependent: evidence from tissue microarray studies on a lymph node-negative cohort. Clin Cancer Res 2008;14:4168–4174.
Tang P, Wang J, Hicks DG et al. A lower Allred score for progesterone receptor is strongly associated with a higher recurrence score of 21-gene assay in breast cancer. Cancer Invest 2010;28:978–982.
Tang P, Wang J, Bourne P . Molecular classifications of breast carcinoma with similar terminology and different definitions: are they the same? Hum Pathol 2008;39:506–513.
Matos I, Dufloth R, Alvarenga M et al. p63, cytokeratin 5, and P-cadherin: three molecular markers to distinguish basal phenotype in breast carcinomas. Virchows Arch 2005;447:688–694.
Carey LA, Perou CM, Livasy CA et al. Race, breast cancer subtypes, and survival in the Carolina Breast Cancer Study. JAMA 2006;295:2492–2502.
Jung S, Jeong J, Shin SH et al. Accumulation of p53 determined by immunohistochemistry as a prognostic marker in node negative breast cancer; analysis according to St Gallen consensus and intrinsic subtypes. J Surg Oncol 2011;103:207–211.
Cheang MC, Chia SK, Voduc D et al. Ki67 index, HER2 status, and prognosis of patients with luminal B breast cancer. J Natl Cancer Inst 2009;101:736–750.
Hugh J, Hanson J, Cheang MC et al. Breast cancer subtypes and response to docetaxel in node-positive breast cancer: use of an immunohistochemical definition in the BCIRG 001 trial. J Clin Oncol 2009;27:1168–1176.
Kim HS, Yom CK, Kim HJ et al. Overexpression of p53 is correlated with poor outcome in premenopausal women with breast cancer treated with tamoxifen after chemotherapy. Breast Cancer Res Treat 2010;121:777–788.
Flanagan MB, Dabbs DJ, Brufsky AM et al. Histopathologic variables predict Oncotype DX recurrence score. Mod Pathol 2008;21:1255–1261.
Wolf I, Ben-Baruch N, Shapira-Frommer R et al. Association between standard clinical and pathologic characteristics and the 21-gene recurrence score in breast cancer patients: a population-based study. Cancer 2008;112:731–736.
Andrulis IL, Bull SB, Blackstein ME et al. neu/erbB-2 amplification identifies a poor-prognosis group of women with node-negative breast cancer. Toronto Breast Cancer Study Group. J Clin Oncol 1998;16:1340–1349.
Mohsin SK, Weiss H, Havighurst T et al. Progesterone receptor by immunohistochemistry and clinical outcome in breast cancer: a validation study. Mod Pathol 2004;17:1545–1554.
O’Malley FP, Parkes R, Latta E et al. Comparison of HER2/neu status assessed by quantitative polymerase chain reaction and immunohistochemistry. Am J Clin Pathol 2001;115:504–511.
Done S, Arneson NC, Ozcelik H et al. p53 protein accumulation in non-invasive lesions surrounding p53 mutation positive invasive breast cancer. Breast Cancer Res Treat 2001;65:111–118.
Langerød A, Zhao H, Borgan Ø et al. TP53 mutation status and gene expression profiles are powerful prognostic markers of breast cancer. Breast Cancer Res 2007;9:R30.
Mamounas EP, Tang G, Fischer B et al. Association between the 21-gene recurrence score assay and risk of locoregional recurrence in node-negative, estrogen receptor-positive breast cancer: results from NSABP B-14 and NSABP B-20. J Clin Oncol 2010;28:1677–1683.
Dowsett M, Cuzick J, Wale C et al. Prediction of risk of distant recurrence using the 21-gene recurrence score in node-negative and node-positive postmenopausal patients with breast cancer treated with anastrozole or tamoxifen: a TransATAC study. J Clin Oncol 2010;28:1829–1834.
Paik S, Shak S, Tang G et al. A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N Engl J Med 2004;351:2817–2826.
Paik S, Tang G, Shak S et al. Gene expression and benefit of chemotherapy in women with node-negative, estrogen receptor–positive breast cancer. J Clin Oncol 2006;24:3726–3734.
Albain KS, Barlow WE, Shak S et al. Prognostic and predictive value of the 21-gene recurrence score assay in postmenopausal women with node-positive, estrogen-receptor-positive breast cancer on chemotherapy: a retrospective analysis of a randomized trial. Lancet Oncol 2010;11:55–65.
Brenton JD, Carey LA, Ahmed AA et al. Molecular classification and molecular forecasting of breast cancer: ready for clinical application? J Clin Oncol 2005;23:7350–7360.
Sotiriou C, Pusztai L . Gene-expression signatures in breast cancer. N Engl J Med 2009;360:790–800.
Asad J, Jacobson AF, Estabrook A et al. Does Oncotype DX recurrence score affect the management of patients with early-stage breast cancer? Am J Surg 2008;196:527–529.
Cuzick J, Dowsett M, Pineda S et al. Prognostic value of a combined estrogen receptor, progesterone receptor, Ki-67, and human epidermal growth factor receptor 2 immunohistochemical score and comparison with the Genomic Health recurrence score in early breast cancer. J Clin Oncol 2011;29:4273–4278.
Dowsett M, Nielsen TO, A’Hern R et al. Assessment of Ki67 in breast cancer: recommendations from the international Ki67 in breast cancer working group. J Natl Cancer Inst 2011;103:1656–1664.
Parker JS, Mullins M, Cheang MC et al. Supervised risk predictor of breast cancer based on intrinsic subtypes. J Clin Oncol 2009;27:1160–1167.
Millar EK, Graham PH, McNeil CM et al. Prediction of outcome of early ER+ breast cancer is improved using a biomarker panel, which includes Ki-67 and p53. Br J Cancer 2011;105:272–280.
Ruiz C, Seibt S, Al Kuraya K et al. Tissue microarrays for comparing molecular features with proliferation activity in breast cancer. Int J Cancer 2006;118:2190–2194.
Nocito A, Kononen J, Kallioniemi OP et al. Tissue microarrays (TMAs) for high-throughput molecular pathology research. Int J Cancer 2001;94:1–5.
Horwitz KB, McGuire WL . Estrogen control of progesterone receptor in human breast cancer. Correlation with nuclear processing of estrogen receptor. J Biol Chem 1978;253:2223–2228.
Linke SP, Bremer TM, Herold CD et al. A multimarker model to predict outcome in tamoxifen-treated breast cancer patients. Clin Cancer Res 2006;12:1175–1183.
Chen X, Ko LJ, Jayaraman L et al. p53 levels, functional domains, and DNA damage determine the extent of the apoptotic response of tumor cells. Genes Dev 1996;10:2438–2451.
Acknowledgements
This work was supported by the Canadian Cancer Society Research Institute and the Canadian Institutes of Health Research (ILA and SBB). We thank the study participants and the contributions of the Toronto Breast Cancer Group to this work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Modern Pathology website
Rights and permissions
About this article
Cite this article
Feeley, L., Mulligan, A., Pinnaduwage, D. et al. Distinguishing luminal breast cancer subtypes by Ki67, progesterone receptor or TP53 status provides prognostic information. Mod Pathol 27, 554–561 (2014). https://doi.org/10.1038/modpathol.2013.153
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2013.153
Keywords
This article is cited by
-
Searching for the “Holy Grail” of breast cancer recurrence risk: a narrative review of the hunt for a better biomarker and the promise of circulating tumor DNA (ctDNA)
Breast Cancer Research and Treatment (2024)
-
Prediction of Oncotype DX recurrence score using deep multi-layer perceptrons in estrogen receptor-positive, HER2-negative breast cancer
Breast Cancer (2020)
-
Tumoral BRD4 expression in lymph node-negative breast cancer: association with T-bet+ tumor-infiltrating lymphocytes and disease-free survival
BMC Cancer (2018)
-
Spinale Mammakarzinommetastasen aus Sicht der Wirbelsäulenchirurgie
Der Orthopäde (2018)
-
St Gallen 2015 subtyping of luminal breast cancers: impact of different Ki67-based proliferation assessment methods
Breast Cancer Research and Treatment (2016)