Abstract
Histone deacetylases (HDACs) have a critical role in epigenetic gene silencing, rendering a compact chromatin structure by removing acetyl groups from lysine residues within the tails of core histones, thereby repressing gene expression. Epigenetic transcriptional dysregulation is an important oncogenic mechanism in some sarcomas associated with translocations, for which antitumor activity by HDAC inhibitors has been shown in preclinical studies. Nevertheless, the expression of the protein targets of these drugs has not yet been broadly surveyed in this neoplasia. In this study, we assess the expression of HDAC1 and 2 by immunohistochemistry in a tissue microarray series of 1332 cases, representing 44 categories of malignant and borderline mesenchymal tumors. HDAC2 was the more highly expressed isoform, and was more strongly expressed in translocation-associated sarcomas than in other mesenchymal tumors or normal tissues. HDAC1, in contrast, displayed lower expression in translocation-associated sarcomas than in other mesenchymal tumors or in normal tissues. These results indicate that HDAC1 and HDAC2 are differentially expressed in mesenchymal neoplasms, and suggest that HDAC2 is the isoform more likely contributing to the pathogenesis of many translocation-associated sarcomas and to their response to HDAC inhibitors.
Similar content being viewed by others
Main
Histone deacetylases (HDACs) are a heterogeneous group of enzymes that act mainly as transcriptional regulators, by deacetylating the lysine residues in the amino-terminal tails of histone proteins. Deacetylation of histones is one of the epigenetic mechanisms regulating gene expression, associated with a compact state of chromatin and transcriptional silencing. Epigenetic deregulation of gene expression is strongly implicated in cancer pathogenesis and can occur through abnormal recruitment of HDACs to gene promoters by, for example, the fusion product of chromosomal translocations in leukemias and sarcomas.1, 2, 3, 4
The classical HDACs comprise a family of 11 HDAC isoforms that share a zinc-dependent catalytic domain with a high degree of homology, and are grouped in three classes on the basis of their structural similarity to homologous proteins in yeast. Class I HDACs (HDAC1, 2, 3 and 8), related to the yeast transcriptional regulator RPD3, are the best characterized and the most important in carcinogenesis.5
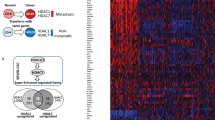
Transcriptional dysregulation through epigenetic mechanisms has been shown to be an important event underlying the pathogenesis of some translocation-associated sarcomas, including synovial sarcoma. In tumor cell lines and in cells engineered to stably express SS18-SSX, this fusion oncoprotein represses the EGR1 tumor suppressor gene through histone modifications and recruitment of Polycomb group-epigenetic repressor complex proteins. Target gene repression is associated with histone-3 deacetylation and Polycomb-mediated repressive histone methylation events, which can be reversed by treatment with the HDAC inhibitor romidepsin.2 Similarly, in endometrial stromal sarcoma, the pathognomonic translocation event JAZF1-JJAZ1 and its variants JAZF1-PHF1 and EPC1-PHF1, all involve rearrangements of Polycomb group genes. The oncogenic mechanism of this fusion oncoprotein is not fully understood,6 but endometrial stromal sarcoma cells highly express HDAC2.7 The HDAC inhibitors suberoylanilide hydroxamic acid and valproate induce cell cycle arrest, differentiation and death in ESS-1 cell lines,7, 8 supporting the involvement of abnormal gene repression by HDAC2 in the pathogenesis of endometrial stromal sarcoma. In Ewing family tumors, the fusion oncoprotein EWS-FLI1 downregulates a greater number of genes than it upregulates.9 In Ewing cell lines, the Polycomb group histone methyltransferase EZH2 is one of the target genes upregulated by EWS-FLI1.10 Transcriptional repression of mesenchymal differentiation mediated by EZH2-containing Polycomb complexes can account in part for the repressive activity of Ewing-associated fusion transcription factors. EZH2 interacts with HDAC2 via EED,11 and EZH2 gene silencing is abrogated by HDAC inhibition in prostate cancer cells.12 Restoration of repressed genes by HDAC inhibitors supports a role for HDAC-mediated transcriptional repression in Ewing family tumors.13 Several HDAC inhibitors have been shown to be effective in Ewing preclinical models,14, 15, 16 and comparable findings have now been reported in other translocation-associated sarcomas, including synovial sarcoma,17 clear cell sarcoma, myxoid liposarcoma and desmoplastic small-blue round-cell tumor.18
Despite this body of preclinical evidence supporting HDAC inhibitors as therapeutic agents in translocation-associated sarcomas, the well-documented involvement of class I HDACs in cancer biology,19 and the recent opening of phase II clinical trials testing these agents in sarcomas, little information exists on the expression of the enzymes targeted by these drugs in mesenchymal neoplasms. In this study, we assay, by immunohistochemistry, the expression of HDAC 1 and 2 isoforms in a broad collection of mesenchymal neoplasms, to assess if there is high expression of specific HDAC isoforms in translocation-associated sarcomas that may help explain their sensitivity to this class of drugs and provide information to help determine which agents may be most effective.
Materials and methods
Tissue Microarrays
Both previously described and more recently constructed tissue microarrays were used in this study, each containing replicate 0.6 mm cores directed to diagnostic, viable areas of tumor by a soft tissue subspecialty pathologist. Published tissue arrays from The Genetic Evaluation Centre (Vancouver, Canada) and Stanford Medical Center (Stanford, CA, USA) used in this study included: MPNST20 (malignant peripheral nerve sheath tumor differential diagnosis array, 176 cases in duplicate), TMA 01-00321 (synovial sarcoma differential diagnosis array, 82 cases in duplicate), TMA 03-00822 (chondroid tumor array, 121 cases in duplicate), TMA 06–00723 (liposarcoma array, 69 cases in triplicate), TMA 06-00124 (gastrointestinal stromal tumors (GIST) array, 713 cases in duplicate), TA-166 and TA-17025 (rebuilds of previous multisarcoma arrays). Four additional tissue arrays not previously published were also employed for this study. TMA09-006 (epithelioid sarcoma differential diagnosis tissue microarray) contains 53 cases in duplicate: 9 epithelioid sarcomas, 3 epithelioid MPNST, 4 epithelioid angiosarcomas, 2 hemangioendotheliomas, 7 melanomas, 8 squamous cell carcinomas, 4 granuloma annulare, 5 necrosis lipodica, 7 rheumatoid nodules, 2 atypical fibroxantomas and 2 clear-cell sarcomas. TMA 03-004 (epithelioid sarcoma array) added six additional cases of epithelioid sarcoma. The TMA 08-019 (endometrial stromal sarcoma array) contains duplicate cores of 21 endometrial stromal sarcomas, 7 undifferentiated endometrial sarcomas, 10 adenosarcomas, 6 adenosarcomas with sarcomatous overgrowth, 2 malignant mixed-müllerian tumors, 2 uterine tumors resembling ovarian sex cord tumors (UTROSCT), 2 cellular leiomyomas, 1 uterine leiomyosarcoma and 7 cases of polypoid endometriosis. Tissue array 10-009 contains triplicate cores of eight cases of alveolar soft part sarcoma, and two cases each of alveolar rhabdomyosarcoma and desmoplastic small round-cell tumor. Only sarcomas, borderline mesenchymal tumors and normal tissues in these arrays were included in this study.
Western Blotting and Immunizing Peptide Blocking Experiments
Antibody specificity was tested by western blotting. Total protein lysates from SYO-1 and DTC1 cell lines18 were prepared using RIPA buffer containing sodium chloride 150 mM/l, 1% NP-40, 0.5% sodium deoxycolate, 0.1% sodium dodecyl sulfate, 50 mM/l Tris pH 8.0 and ready-to-use complete protease inhibitor cocktail (Roche, Mannheim, Germany). Protein extracts were run in SDS–PAGE and analyzed on western blots incubated with primary mouse monoclonal anti-HDAC1 1:2500 (Millipore, Billerica, MA, USA; 05-100), mouse monoclonal anti-HDAC2 1:4000 (Abcam, Cambridge, MA, USA; ab12169), mouse monoclonal anti-HDAC3 1:1000 (Becton Dickinson, Franklin Lakes, CA, USA; 611124), rabbit monoclonal anti-HDAC3 1:7500 (Abcam; ab32369) and rabbit polyclonal anti-HDAC6 1:200 (Abcam; ab1440). For HDAC1, HDAC2 and HDAC6, before neutralization for 30 minutes at room temperature with HDAC1 (1:1000), HDAC2 (1:1000) and HDAC6 (1:1000) peptides (Abcam) that correspond to the epitope recognized by the primary antibodies was performed. Horseradish peroxidase-coupled goat anti-mouse or mouse anti-rabbit IgG secondary antibodies were used. Protein bands were detected using the Pierce ECL Western Blotting Substrate (Thermo Fisher Scientific, Rockford, IL, USA) in an automated x-ray film developer.
Immunohistochemistry
Sections of 4 μm from these tissue microarrays were stained with anti-HDAC1 (1:35) and anti-HDAC2 (1:2500). Immunostaining was performed on an automated Ventana Discovery XT staining platform (Tucson, AZ, USA) using the standard CC1 heat-induced epitope retrieval protocol (1 h at 95 °C). Discovery Universal Secondary Antibody (Tucson, AZ, USA), Universal DAB Detection Kit (Tucson, AZ, USA) and Gill's hematoxylin counterstain were employed for visualization.
Scoring
Immunohistochemical labeling was assessed with a × 10-magnification objective and was based on the percentage of positive nuclei staining above the background, as well as the staining intensity. Cores with less than 50 viable tumor cells were excluded. On the basis of the assessed percentage of positive nuclei (averaged across replicate cores), the following score categories were assigned: no visible staining=score 0; 1–50%=score 1; 51–75%=score 2; >75%=score 3. The intensity of staining was scored as 0 if no visible staining, 1 if weak/equivocal, 2 if moderate, or 3 if intense. When there was discrepancy in the intensity of staining among replicate cores, the higher intensity was used. The sum of these individual scores then defined the final score of each sample.
Statistical Analysis
R software version 2.11.1 was used for statistical analysis. Comparison of the distribution of the staining scores between groups (translocation-associated sarcomas vs sarcomas, and borderline tumors not associated with translocations vs normal tissues) was made using the Wilcoxon rank-sum test. GISTs were assessed as a separate category due to the large numbers of cases included. For comparing intensity of staining among groups, Pearson's χ2-test was used. P<0.05 was considered statistically significant.
Results
The antibodies used in this study were identified on the basis of their reported good performance for IHC analysis in FFPE tissue in prior publications, and according to the suppliers’ data sheets. HDAC1 and HDAC2 antibodies showed single bands of the expected molecular weights (60 kDa and 55 kDa, respectively) on western blot of sarcoma cell-line lysates. These bands were absent in the presence of blocking peptide, confirming antibody specificity. In contrast, protein immunoblots with commercial mouse monoclonal anti-HDAC3, rabbit monoclonal anti-HDAC3 and rabbit polyclonal anti-HDAC6 revealed extra bands, suggesting a reaction with multiple antigenic targets, so these reagents were not used in our subsequent immunohistochemical experiments.
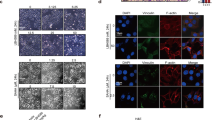
On formalin-fixed, paraffin-embedded tissue microarray specimens, HDAC1 and HDAC2 immunostaining was nuclear and readily interpretable in virtually all cases (Figures 1 and 2). HDAC1 staining in the cytoplasm was seen in some normal tissues such as pancreatic and hepatic parenchyma, small bowel and gastric mucosa, renal tubules, and trophoblast. No cytoplasmic staining for HDAC2 was seen in normal tissues. Also, 6 of the 10 tested cases of alveolar soft part sarcoma showed granular and globular cytoplasmic staining for HDAC1 (Figure 3). HDAC1 cytoplasmic immunoreactivity was seen in none of the other types of tumor in this study, nor was any cytoplasmic staining seen for HDAC2 in alveolar soft part sarcoma. Histograms showing the scoring distributions are presented as Figure 4, and results of all specimen types by diagnosis are shown in Table 1.
A total of 1196 mesenchymal tumors, representing 44 diagnostic entities were analyzed for HDAC1. Of this total, 210 were translocation-associated sarcomas and 986 were mesenchymal tumors not associated with known fusion transcription factor oncoproteins, including a set of 486 GIST. In addition, a total of 136 samples representing 39 different types of normal tissues were analyzed, including epithelial, mesenchymal and embryofetal tissues. The mean HDAC1 score among normal tissues was 1.8 (95% C.I=1.5–2.0). In this group, the mesenchymal tissues showed the lowest mean score (1.05) and the epithelial tissues showed the highest (1.94). The mean HDAC1 score among GISTs was 1.8 (C.I=1.6–2.0), among translocation-associated sarcomas was 0.8 (C.I=0.6–1.0), and among other mesenchymal tumors was 1.3 (C.I=1.1–1.4). This distribution of staining scores was not significantly different between normal tissues and GIST, but both of these categories had significantly higher staining scores than translocation-asociated sarcomas (P<10−7) or the group of other mesenchymal tumors (P=0.003).
A total of 1195 tumors were scored for HDAC2. The mean HDAC2 staining score was 3.9 (C.I=3.7–4.1) in normal tissues. In this group, the mesenchymal tissues showed the lowest mean score (3.37) and the embryofetal tissues the highest (4.31). The mean HDAC2 score in GIST was 3.4 (C.I=3.2–3.6), as compared with 4.5 (C.I=4.3–4.7) in translocation-associated sarcomas, and 4.1 (C.I=3.9–4.3) among other mesenchymal tumors. These staining scores were significantly higher in translocation-associated sarcomas (P<10−7) and other non-GIST mesenchymal tumors (P=0.003) compared with normal tissues. Indeed, the staining scores in translocation-associated sarcomas were also significantly higher than in the next highest scoring group, the other non-GIST mesenchymal tumors (P=0.01). GIST in contrast had the lowest scores for HDAC2, significantly lower than in normal tissues (P=0.01). Among the sarcomas associated with fusion transcription factors, all the individual diagnostic entities showed mean HDAC2 score values considerably higher than that for normal tissues. The only exception to this observation was low-grade fibromyxoid sarcoma, for which the mean score value was 2.3, much lower than in the rest of this group. The translocation-associated sarcomas with the highest mean staining scores for HDAC2 are alveolar rhabdomyosarcoma, synovial sarcoma, desmoplastic small round-cell tumor and myxoid liposarcoma. In the group of other mesenchymal tumors the entities that showed mean HDAC2 scores higher than that for normal tissues included embryonal rhabdomyosarcoma, neuroblastoma, epithelioid angiosarcoma, dermatofibrosarcoma protuberans with fibrosarcomatous differentiation, and myxofibrosarcoma (Table 1).
Staining intensity and percentage of positive-tumor cells were also assessed as separate parameters. The proportion of translocation-associated sarcomas with moderate or strong staining intensity for HDAC1, 4%, was lower than among normal tissues (15%; P=0.0003), GIST (12%; P=0.0010) and other mesenchymal tumors (11%; P=0.0021). In contrast, for HDAC2, the translocation-associated sarcomas exhibited moderate or strong staining intensity in 69% of the cases vs 46% of examined normal tissues (P<0.0001), 33% of GIST (P<0.0001) and 61% of other mesenchymal tumors (P=0.043). Similarly, the mean percentage of stained nuclei with HDAC1 antibody was 12% in translocation-associated sarcomas vs 28% in normal tissues (P<0.0001), 36% in GIST (P<0.0001) and 19% in other mesenchymal tumors (P<0.0001). The mean percentage of nuclei staining positively for HDAC2 was 86% in translocation-associated sarcomas vs 82% in normal tissues (P=0.035), 70% in GIST (P<0.0001) and 77% in other mesenchymal tumors (P=0.0026). Thus, findings for not only the combined immunohistochemical scores, but also their components of intensity and percentage of positive nuclei are broadly similar.
Discussion
Despite the wide use of HDAC inhibitors in cell culture, animal models and early-phase clinical trials, little is known about the expression of their targets in human tumor specimens.26 Most of the research on HDACs deals with functional aspects of this group of enzymes. Only recently have a few translational studies been published describing HDAC expression in different types of cancers (reviewed in Weichert27). To date, no study had described the in situ expression of different HDAC proteins in clinical samples of translocation-associated sarcomas, other than the work of Hrzenjak et al7 in endometrial stromal sarcomas, where it was shown that HDAC2 expression is consistently high.
In this study, we found that HDAC2 is present and intensely expressed in a high proportion of mesenchymal tumors, and most prominently, among the group of sarcomas associated with fusion transcription factors. The sole exception is low-grade fibromyxoid sarcoma, which although bearing a fusion oncoprotein (FUS-CREB3L2) similar in general structure to other fusion transcription factor-associated sarcomas, has a more well-differentiated histological appearance and a more indolent clinical behavior. These features are conceivably related to its lower level of HDAC expression and potentially more limited capacity to silence tumor suppressor genes.28 HDAC1, consistent with published literature,5 is less expressed than HDAC2 in normal tissues, and HDAC1 protein expression levels among mesenchymal tumors appear to be lower still.
High immunohistochemical expression of class I HDACs has been recently reported in epithelial neoplasms. In gastric carcinoma, HDAC1, HDAC2 and HDAC3 were found to be highly expressed in a subgroup of cases, and associated with nodal spread.29 In the same study, high expression of HDAC2 had independent negative prognostic significance for overall survival. In our study, the same HDAC3 antibody was tested, found unreliable (yielding multiple bands) and so, was excluded. In a study of colorectal cancer, a subset of cases (particularly highly proliferating and dedifferentiated tumors) showed high expression of class I HDAC isoforms, and HDAC2 expression was independently prognostic for diminished overall survival.30 In prostate cancer, class I HDAC isoforms are strongly expressed in the majority of cases, with high HDAC1, and two correlating with high Gleason grade, proliferative capacity and reduced disease free-survival.31, 32 High expression of class I HDACs has also been shown in most ovarian and endometrial carcinomas.33 In all these studies, the expression of class I HDAC isoforms showed a high degree of concordance with each other, suggesting a shared regulation. The marked difference in expression between HDAC1 and HDAC2 isoforms in our current study suggests that they might be regulated differently in mesenchymal tissues.
Although there is convincing evidence for the involvement of HDACs in the development of cancer, the specic roles of individual HDACs in the regulation of cell proliferation, apoptosis and the cell cycle are as yet unclear. Systematic investigations into the function of each HDAC isoform in tumor models are lacking, but some specific actions of HDAC2 have been reported in cell lines. For example, in endometrial stromal sarcoma ESS-1 cells transfected with siRNAs to specifically knock down HDAC2, the concentration of p21 increases in a time-dependent manner, supporting the concept that HDAC2 specifically contributes to cell cycle progression.7 Knockdown of HDAC2 in HeLa cells induces apoptosis associated with p53-independent p21 upregulation and cell differentiation.34 In MCF7 breast cancer cells, HDAC2 knockdown increases the DNA-binding activity of p53 associated with inhibition of proliferation and induction of cellular senescence.35 By analogy, a dependence on HDAC2 activity for cell cycle progression in translocation-associated sarcomas could contribute to their particular sensitivity to HDAC inhibitors.18
In summary, HDAC2, one of the HDAC isoforms and a therapeutic target for HDAC inhibitors, is present and intensely expressed in a very high proportion of mesenchymal tumors. Expression is highest among sarcomas associated with fusion transcription factors. These results are of particular interest, as a systematic evaluation of the expression of different HDAC isoforms has not been undertaken in this class of tumors before, and because HDAC inhibitors are currently promising agents for cancer therapy, being actively evaluated in clinical trials in several types of malignancies.36, 37 A phase II clinical trial of the hydroxamic acid-based HDAC inhibitor SB93938 in recurrent and metastatic translocation-associated sarcomas has been opened in Canada (NCIC-CTG IND.200). Also underway is a phase II study of vorinostat in German patients with advanced, metastatic soft tissue sarcoma (NCT00918489), and a phase II clinical trial of belinostat in combination with doxorubicin in soft tissue sarcomas is approaching completion in Denmark and the UK (NCT00878800). On the basis of our results, HDAC2 expression could be evaluated as a candidate predictive biomarker for therapeutic response to HDAC inhibitors in this particular group of neoplasia, as the determination of HDAC expression in tumor tissue is feasible even on small, formalin-fixed paraffin-embedded tissue cores.
References
Martens J, Stunnenberg H . The molecular signature of oncofusion proteins in acute myeloid leukemia. FEBS Lett 2010;584:2662–2669.
Lubieniecka JM, de Bruijn DR, Su L, et al. Histone deacetylase inhibitors reverse SS18-SSX-mediated polycomb silencing of the tumor suppressor early growth response 1 in synovial sarcoma. Cancer Res 2008;68:4303–4310.
Grignani F, De Matteis S, Nervi C, et al. Fusion proteins of the retinoic acid receptor-a recruit histone deacetylase in promyelocytic leukaemia. Nature 1998;391:815–817.
Wang J, Hoshino T, Redner R, et al. ETO, fusion partner in the (8; 21) acute myeloid leukemia, represses transcription by interaction with the human N-CoR/mSin3/HDAC1 complex. Proc Natl Acad Sci USA 1998;95:10860–10865.
De Ruijter A, Van Gennip A, Caron H, et al. Histone deacetylases (HDACs): characterization of the classical HDAC family. Biochem J 2003;370:737–749.
Li H, Ma X, Wang J, et al. Effects of rearrangement and allelic exclusion of JJAZ1/SUZ12 on cell proliferation and survival. Proc Natl Acad Sci USA 2007;104:20001–20006.
Hrzenjak A, Moinfar F, Kremser ML, et al. Valproate inhibition of histone deacetylase 2 affects differentiation and decreases proliferation of endometrial stromal sarcoma cells. Mol Cancer Ther 2006;5:2203–2210.
Hrzenjak A, Kremser M, Strohmeier B, et al. SAHA induces caspase independent, autophagic cell death of endometrial stromal sarcoma cells by influencing the mTOR pathway. J Pathol 2008;216:495–504.
Smith R, Owen L, Trem D, et al. Expression profiling of EWS/FLI identifies NKX2. 2 as a critical target gene in Ewing's sarcoma. Cancer Cell 2006;9:405–416.
Richter G, Plehm S, Fasan A, et al. EZH2 is a mediator of EWS/FLI1 driven tumor growth and metastasis blocking endothelial and neuro-ectodermal differentiation. Proc Natl Acad Sci USA 2009;106:5324–5329.
van der Vlag J, Otte A . Transcriptional repression mediated by the human polycomb-group protein EED involves histone deacetylation. Nat Genet 1999;23:474–478.
Varambally S, Dhanasekaran S, Zhou M, et al. The polycomb group protein EZH2 is involved in progression of prostate cancer. Nature 2002;419:624–629.
Owen L, Kowalewski A, Lessnick S . EWS/FLI mediates transcriptional repression via NKX2. 2 during oncogenic transformation in Ewing's sarcoma. PLoS One 2008;3:e1965.
Sakimura R, Tanaka K, Nakatani F, et al. Antitumor effects of histone deacetylase inhibitor on Ewing's family tumors. Int J Cancer 2005;116:784–792.
Jaboin J, Wild J, Hamidi H, et al. MS-27-275, an inhibitor of histone deacetylase, has marked in vitro and in vivo antitumor activity against pediatric solid tumors. Cancer Res 2002;62:6108–6115.
Sonnemann J, Dreyer L, Hartwig M, et al. Histone deacetylase inhibitors induce cell death and enhance the apoptosis-inducing activity of TRAIL in Ewinǵs sarcoma cells. J Cancer Res Clin Oncol 2007;133:847–858.
Ito T, Ouchida M, Morimoto Y, et al. Significant growth suppression of synovial sarcomas by the histone deacetylase inhibitor FK228 in vitro and in vivo. Cancer Lett 2005;224:311–319.
Liu S, Cheng H, Kwan W, et al. Histone deacetylase inhibitors induce growth arrest, apoptosis, and differentiation in clear cell sarcoma models. Mol Cancer Ther 2008;7:1751–1761.
Bolden J, Peart M, Johnstone R . Anticancer activities of histone deacetylase inhibitors. Nat Rev Drug Discov 2006;5:769–784.
Terry J, Saito T, Subramanian S, et al. TLE1 as a diagnostic immunohistochemical marker for synovial sarcoma emerging from gene expression profiling studies. Am J Surg Pathol 2007;31:240–246.
Nielsen TO, Hsu FD, O’Connell JX, et al. Tissue microarray validation of epidermal growth factor receptor and SALL2 in synovial sarcoma with comparison to tumors of similar histology. Am J Pathol 2003;163:1449–1456.
Ng TL, Gown AM, Barry TS, et al. Nuclear beta-catenin in mesenchymal tumors. Mod Pathol 2005;18:68–74.
Cheng H, Dodge J, Mehl E, et al. Validation of immature adipogenic status and identification of prognostic biomarkers in myxoid liposarcoma using tissue microarrays. Hum Pathol 2009;40:1244–1251.
Steigen SE, Straume B, Turbin D, et al. Clinicopathologic factors and nuclear morphometry as independent prognosticators in KIT-positive gastrointestinal stromal tumors. J Histochem Cytochem 2008;56:139–145.
West RB, Harvell J, Linn SC, et al. Apo D in soft tissue tumors: a novel marker for dermatofibrosarcoma protuberans. Am J Surg Pathol 2004;28:1063–1069.
Witt O, Deubzer H, Milde T, et al. HDAC family: what are the cancer relevant targets? Cancer Lett 2009;277:8–21.
Weichert W . HDAC expression and clinical prognosis in human malignancies. Cancer Lett 2009;280:168–176.
Su L, Cheng H, Sampaio AV, et al. EGR1 reactivation by histone deacetylase inhibitors promotes synovial sarcoma cell death through the PTEN tumor suppressor. Oncogene 2010;29:4352–4361.
Weichert W, Röske A, Gekeler V, et al. Association of patterns of class I histone deacetylase expression with patient prognosis in gastric cancer: a retrospective analysis. Lancet Oncol 2008;9:139–148.
Weichert W, Röske A, Niesporek S, et al. Class I histone deacetylase expression has independent prognostic impact in human colorectal cancer: specific role of class I histone deacetylases in vitro and in vivo. Clin Cancer Res 2008;14:1669–1677.
Song YH, Shiota M, Tamiya S, et al. The significance of strong histone deacetylase 1 expression in the progression of prostate cancer. Histopathology 2011;58:773–780.
Weichert W, Röske A, Gekeler V, et al. Histone deacetylases 1, 2 and 3 are highly expressed in prostate cancer and HDAC2 expression is associated with shorter PSA relapse time after radical prostatectomy. Br J Cancer 2008;98:604–610.
Weichert W, Denkert C, Noske A, et al. Expression of class I histone deacetylases indicates poor prognosis in endometrioid subtypes of ovarian and endometrial carcinomas. Neoplasia 2008;10:1021–1027.
Huang B, Laban M, Leung C, et al. Inhibition of histone deacetylase 2 increases apoptosis and p21Cip1/WAF1 expression, independent of histone deacetylase 1. Cell Death Differ 2005;12:395–404.
Harms K, Chen X . Histone deacetylase 2 modulates p53 transcriptional activities through regulation of p53-DNA binding activity. Cancer Res 2007;67:3145–3152.
Lane A, Chabner B . Histone deacetylase inhibitors in cancer therapy. J Clin Oncol 2009;27:5459–5468.
Batty N, Malouf G, Issa J . Histone deacetylase inhibitors as anti-neoplastic agents. Cancer Lett 2009;280:192–200.
Novotny-Diermayr V, Sangthongpitag K, Hu C, et al. SB939, a novel potent and orally active histone deacetylase inhibitor with high tumor exposure and efficacy in mouse models of colorectal cancer. Mol Cancer Ther 2010;9:642–652.
Acknowledgements
We thank Christine Chow and Dr Hongwei Cheng for their technical assistance with immunohistochemistry and western blotting, respectively; Samuel Leung and Dr Walton Green for their assistance with statistical analysis; and Dr Robert B West and Dr Matt van de Rijn for their collaboration in providing tissue microarrays for this study. This work is supported by an operating grant from the Canadian Cancer Society (Grant number 018355) and an unrestricted educational grant from Sanofi-aventis Canada. Torsten Nielsen is a senior scholar of the Michael Smith Health Research Foundation. Dr Pacheco was supported by the Terry Fox Foundation STIHR in Molecular Oncologic Pathology at CIHR.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Pacheco, M., Nielsen, T. Histone deacetylase 1 and 2 in mesenchymal tumors. Mod Pathol 25, 222–230 (2012). https://doi.org/10.1038/modpathol.2011.157
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2011.157
Keywords
This article is cited by
-
Upregulation of ERK-EGR1-heparanase axis by HDAC inhibitors provides targets for rational therapeutic intervention in synovial sarcoma
Journal of Experimental & Clinical Cancer Research (2021)
-
Evaluation of glucocorticoid-induced TNF receptor (GITR) expression in breast cancer and across multiple tumor types
Modern Pathology (2020)
-
Expression of lymphocyte immunoregulatory biomarkers in bone and soft-tissue sarcomas
Modern Pathology (2019)
-
Establishment and characterization of a new human myxoid liposarcoma cell line (DL-221) with the FUS-DDIT3 translocation
Laboratory Investigation (2016)
-
NY-ESO-1 (CTAG1B) expression in mesenchymal tumors
Modern Pathology (2015)