Abstract
Homozygous deletion of CDKN2A (p16) is one of the most common genetic alterations in pleural mesotheliomas, occurring in up to 74% of cases. MTAP resides in the same gene cluster of the 9p21 region and is co-deleted in the majority of CDKN2A deleted cases. This study examines the genetic alterations in peritoneal mesotheliomas, which may have a different pathogenesis than their pleural counterparts. Twenty-six cases of peritoneal mesotheliomas in a triplicate tissue microarray were studied. Dual-color fluorescence in situ hybridization was performed with CDKN2A and MTAP locus-specific probes. Nine of 26 (35%) peritoneal mesotheliomas had homozygous deletion of CDKN2A; MTAP was co-deleted in every case. All cases with CDKN2A deletions had loss of p16 protein expression; five cases had loss of p16 protein without evidence of CDKN2A deletions. All patients with CDKN2A deletions were men (P, NS) and were significantly older (mean, 63 years) than the patients with no deletions (mean, 52 years) (P=0.033, t-test). An association with asbestos exposure could not be proved in this study. Similar to pleural mesotheliomas, patients with CDKN2A deletions and loss of p16 protein expression had worse overall and disease-specific survival (P=0.010 and 0.006, respectively; Kaplan–Meier log rank). Detection of CDKN2A-MTAP co-deletion in peritoneal mesotheliomas, coupled with a p16 immunohistochemical stain as an inexpensive screening tool, can help identify those patients who may have an unfavorable outcome after aggressive cytoreductive surgery combined with hyperthermic intraperitoneal chemotherapy and those who may respond to targeted therapy of the MTAP pathway.
Similar content being viewed by others
Main
Malignant mesotheliomas are aggressive tumors that arise from serosal surfaces. The median survival of untreated cases ranges between 6 and 18 months.1 The majority arise from the pleura, but about 10–20% of the cases in the Unites States arise from the peritoneal surfaces.1, 2 The median age at diagnosis is 65–69 years for peritoneal mesotheliomas, which is slightly younger than the median age at diagnosis of 72 years for pleural mesotheliomas.1, 2 Interestingly, there is a gender difference between the two sites. Pleural mesotheliomas occur primarily in men (at about a ratio of 5:1 men to women), whereas only about 55% of the cases of peritoneal mesotheliomas occur in men.1, 2 Furthermore, peritoneal mesotheliomas account for 7% of all mesotheliomas in men, but 17% of all mesotheliomas in women.3 Although peritoneal mesotheliomas are associated with asbestos exposure,4 the association is not as strong as with their pleural counterparts. This may be related to the observation that cases of peritoneal mesothelioma that occur in women are not associated with asbestos exposure.
Without aggressive therapy, diffuse peritoneal mesotheliomas are, in general, rapidly fatal malignancies with a median survival of about 1 year.5, 6, 7, 8 With aggressive cytoreductive surgery (CS) combined with hyperthermic intraperitoneal chemotherapy (HIPEC), recent studies have reported survival rates that approach 5 years.8, 9, 10 Owing to major morbidly after such aggressive therapy, patient selection has a critical role in a successful outcome and improved survival.6, 11 A preoperative prognostic classification for diffuse peritoneal mesotheliomas has been proposed and involves the following parameters: (1) presence of extra-abdominal and/or hepatic metastases; (2) disease potentially suitable for complete surgical cytoreduction at preoperative imaging studies; and (3) prognostic factors, including histologic subtype, nuclear grade and mitotic count.6 As only a few studies have investigated potential biological and pathological prognostic factors for patients with diffuse peritoneal mesotheliomas treated by CS and perioperative intraperitoneal chemotherapy,9, 10, 12 additional studies are warranted.
Cytogenetic and molecular changes in malignant mesotheliomas have been well studied, primarily in pleural mesotheliomas.13 Homozygous deletion of CDKN2A (p16) and co-deletion of MTAP are two of the most common genetic alterations. Both genes reside in the same gene cluster of the 9p21 locus. CDKN2A is deleted in 60–75% of primary tumors14, 15, 16, 17 and up to 100% of cell lines.18, 19 The detection of homozygous deletion of CDKN2A by fluorescence in situ hybridization (FISH) can be a useful diagnostic tool to distinguish between malignant mesotheliomas and reactive mesothelial proliferations,14, 20 and as a prognostic indicator of a poor outcome in pleural mesotheliomas.17, 21, 22 In a previous study from our institution that examined the diagnostic utility of CDKN2A deletions in malignant mesotheliomas, a subset of cases were from the peritoneum and only 5 of the 20 (25%) cases harbored CDKN2A deletions.14 This is the only study that has examined CDKN2A deletions in peritoneal mesotheliomas. This difference in CDKN2A deletions between pleural and peritoneal mesotheliomas prompted this study on an additional cohort of peritoneal mesotheliomas.
Loss of p16 protein expression has also been shown to predict poor survival in patients with malignant peritoneal mesothelioma.23 In pleural mesotheliomas, loss of p16 protein expression is typically associated with CDKN2A deletion, but a subset of cases showed loss of p16 due to inactivation of the gene by methylation.24, 25, 26 The correlation between CDKN2A deletion, loss of p16 protein expression and survival has not been made in peritoneal mesotheliomas.
MTAP encodes methylthioadenosine phosphorylase, an enzyme essential in the salvage of cellular adenine and methionine. MTAP is co-deleted in the majority of CDKN2A deleted cases and all cases with deletion of MTAP also have deletion of CDKN2A.15, 21 Mesotheliomas that show loss of MTAP could be treated with targeted therapy using inhibitors of the AMP synthesis pathway.21 However, deletion of MTAP has not been studied in peritoneal mesotheliomas.
The aim of this study was to determine the prevalence of homozygous deletion of CDKN2A and/or MTAP in a series of peritoneal mesotheliomas, to correlate CDKN2A deletions by FISH with loss of protein expression by IHC and to determine the prognostic significance of CDKN2A deletions in this study population.
Materials and methods
All patients with malignant peritoneal mesotheliomas who underwent CS combined with HIPEC between 2001 and 2007 at the University of Pittsburgh Medical Center were reviewed and the 26 cases with available archived tissue were included in the study (eight of these patients were included in our earlier study14). The diagnosis of malignant mesothelioma was based on the combination of clinicoradiographic findings, intraoperative observations and routine H&E histology. The diagnosis was confirmed by immunohistochemistry using routine antibodies such as calretinin, WT-1, D2-40, cytokeratin 5/6, Ber-EP4, B72.3, p63, CEA. This study was approved by the University of Pittsburgh Institutional Review Board (IRB #PRO08080411).
Tissue Microarray Constructions
A high-density tissue microarray was constructed from archival formalin-fixed, paraffin-embedded samples. For each sample, three representative areas rich in tumor cells were identified by light microscopic examination and marked on the H&E sections. Three cores measuring 0.1 cm in diameter were taken from the donor paraffin tissue blocks of each case and were arranged in a recipient paraffin tissue array block by using a Manual Tissue Arrayer (MTA-1, Beecher Instruments, Sun Prairie, WI, USA).
Fluorescence In Situ Hybridization
Dual-color FISH was performed as previously described using a Spectrum Green-labeled chromosome 9 centromeric (CEP9) probe and either a CDKN2A (Abbott Molecular, Des Plains, IL, USA) and/or MTAP (RP11-70L8, CHORI, Oakland, CA, USA).14 In brief, paraffin sections of the tissue microarray were de-paraffinized, dehydrated in ethanol and air-dried. The slides were digested with protease K (0.5 mg/ml) at 37°C for 28 min, denatured at 75°C for 5 min in 70% Formamide (Chemicon, Billerica, MA, USA) and dehydrated in ethanol. The probes were denatured for 5 min at 75°C before hybridization. Slides were hybridized overnight at 37°C and washed in 2 × SSC/0.3% Igepal (Sigma, St Louis, MO, USA) at 72°C for 2 min. Nuclei were counterstained with DAPI/antifade (Vysis, Downers Grove, IL, USA). Each FISH assay included positive and negative controls. Analyses were performed using a fluorescence microscope (Nikon Eclipse E600) and Cytovysion Workstation (Applied Imaging, Santa Clara, CA, USA) equipped with filter sets with single- and dual-band excitors for Spectrum Green, Spectrum Orange and DAPI (UV 360 nm). Each core on the tissue microarray was identified and only individual and well-delineated cells were scored; overlapping cells were excluded from the analysis. At least 60 cells were scored for each case and control. Each tumor was assessed by the average and the maximum numbers of copies of the gene (CDKN2A or MTAP) per cell and the average ratio of the gene to CEP9 copy numbers. Homozygous deletion was defined by loss of both gene signals in at least 20% of nuclei that showed at least one signal for the CEP9 probe.14
Immunohistochemistry
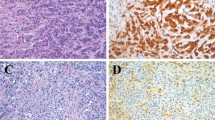
Immunohistochemistry was performed using anti-p16 mouse monoclonal antibody at a dilution of 1:200 (BD PharMingen, San Diego, CA, USA) according to the standard avidin–biotin–peroxidase complex method. The p16 immunostain for each case was interpreted as previously described.23 Briefly, nuclear and cytoplasmic p16 immunoreactivity was scored for intensity and extent. Intensity of immunoreactivity was scored as 0, no staining; 1+, weak staining; 2+, strong staining. The extent of immunoreactivity was assessed as focal (<5%) or multifocal/diffuse. Cases were considered negative if scored as 0 or 1+ and focal; all 2+ cases and cases scored as 1+ and multifocal/diffuse were considered positive (Figure 1). Sections of cervical biopsy previously identified as immunoreactive for p16 protein served as a positive control. Negative controls included omitting the primary antibody and its substitution with normal serum.
Statistical Analysis
For the comparisons of means, a t-test for independent variables was used for continuous data, and Fisher's exact test was used for categorical data. For median survival data, Kaplan–Meier survival analysis was performed using a log-rank statistical method.
Results
Of the 26 patients included in this study, 22 were males (M/F ratio 5.5:1) and the mean age at the time of surgery was 56 years (range 23–76 years). Seven patients had a known history of asbestos exposure; all were men with a mean age of 60 years (range, 37–69 years). All patients underwent CS combined with HIPEC.27 At the completion of CS, the amount of residual disease was recorded as follows: R0 (complete) resection was achieved in 6 (23%) of cases, R1 (remaining peritoneal implants <5 mm) in 12 (46%) and R2 (remaining peritoneal implants >5 mm) in 8 (31%). Median time from tumor diagnosis to CS-HIPEC procedure was 254 days (range, 5–1894 days). Median follow-up after surgery for this cohort of patients was 450 days (mean, 635 days; range, 15–2647 days); median follow-up after diagnosis was 553 days (mean, 885 days; range, 46–2788 days). Mortality in this cohort was 69%; 10 patients (38%) died from their mesothelioma; 3 (11%) died from late complications of surgery (renal, respiratory and/or hepatic failure) and all 3 patients had gross residual disease; 5 (19%) with residual/recurrent disease were lost to follow-up and died; and 1 patient was lost to follow-up and died. The high mortality rate from late complications of surgery is not reflective of our overall results with the HIPEC procedure27 or our previous results with mesothelioma patients,12 and we believe represents a random anomaly within this subgroup. Of the five patients who are still alive, three are free of disease and three are alive with disease. One patient was lost to follow-up soon after surgery. Histopathologically, 25 (96%) of the tumors were classified as epithelioid type (Figure 1a and c); only one tumor was a sarcomatoid mesothelioma.
The results of CDKN2A and MTAP deletions assessed by FISH are shown in Table 1. FISH analysis demonstrated homozygous deletion of CDKN2A in 9 of 26 cases (35%) of peritoneal mesotheliomas (Figure 2b). The one sarcomatoid mesotheliomas in this study did have the homozygous deletion of CDKN2A. All cases showed homozygous deletion in >89% of the analyzed cells (mean, 96.8%; range, 89.5–100%). Hemizygous loss was not identified in the 16 cases that were analyzed.
CDKN2A and MTAP deletions by FISH in peritoneal mesothelioma. (a) Normal cells are expected to show two green signals (chromosome 9 centromeric probe) and two red signals (locus-specific probes to either CDKN2A (shown) or MTAP (not shown)). (b) Homozygous deletion of CDKN2A (loss of both red signals in at least 20% of nuclei). (c) Homozygous deletion of MTAP in the same case.
MTAP showed homozygous deletion in nine cases (Figure 2c), the same nine cases that also had CDKN2A deletions. Homozygous deletions were present in 85.3–100% of the analyzed cells (mean, 93.2%). The correlation between MTAP and CDKN2A deletions was 100%. The nine patients with CDKN2A-MTAP deletions were significantly older (mean, 63 years) than the patients with no deletions (mean, 52 years) (P=0.033, t-test), and although all nine patients were male, this did not reach statistical significance. Asbestos exposure did not correlate with CDKN2A-MTAP deletions (four of the seven men with asbestos exposure had CDKN2A-MTAP deletions, which did not reach statistical significance).
By immunohistochemistry, 14 cases (54%) showed loss of p16 expression (Figure 1d) and p16 loss was strongly associated with CDKN2A-MTAP deletions. All nine cases with CDKN2A-MTAP deletions showed loss of the p16 protein, whereas only 5 of the 17 non-deleted cases showed p16 protein loss (P=0.0007).
Patients with CDKN2A-MTAP deletions had worse overall survival, both from the time of surgery at our institution and from the time of initial diagnosis (P=0.010 for both). Patients with CDKN2A-MTAP deletions also had worse disease-specific survival, both from the time of surgery at our institution and from the time of initial diagnosis (P=0.006 for both). Survival curves from time of surgery are shown in Figure 3. All nine patients with CDKN2A-MTAP deletions died with disease (six died of the disease, two who had residual disease were lost to follow-up and died, and one patient with gross residual disease died of renal and hepatic failure 2 months after surgery). The amount of post-operative residual disease did not correlate with overall survival or disease-specific survival in this study (P=NS). Patients with evidence of p16 protein loss by IHC also had worse overall survival, both from time of diagnosis (P=0.010) and from time of surgery (P=0.044), and worse disease-specific survival from time of diagnosis (P=0.038) with a trend toward a worse prognosis from time of surgery (P=0.066) (graphs not shown). Of the five patients with p16 protein loss but no evidence of CDKN2A-MTAP deletions, two had evidence of disease but were lost to follow-up and died, one died of the disease, one is alive with no evidence of disease 1138 days after surgery, and one was lost to follow-up.
Discussion
CDKN2A deletions were found in 35% of peritoneal mesotheliomas in our series. Compared with earlier studies that used FISH, this percentage is lower than the 60–74% deletion rate seen in pleural mesotheliomas14, 15, 17 and similar to a different cohort of peritoneal mesotheliomas previously reported by us (25%).14 It is unclear why this difference in CDKN2A deletions exists between pleural and peritoneal mesotheliomas. It is possible that CDKN2A deletions are associated with increasing age. In our study, patients with CDKN2A deletions were significantly older (mean age of 63 years compared with patients with non-deleted tumors with a mean age of 52 years). And, in general, patients with pleural mesotheliomas are older than patients with peritoneal disease.1, 2 It is also possible that CDKN2A deletions are associated with gender. All of our cases with CDKN2A deletions occurred in men, but due to the small number of women in this cohort, this did not reach statistical significance. In general, women with peritoneal mesotheliomas appear to have a better survival.28 In a recent study of pleural mesotheliomas, gender and FISH result predicted survival. In that study, patients who did not have CDKN2A deletions were more frequently women who survived more than 3 years.17 And finally, it is possible that CDKN2A deletions are associated with asbestos exposure. On the basis of our study, there does not appear to be an association between asbestos exposure and CDKN2A deletions in peritoneal mesotheliomas as four of the seven patients with known asbestos exposure were found to have CDKN2A deletions. However, the small size of the cohort and the difficulty in determining an exposure history in this retrospective study limit this assessment.
Owing to its proximity to CDKN2A in the 9p21 locus, MTAP deletions have also been studied in pleural mesotheliomas. Using FISH, Illei et al15 found MTAP co-deletions in 91% of pleural mesotheliomas that harbored CDKN2A deletions and that all MTAP deletions occurred with CDKN2A deletions. In our series of peritoneal mesotheliomas, there was 100% correlation between CDKN2A and MTAP deletions: all nine cases with CDKN2A deletions also had MTAP deletions and all MTAP deletions occurred in cases with CDKN2A deletions. Our study further supports the finding in other tumors that MTAP is only deleted when CDKN2A is deleted.15, 29, 30, 31
Tumors with MTAP deletions are potentially susceptible to inhibitors of de novo purine synthesis. Normal cells can synthesize adenine both de novo and through the salvage pathway where cleavage of methylthioadenosine (MTA) by MTAP produces adenosine triphosphate. As MTAP-deficient tumors are dependent on de novo purine synthesis, inhibitors of de novo adenine synthesis, such as L-alanosine, could target these tumors. Unfortunately, one Phase II clinical trial that treated MTAP-deficient tumors (detected by IHC) with L-alanosine was ineffective in patients with advanced MTAP-deficient tumors.32 However, other strategies have been proposed. One strategy involves administering both MTA and a toxic adenine analog, such as 2,6-diaminopurine, 6-methylpurine, or 2-fluoroadenine, or possibly even 5-fluorouracil or 6-thioguanine. In MTAP-positive cells, abundant adenine is generated from the supplied MTA, which competitively blocks the conversion of the analog to its active nucleotide form. In MTAP-negative tumor cells, the supplied MTA cannot generate adenine and hence cannot block conversion of the analog to its active form.33
In addition to detecting CDKN2A deletions by FISH, we also performed IHC for the p16 protein using a tissue microarray. More of our cases had loss of p16 by IHC (54%) than loss of CDKN2A by FISH (25%), but all cases with CDKN2A deletions showed loss of p16 protein. This percentage of p16 loss is similar to a previous report that found p16 loss in 48% of peritoneal mesotheliomas.23 A slightly higher percentage (61%) of p16 protein loss was reported in a series of pleural mesotheliomas, and loss of p16 protein in this study was strongly associated with CDKN2A deletions.17 It is possible that loss of p16 protein expression without homozygous loss of the CDKN2A gene by FISH is due to methylation of the CDKN2A gene. Similar to our findings, a recent study of pleural mesotheliomas showed that all 21 cases with CDKN2A deletion had loss of p16 expression by IHC, and that two of the three cases with p16 loss without deletion showed CDKN2A gene methylation.25 This and other studies have shown that, at least in a small percentage of pleural mesotheliomas (11–21%), the CDKN2A gene is inactivated by methylation.24, 25, 26 Although we did not test for aberrant methylation of the CDKN2A gene, a similar percentage of our peritoneal mesothelioma cases (19%) showed loss of p16 expression without loss of the CDKN2A gene.
One of the most striking observations in this study is the strong correlation of CDKN2A-MTAP deletions with a poor prognosis. There was also a correlation of p16 protein loss and a poor prognosis, but the association was not as strong. Our study is in agreement with earlier publications that have associated p16 abnormalities with poor outcome in mesotheliomas. One publication that studied peritoneal mesotheliomas reported that p16 protein loss (assessed by IHC) is associated with increased risk of death. Studies on pleural mesotheliomas have also documented the prognostic significance of CDKN2A deletions17, 22 and loss of p16 protein.17 In fact, the study by Dacic et al17 showed that cases with positive (intact) p16 protein expression as assessed by IHC and no evidence of CDKN2A deletions as determined by FISH had the best survival time.
If left untreated, diffuse malignant mesothelioma is a rapidly lethal disease. Although aggressive therapy that includes CS combined with HIPEC can significantly improve survival in these patients,8, 9, 10 it comes at a cost of major morbidity. Hence, proper patient selection is crucial for both successful outcome and improved survival.6, 11 Prognostic factors, such as histologic subtype, nuclear grade and mitotic count, are included in a preoperative prognostic classification system for diffuse peritoneal mesotheliomas.6 However, these traditional pathologic variables have either not been well studied or have shown mixed results as predictors of survival.9, 10, 12, 23, 34, 35 In addition to finding strong and useful predictors of survival, new target for therapy are needed.
Our study demonstrates that detection of CDKN2A deletions by FISH is a very strong predictor of survival in patients with peritoneal mesotheliomas. Loss of p16 protein as assessed by IHC is also a predictor of a poor outcome. It appears that, at least for peritoneal mesotheliomas, IHC for p16 could be used as a first-line test for CDKN2A deletions, but loss of p16 staining would have to be verified by FISH to determine those patients at highest risk of a poor outcome. Our study also demonstrates that CDKN2A deletions are strongly associated with MTAP deletions (100% correlation). These tests could easily be incorporated into the preoperative prognostic classification system for diffuse peritoneal mesotheliomas and could also be used to include patients into new novel-targeted chemotherapeutic protocols.
References
Tsao AS, Wistuba I, Roth JA, et al. Malignant pleural mesothelioma. J Clin Oncol 2009;27:2081–2090.
Hesdorffer ME, Chabot J, DeRosa C, et al. Peritoneal mesothelioma. Curr Treat Options Oncol 2008;9:180–190.
Price B, Ware A . Mesothelioma trends in the United States: an update based on Surveillance, Epidemiology, and End Results Program data for 1973 through 2003. Am J Epidemiol 2004;159:107–112.
Browne K, Smither WJ . Asbestos-related mesothelioma: factors discriminating between pleural and peritoneal sites. Br J Ind Med 1983;40:145–152.
Antman KH, Pomfret EA, Aisner J, et al. Peritoneal mesothelioma: natural history and response to chemotherapy. J Clin Oncol 1983;1:386–391.
Deraco M, Bartlett D, Kusamura S, et al. Consensus statement on peritoneal mesothelioma. J Surg Oncol 2008;98:268–272.
Eltabbakh GH, Piver MS, Hempling RE, et al. Clinical picture, response to therapy, and survival of women with diffuse malignant peritoneal mesothelioma. J Surg Oncol 1999;70:6–12.
Sugarbaker PH, Yan TD, Stuart OA, et al. Comprehensive management of diffuse malignant peritoneal mesothelioma. Eur J Surg Oncol 2006;32:686–691.
Deraco M, Nonaka D, Baratti D, et al. Prognostic analysis of clinicopathologic factors in 49 patients with diffuse malignant peritoneal mesothelioma treated with cytoreductive surgery and intraperitoneal hyperthermic perfusion. Ann Surg Oncol 2006;13:229–237.
Yan TD, Brun EA, Cerruto CA, et al. Prognostic indicators for patients undergoing cytoreductive surgery and perioperative intraperitoneal chemotherapy for diffuse malignant peritoneal mesothelioma. Ann Surg Oncol 2007;14:41–49.
Glockzin G, Schlitt HJ, Piso P . Peritoneal carcinomatosis: patients selection, perioperative complications and quality of life related to cytoreductive surgery and hyperthermic intraperitoneal chemotherapy. World J Surg Oncol 2009;7:5.
Feldman AL, Libutti SK, Pingpank JF, et al. Analysis of factors associated with outcome in patients with malignant peritoneal mesothelioma undergoing surgical debulking and intraperitoneal chemotherapy. J Clin Oncol 2003;21:4560–4567.
Musti M, Kettunen E, Dragonieri S, et al. Cytogenetic and molecular genetic changes in malignant mesothelioma. Cancer Genet Cytogenet 2006;170:9–15.
Chiosea S, Krasinskas A, Cagle PT, et al. Diagnostic importance of 9p21 homozygous deletion in malignant mesotheliomas. Mod Pathol 2008;21:742–747.
Illei PB, Rusch VW, Zakowski MF, et al. Homozygous deletion of CDKN2A and codeletion of the methylthioadenosine phosphorylase gene in the majority of pleural mesotheliomas. Clin Cancer Res 2003;9:2108–2113.
Xio S, Li D, Vijg J, et al. Codeletion of p15 and p16 in primary malignant mesothelioma. Oncogene 1995;11:511–515.
Dacic S, Kothmaier H, Land S, et al. Prognostic significance of p16/cdkn2a loss in pleural malignant mesotheliomas. Virchows Arch 2008;453:627–635.
Cheng JQ, Jhanwar SC, Klein WM, et al. p16 alterations and deletion mapping of 9p21-p22 in malignant mesothelioma. Cancer Res 1994;54:5547–5551.
Prins JB, Williamson KA, Kamp MM, et al. The gene for the cyclin-dependent-kinase-4 inhibitor, CDKN2A, is preferentially deleted in malignant mesothelioma. Int J Cancer 1998;75:649–653.
Illei PB, Ladanyi M, Rusch VW, et al. The use of CDKN2A deletion as a diagnostic marker for malignant mesothelioma in body cavity effusions. Cancer 2003;99:51–56.
Ladanyi M . Implications of P16/CDKN2A deletion in pleural mesotheliomas. Lung Cancer 2005;49 (Suppl 1):S95–S98.
Lopez-Rios F, Chuai S, Flores R, et al. Global gene expression profiling of pleural mesotheliomas: overexpression of aurora kinases and P16/CDKN2A deletion as prognostic factors and critical evaluation of microarray-based prognostic prediction. Cancer Res 2006;66:2970–2979.
Borczuk AC, Taub RN, Hesdorffer M, et al. P16 loss and mitotic activity predict poor survival in patients with peritoneal malignant mesothelioma. Clin Cancer Res 2005;11:3303–3308.
Destro A, Ceresoli GL, Baryshnikova E, et al. Gene methylation in pleural mesothelioma: correlations with clinico-pathological features and patient's follow-up. Lung Cancer 2008;59:369–376.
Kobayashi N, Toyooka S, Yanai H, et al. Frequent p16 inactivation by homozygous deletion or methylation is associated with a poor prognosis in Japanese patients with pleural mesothelioma. Lung Cancer 2008;62:120–125.
Wong L, Zhou J, Anderson D, et al. Inactivation of p16INK4a expression in malignant mesothelioma by methylation. Lung Cancer 2002;38:131–136.
Gusani NJ, Cho SW, Colovos C, et al. Aggressive surgical management of peritoneal carcinomatosis with low mortality in a high-volume tertiary cancer center. Ann Surg Oncol 2008;15:754–763.
Yan TD, Popa E, Brun EA, et al. Sex difference in diffuse malignant peritoneal mesothelioma. Br J Surg 2006;93:1536–1542.
Della Ragione F, Russo G, Oliva A, et al. 5′-Deoxy-5′-methylthioadenosine phosphorylase and p16INK4 deficiency in multiple tumor cell lines. Oncogene 1995;10:827–833.
Stadler WM, Olopade OI . The 9p21 region in bladder cancer cell lines: large homozygous deletion inactivate the CDKN2, CDKN2B and MTAP genes. Urol Res 1996;24:239–244.
Zhang H, Chen ZH, Savarese TM . Codeletion of the genes for p16INK4, methylthioadenosine phosphorylase, interferon-alpha1, interferon-beta1, and other 9p21 markers in human malignant cell lines. Cancer Genet Cytogenet 1996;86:22–28.
Kindler HL, Burris III HA, Sandler AB, et al. A phase II multicenter study of L-alanosine, a potent inhibitor of adenine biosynthesis, in patients with MTAP-deficient cancer. Invest New Drugs 2009;27:75–81.
Lubin M, Lubin A . Selective killing of tumors deficient in methylthioadenosine phosphorylase: a novel strategy. PLoS One 2009;4:e5735.
Goldblum J, Hart WR . Localized and diffuse mesotheliomas of the genital tract and peritoneum in women. A clinicopathologic study of nineteen true mesothelial neoplasms, other than adenomatoid tumors, multicystic mesotheliomas, and localized fibrous tumors. Am J Surg Pathol 1995;19:1124–1137.
Kerrigan SA, Turnnir RT, Clement PB, et al. Diffuse malignant epithelial mesotheliomas of the peritoneum in women: a clinicopathologic study of 25 patients. Cancer 2002;94:378–385.
Author information
Authors and Affiliations
Corresponding author
Additional information
Disclosure/conflict of interest
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Krasinskas, A., Bartlett, D., Cieply, K. et al. CDKN2A and MTAP deletions in peritoneal mesotheliomas are correlated with loss of p16 protein expression and poor survival. Mod Pathol 23, 531–538 (2010). https://doi.org/10.1038/modpathol.2009.186
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2009.186
Keywords
This article is cited by
-
MTAP immunohistochemistry is an accurate and reproducible surrogate for CDKN2A fluorescence in situ hybridization in diagnosis of malignant pleural mesothelioma
Modern Pathology (2020)
-
Molecular characterization of diffuse malignant peritoneal mesothelioma
Modern Pathology (2020)
-
Malignant mesothelioma in situ: morphologic features and clinical outcome
Modern Pathology (2020)
-
5-hydroxymethylcytosine but not MTAP methylation status can stratify malignant pleural mesothelioma based on the lineage of origin
Multidisciplinary Respiratory Medicine (2018)
-
The prognostic significance of BAP1, NF2, and CDKN2A in malignant peritoneal mesothelioma
Modern Pathology (2016)