Abstract
We propose a novel on-chip 3D cell rotation method based on a vibration-induced flow. When circular vibration is applied to a microchip with micropillar patterns, a highly localized whirling flow is induced around the micropillars. The direction and velocity of this flow can be controlled by changing the direction and amplitude of the applied vibration. Furthermore, this flow can be induced on an open chip structure. In this study, we adopted a microchip with three micropillars arranged in a triangular configuration and an xyz piezoelectric actuator to apply the circular vibration. At the centre of the micropillars, the interference of the vibration-induced flows originating from the individual micropillars induces rotational flow. Consequently, a biological cell placed at this centre rotates under the influence of the flow. Under three-plane circular vibrations in the xy, xz or yz plane, the cell can rotate in both the focal and vertical planes of the microscope. Applying this 3D cell rotation method, we measured the rotational speeds of mouse oocytes in the focal and vertical planes as 63.7 ± 4.0° s−1 and 3.5 ± 2.1° s−1, respectively. Furthermore, we demonstrated the transportation and rotation of the mouse oocytes and re-positioned their nuclei into a position observable by microscope.
Similar content being viewed by others
Introduction
Cell rotation is a fundamental technique in modern bioscience. For example, it enables three-dimensional (3D) observation of cellular structures and the 3D manipulations required in cellular surgery. Cell rotation is necessary to properly orient the target cell in the focal plane of the microscope for 3D observations. Such orientation control is especially important for observing non-uniform 3D cellular structures under the microscope. In addition, 3D cell manipulation is required for the removal or transfer of nuclei in transgenic animal research and for quality control purposes in the livestock industry1,2. Because the nucleus of an oocyte is anisotropically located at the cell edge, it should be re-positioned to an observable location in the focal plane during cellular surgery. Therefore, the cell rotation technique plays an important role in 3D cell observation and manipulation and has become a powerful tool.
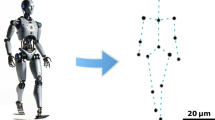
Cell rotation involves two essential motions: rotation in the focal plane (xy plane) and rotation in the plane perpendicular to the focal plane (xz or yz plane). These motions, illustrated in Figure 1a, are referred to as focal plane rotation (FPR) and vertical plane rotation (VPR), respectively. Conventionally, both motions are performed using dual micromanipulators3,4. One micromanipulator holds the target cell, while the other applies a torque for the rotation. FPR and VPR are achieved via the 3D operation of the micromanipulators under a microscope. However, this operation requires a complex system set-up and expert operational skills, which reduces its throughput and repeatability.
Cell rotation based on vibration-induced flow: (a) Schematic of vertical plane rotation (VPR). (b) Schematic of focal plane rotation (FPR). (c) Local flow around a single micropillar induced by rectilinear vibration. (d) Local whirling flow around a single micropillar induced by circular vibration. (e) Schematic of cell rotation based on vibration-induced flow. (f) Schematic of system set-up in the proposed method. (g) Schematic of FPR. (h) Schematic of VPR.
With recent progress in microtechnology, microfluidic chips have emerged as promising cell rotation devices. The on-chip cell rotation technique integrates rotation mechanisms into the chip, thereby realizing high throughput and a highly repeatable, low-cost process. On-chip cell rotation studies have reported the use of electric, magnetic and optical forces5–14. In the electrorotation technique, the torque is induced by applying a dielectric force on the target cell. FPR by electrorotation is easily achieved by patterning multiple electrodes in a two-dimensional (2D) configuration, but VPR requires multiple electrodes in a 3D configuration. This 3D structure is realized by patterning the electrodes on a substrate layer and a cover layer; the layers are then bonded10. However, this closed chip structure is unsuitable for applications such as nuclear treatments, which require that external probes access the rotation point. An open-type cell rotation chip has been recently constructed from four vertical wall electrodes and two bottom electrodes11, but the fabrication process remains complex and requires multiple wires that connect to each electrode on the chip. Cell rotation via a magnetic force is achieved by inserting magnetic nanoparticles into the target cells and subjecting the cells to magnetic fields12. The magnetic fields are supplied by multiple Helmholtz coils placed outside the chip. However, these coils require a complex 3D arrangement to achieve both FPR and VPR. Alternatively, optical forces may supply the torque for cell rotation; that is, different points on the target cell may be irradiated with dual lasers13 or a rod-shaped target cell may be laser-scanned along its longitudinal axis14. Although optical forces successfully rotate small cells (≈10 μm), they generate insufficient torque to rotate large cells, such as oocytes (≈100 μm). Furthermore, like their electrically and magnetically driven counterparts, optical systems that realize both FPR and VPR require a complex system configuration.
Thus, we aim to achieve 3D cell rotation (FPR and VPR) on a simply configured open structure chip. To this end, we propose a novel on-chip cell rotation method based on vibration-induced local flow. The substrate surface of the chip is embellished with micropillar patterns, around which local whirling flows are induced by applying circular vibrations to the chip (see Figure 1b). The triangular configuration of the micropillars achieves stable cell rotation (Figure 1d).
The advantages of the proposed method are threefold. (1) Capability of both FPR and VPR: the method achieves FPR and VPR by applying circular vibrations to the chip in both the focal and vertical plane (see Figure 1f and 1g, respectively). (2) System simplicity: the system is simplified, consisting only of the microfluidic chip, a chip holder (to provide a connection to the piezoelectric actuators that supply the vibrations) and a microscope for observations (see Figure 1e). The microfluidic chip is fabricated via a one-step photolithography process. The experimental set-up of cell rotation is completed merely by placing the chip in the holder. (3) Open chip structure: the proposed method enables cell rotation on an open chip, rendering it suitable for applications that require external probe access. Furthermore, the sample is easily introduced by dropping the culture medium containing the sample cells into the chip holder, as shown in Figure 1e.
Materials and methods
Vibration-induced flow
When a vibrating object is placed in a microfluidic environment, it induces a local flow in its vicinity15–17. To mix or pump fluids in a microfluidic channel or trap micro-objects, some researchers have generated local flows by applying rectilinear vibrations to the chip16–19. These flows can be induced on open chips or in closed-chip environments. In our previous study, we applied 2D circular vibrations to responsive micropillars20. This configuration induces a highly localized whirling flow around the micropillar, as shown in Figure 1c. By patterning a micropillar array and applying 2D circular vibrations to the chip, we induced flow along the micropillar array and transported small cells (≈10 μm), even in an open chip environment.
In the present study, this 2D flow control method is expanded to 3D cell rotation by adding one degree of freedom to the piezoelectric actuator. We patterned three micropillars in a triangular configuration on a glass substrate, as shown in Figure 1d. For FPR, we applied a circular vibration to the xy plane and induced local whirling flow around each micropillar, as shown in Figure 1f. Eventually, the flows around the individual micropillars interfered to generate a rotational flow at the combined centre of the micropillars. For VPR, we applied a circular vibration to the xz (or yz) plane, which induced a local vertical flow around the micropillars (see Figure 1g). These vertical flows supplied a vertical torque to the target cell, enabling VPR. In this way, vibration-induced flow can realize both FPR and VPR.
Theoretical analysis
To evaluate the design and input parameters, we theoretically analysed the vibration-induced flow around a single micropillar. Previously, we evaluated the effects of the frequency of the applied vibrations20. Here, we evaluate the effects of micropillar radius and vibration amplitude. The coordinate system is illustrated in Figure 2a. Our analysis is based on the stream equation of a 2D incompressible fluid with constant viscosity, given by
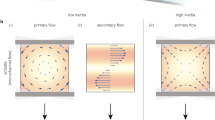
Theoretical analysis of vibration-induced flow: (a) Coordinate system. (b) Calculated flow velocity distribution of vibration-induced flow around a micropillar with f = 100 Hz, a = 50 μm and A = 1.0 μm (black arrows indicate the flow direction). (c) Radial flow velocity profiles under different conditions: (i) f = f0, a = a0, A = A0 (black solid line), (ii) f = 2f0, a = a0, A = A0 (red broken line), (iii) f = f0, a = 2a0, A = A0 (green dashed-dotted line), (iv) f = f0, a = a0, A = 2A0 (blue dashed double-dotted line). (d) Normalized maximum velocity versus normalized frequency f = f0 (red triangles; other parameters fixed to a = a0 and A = A0), radius a/a0 (green circles; other parameters set to f = f0 and A = A0) and amplitude A/A0 (blue squares; other parameters set to f = f0 and a = a0). (e) Normalized distance to velocity peak versus the normalized parameters explained in Figure 2d.
In Equation (1), , and v denote the stream function, kinematic viscosity and flow velocity vector, respectively. In our previous work, we derived the first-order approximate solution to the stream function (details are provided in Refs. 15 and 20). The steady-state solution, , dominates the vibration-induced flow and is given by
where and are the frequency and amplitude of the applied vibration, respectively. , and are n‐th order Hankel functions of the first kind, . Explicitly, these functions are defined as follows:
where ε is a length-scale parameter with dimensions of inverse length. Using the relationship between the stream function and flow velocity (Equation (7)), we can calculate the 2D flow velocity distribution around each micropillar. A typical velocity distribution is presented in Figure 2b.
The dominant parameters in Equations (2) through (4) are the frequency, f, and amplitude, A, of the applied vibration and the radius, a, of the micropillar. Thus, we varied these parameters and compared the resulting flow velocity profiles with those obtained under basic conditions (defined as f0 ≡ 100 Hz, a0 ≡ 50 μm, A0 ≡ 1.0 μm). The velocity profiles under the basic and modified conditions are presented in Figure 2c. In plots (ii), (iii) and (iv) of this figure, f = 200 Hz, a = 100 μm and A = 2.0 μm, respectively, while the other parameters were set to their base values.
We first evaluated the effect of f by comparing plots (i) and (ii) in Figure 2c. The maximum flow velocity is approximately three times larger in (ii) than in (i). We also evaluated the value of δ, the distance from the micropillar surface to the point of the flow velocity peak, which indicates the shape change of the velocity profile under various conditions. In plots (i) and (ii) of Figure 2c, the approximate δ values are 100 μm and 70 µm, respectively. These features can be understood from Equations (3), (5) and (6). The amplitude of ρ increases with f3, whereas the length-scale parameter ε is proportional to . Because ε determines the radial scale of ρ, larger values of ε indicate a flow distribution that is more localized around the micropillar. Thus, f strongly affects both the absolute values and the radial profiles of the flow velocity.
Second, we evaluated the effects of a by comparing plots (i) and (iii). Although the velocity is higher in (iii) than in (i), the difference is small relative to the change induced by doubling f, and both flow velocity profiles exhibit similar shapes. Consistent with these observations, δ is approximately 98 μm and 101 μm in plots (i) and (iii), respectively. These results support the hypothesis of this study: that we can exploit the highly localized flow surrounding the pillar surface generated by local interactions between the fluid and the micropillar surface.
Finally, we evaluated the effects of A by comparing plots (i) and (iv). The velocity in plot (iv) is approximately four times larger than that in (i), yet the profiles exhibit similar shapes and δ values. Equation (3) indicates that A affects the amplitude of ρ (specifically, ρ is proportional to A2) but not the radial profile. Therefore, the flow velocity can be changed by tuning A without changing δ or the flow velocity profile.
To confirm these behaviours of the flow velocity and its profile, we evaluated the maximum velocity, , and δ as functions of f, a and A. To this end, we introduce the normalized parameters a/a0, A/A0 and f/f0. We also normalized the velocity, , and the distance, δ/δ0, where and δ0 are the maximum velocity and the distance from the micropillar surface to the maximum velocity point under the basic conditions, respectively. and δ/δ0 are plotted as functions of each normalized parameter in Figure 2d and 2e, respectively. The variables and δ/δ0 behave as expected from our analysis of Figure 2c.
Design
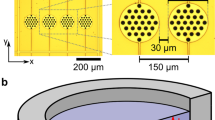
To achieve stable rotation, we arranged three micropillars in a triangular configuration. The geometric details are presented in Figure 3a. First, the micropillar height was designed to cover the target cell. In this study, the target cell was a mouse oocyte, whose size varies from approximately 70 µm to 100 μm. Thus, we set the micropillar height to 200 μm. Second, we determined the micropillar radius, a. Considering the processing limits of the photoresist and the size of the microscope sensing area, we set a to 60 μm. Finally, we determined the distance between the micropillars, d1. Because d2 is geometrically determined by , the necessary condition is given by
The upper limit of Equation (8) avoids any overlap between the target cell and the micropillars, while the lower limit prevents the overlap of the micropillar patterns. Considering the size variability of the target cell, the minimum is 226 μm. Because this value exceeds a (=60 μm), we applied the following upper limit: μm. Thus, d2 was determined as 312 μm by setting d1 to 180 μm. In this micropillar geometry, the natural frequency of the micropillar is approximately 1.2 MHz when using SU-8 photoresist (SU-8 3000, Nippon Kayaku Co. Ltd., Tokyo, Japan). Because this frequency far exceeds the frequency of the applied vibration, micropillar resonance is negligible. This configuration successfully induced rotational flow at the centre of the triangle, as shown in Figure 3b.
Experiments
System set-up
The proposed cell rotation system consists of a microfluidic chip, piezoelectric actuators (PAC166J, Nihon Ceratec Co. Ltd., Sendai, Japan) to apply the circular vibration and an inverted microscope (Eclipse Ti, Nikon Corporation, Tokyo, Japan) for observations. Microscopic images of the target oocyte were acquired with a CCD camera (Flea 3, ViewPLUS Inc., Tokyo, Japan) attached to the microscope and recorded on a control PC. The micropillars on the chip surface were fabricated from SU-8 photoresist applied to a glass substrate with a one-step photolithography process. The driving voltages of the piezoelectric actuators were output from a DA board (PCI-3340, Interface Corporation, Hiroshima, Japan) connected to the control PC. These output signals were amplified by a high-voltage amplifier (9400, Toyo Corporation, Tokyo, Japan). The piezoelectric actuators were driven by sinusoidal signals. The oocyte's nucleus was observed under a confocal laser scanning microscope (CLSM) (CSU-X, Yokogawa Electric Corporation, Tokyo, Japan).
Sample cell preparation
Sample mouse oocytes were purchased from Japan SLC Inc., Shizuoka, Japan. Immediately prior to the experiment, the oocytes were cryopreserved and thawed following the protocols of the Ref. 21. The nucleus was stained with SYTO13 Green Fluorescent Nucleic Acid Stain in preparation for observation (S7575, Life Technologies Japan Ltd., Tokyo, Japan). Additionally, the oocyte cytoplasm was stained with Cellstain CytoRed (C410, Dojindo Laboratories, Kumamoto, Japan). We used Medium 199 (11150-059, Life Technologies Japan Ltd., Tokyo, Japan) as the oocyte culture medium for all experiments. For oocytes that were fluorescently stained, we used Medium 199 without phenol red (11043-023, Life Technologies Japan Ltd., Tokyo, Japan) to reduce the background autofluorescence due to the phenol red.
Results
Figure 4a shows a series of photographs of rotating mouse oocytes acquired during FPR evaluations. As a visual guide to the rotation angle, the characteristic point of the oocyte is enclosed by a dotted circle. According to the photographs, the oocyte rotated in the focal plane of the microscope. Its rotation was governed by vibration-induced flow at the centre of the triangular configuration of the micropillars. We first evaluated the acceleration and deceleration periods of the FPR speed. The acceleration period is defined as the time required for the rotation to reach a stable speed after the vibration is first applied. Similarly, the deceleration period is defined as the time required for the rotation to cease (i.e. to reach 0.0° s−1) after the vibration is removed. The stable speed is defined as the speed at which the deviations are stabilized within ±10° s−1. Figure 4b plots the response characteristics of the clockwise (CW) and counterclockwise (CCW) FPR speeds under applied voltages of 1.0, 2.0 and 3.0 V. The frequency of the applied vibration was fixed at 200 Hz. The time at which vibration was first applied is considered 0 ms, and vibrations were applied for 5000 ms. Figure 4b shows that the acceleration and deceleration periods were less than 200 ms for all conditions and that the rotation direction can be controlled by switching the direction of the applied vibration. Second, we evaluated the stable CW and CCW FPR speeds for various amplitudes of the applied vibration. The results are plotted in Figure 4c. As the amplitude of the applied vibration increased from 1.0 to 3.0 V, the stable speed increased from 5.8° s−1 to 63.7° s−1. Furthermore, the rotational speed qualitatively depended on the amplitude, as predicted by the theoretical analysis, indicating that the flow velocity was proportional to the square of the amplitude of the applied vibration. The differences between the CW and CCW speeds likely originate from differences in the chip mounting angles or tolerances of the piezoelectric actuators. These results indicate that FPR was successful for the mouse oocytes.
Results of FPR: (a) A series of photographs of mouse oocytes during FPR. The dotted circle encloses the characteristic point for detecting the rotation angle. (b) Step response of rotational velocity. (c) Dependence of stable velocity on applied voltage. In (b) and (c), red square, green circle and blue triangles indicate the results for applied voltages of 3.0 V, 2.0 V and 1.0 V, respectively. Closed and open symbols for each condition indicate the results of clockwise (CW) and counterclockwise (CCW) rotation, respectively.
In the VPR evaluation, the relative 3D position of the nucleus in the target oocyte was evaluated using CLSM. Figure 5b and 5c was obtained scanning the oocyte in 1.0 μm steps in the z-direction before and after VPR, respectively. The frequency, amplitude and duration of the applied vibration were 200 Hz, 3.0 V and 10 s, respectively. The vibration was applied in the normal and opposite directions (see Figure 5a for a definition). The positions of the nucleus before the vibration () and after the vibration () were measured, and the VPR angles were calculated as and . Figure 5b and 5c shows the cross-sectional views of the 3D images of the target oocyte before and after VPR, respectively. The cytoplasm and nucleus are stained red and green, respectively. These figures show a shift in the relative position of the nucleus, from which the rotation angles were measured as = −16.5° s−1 and = 24.3° s−1. This step was repeated five times to obtain the average VPR speed in θ (vθ) and ϕ (vϕ) (see Figure 5d). The average rotational speeds in the normal and opposite direction were 3.5 ± 2.1° s−1 and 2.2 ± 1.1° s−1, respectively.
Results of VPR: (a) Definition of the coordinate system, VPR angle and direction of the applied vibrations (normal and opposite). (b) and (c): Cross-sectional views of fluorescent images of the target oocyte before and after VPR, respectively. (d) Calculated VPR speeds of mouse oocytes. Grey bar and white bar indicate vθ and vϕ, respectively.
Thus, we successfully achieved the VPR of oocytes and altered the direction by switching the direction of the applied vibration.
Discussion
The experimental results indicate that the proposed cell rotation method based on vibration-induced flow can achieve both the VPR and FPR of mouse oocytes. Figure 5d indicates that of FPR (vϕ) coupled with VPR (vθ ) when VPR was performed by applying a circular vibration in the vertical plane. However, little coupling between VPR and FPR was observed when FPR was performed as shown in Supplementary File 1. Therefore, we can apply the proposed method to control the orientation with the following procedure:
-
1
First, vertically rotate the target cellular structure (ex. nucleus) to the focal plane using VPR.
-
2
Second, horizontally rotate the target cellular structure in the focal plane using FPR.
Here, we demonstrate the orientation control of mouse oocytes and discuss further applications of the proposed method combined with the transportation method by using the vibration-induced flow. This technique enables the orientation control of a mouse oocyte with one-time pipetting, achieved by merely dropping the sample medium containing a target oocyte into the chip. In our previous study, we proposed that cell transportation is also possible via vibration-induced flow20. This transportation can be simply integrated into the rotation chip by constructing a micropillar array around the proposed triangular configuration, as shown in Figure 6a. The vibration-induced flow then transports the dropped target oocyte along the micropillar array. The spiral micropillar pattern guides the target cell towards the centre of the spiral from any area covered by the pattern. Once the target cell has reached the centre of the triangular configuration, it can be rotated using the proposed method. In the micropillar array design, we set the micropillar radius and pitch to 100 μm and 50 μm, respectively. In the triangular configuration, the width of the entrance to the rotation point (d3) was designed as 180 μm, which was sufficiently large for the oocyte to pass between the micropillars. Regarding the rotation, the internal radius of the micropillars, a1, was calculated using Equation (8), and a2 was determined to be 100 μm. The oocyte transportation and orientation are demonstrated in Figure 6c. The sample medium containing the target oocyte was dropped into the chip, and the oocyte was transported along the micropillar array towards the spiral centre. This mechanism automatically guided the target oocyte into the rotation point. The orientation control of mouse oocytes is demonstrated in Figure 7. To orient the nucleus of the oocyte in an arbitrary position in the focal plane, the nucleus was first positioned in the focal plane by VPR, as shown in Figure 7a. Once positioned, circular vibrations were repeatedly applied for 1 s, as shown in Figure 7b. We successfully positioned the oocyte nucleus at 39.4 ± 7.4°. Thus, we integrated the cell transportation and proposed cell rotation method to control the orientation of a mouse oocyte with one-time pipetting.
Finally, we assessed the effect of the proposed method on the viability of cells. We transported and rotated five oocytes with our combined system. The target oocytes were dropped onto the chip, and circular vibrations were applied. The vibrations rotated the target oocytes for five minutes. The target oocytes were collected with a micropipette and incubated in a culture well at 37 °C in a 5% CO2 atmosphere for three hours. The frequency and amplitude of the applied voltage were 200 Hz and 2 V, respectively. We also prepared control samples, which were thawed at the same time as the experimental samples and incubated under identical conditions. The viability of the incubated cells was then assessed using the LIVE/DEAD Viability Kit (L-3224, Life Technologies Japan Ltd., Tokyo, Japan). All experimental and control samples remained viable after the incubation. Thus, the proposed method did not affect cell viability.
The proposed method enables 3D cell rotation on an open chip structure. Due to the simplicity and open structure of the system set-up (it is easily accessed by a micropipette), this method is expected to become a user-friendly on-chip cell manipulation technique. The set-up is completed merely by attaching the chip to the chip holder. In addition to its simplicity, the method demonstrates excellent performance and will likely become a valuable tool in biotechnological research.Footnote 1
Notes
Supplementary information for this article can be found on the Microsystems & Nanoengineering website (http://www.nature.com/micronano).
References
Campbell HK . Nuclear transfer in farm animal species. Seminars in Cell & Developmental Biology 1999; 10: 245–252.
Rudolf Jaenisch . Transgenic animals. Science 1988; 240: 1468–1474.
Yanagida K, Katayose H, Yazawa H, et al. The usefulness of a piezo-micromanipulator in intracytoplasmic sperm injection in humans. Human Reproduction 1999; 14: 448–453.
Ramadan A, Inoue K, Arai T, et al. New architecture of a hybrid two-fingered micro–nano manipulator hand: optimization and design. Advanced Robotics 2008; 22: 235–260.
Turcu I, Lucaciu CM . Electrorotation: a spherical shell model. Journal of Physics A: Mathematical and General 1989; 22: 995–1003.
Hölzel R, Lamprecht I . Dielectric properties of yeast cells as determined by electrorotation. Biochimica et Biophysica Acta (BBA)-Biomembranes 1992; 1104: 195–200.
Huang Y, Wang XB, Holzel R, et al. Electrorotational studies of the cytoplasmic dielectric properties of friend murine erythroleukaemia cells. Physics in Medicine and Biology 1995; 40: 1789–1806.
Jones TB . Basic theory of dielectrophoresis and electrorotation. IEEE Engineering in Medicine and Biology Magazine 2003; 22: 33–42.
Pethig R, Jakubek LM, Sanger RH, et al. Electrokinetic measurements of membrane capacitance and conductance for pancreatic β-cells. IEE Proceedings-Nanobiotechnology 2005; 152: 189–193.
Han SI, Don Joo Y, Han KH . An electrorotation technique for measuring the dielectric properties of cells with simultaneous use of negative quadrupolar dielectrophoresis and electrorotation. Analyst 2013; 138: 1529–1537.
Benhal P, Chase JG, Gaynor P, et al. AC electric field induced dipole-based on-chip 3D cell rotation. Lab on a Chip 2014; 14: 2717–2727.
Elbez R, McNaughton BH, Patel L, et al. Nanoparticle induced cell magneto-rotation: monitoring morphology, stress and drug sensitivity of a suspended single cancer cell. PloS One 2011; 6: e28475.
Kreysing MK, Fritsch A, Dietrich C, et al. The optical cell rotator. Optics Express 2008; 16: 16984–16992.
Carmon G, Feingold M . Rotation of single bacterial cells relative to the optical axis using optical tweezers. Optics Letters 2011; 36: 40–42.
Holtsmark J, Johnsen I, Sikkeland T, et al. Boundary layer flow near a cylindrical obstacle in an oscillating, incompressible fluid. The Journal of the Acoustical Society of America 1954; 26: 26–39.
Hagiwara M, Kawahara T, Arai F . Local streamline generation by mechanical oscillation in a microfluidic chip for noncontact cell manipulations. Applied Physics Letters 2012; 101: 074102.
Lutz BR, Chen J, Schwartz DT . Hydrodynamic Tweezers: 1. Noncontact trapping of single cells using steady streaming microeddies. Analytical Chemistry 2006; 78: 5429–5435.
Huang P-H, Xie Y, Ahmed D, et al. An acoustofluidic micromixer based on oscillating sidewall sharp-edges. Lab on a Chip 2013; 13: 3847–3852.
Huang P-H, Nama N, Mao Z, et al. A reliable and programmable acoustofluidic pump powered by oscillating sharp-edge structures. Lab on a Chip 2014; 14: 4319–4323.
Hayakawa T, Sakuma S, Fukuhara T, et al. A single cell extraction chip using vibration-induced whirling flow and a thermo-responsive gel pattern. Micromachines 2014; 5: 681–696.
Nakao K, Nakagata N, Katsuki M . Simple and efficient procedure for cryopreservation of mouse embryos by simple vitrification. Experimental Animals 1997; 46: 231–234.
Acknowledgements
This study was financially supported by Grant-in-Aid for JSPS Fellows Number 13J03580 and Grant-in-Aid for Scientific Research on Innovative Areas (No. 23106002) and (No. 26630094).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Supplementary information
Figure S1
Results of FPR: (a) and (b) Cross-sectional views of fluorescent images of the target oocyte before and after FPR, respectively. (c) Calculated FPR speed of mouse oocyte. Red horizontal-striped bar and blue vertical-striped bar indicate vθ and vϕ, respectively. (PDF 289 kb)
Supplementary Video
Rotation_Movie (MP4 4141 kb)
Rights and permissions
This license allows readers to copy, distribute and transmit the Contribution as long as it attributed back to the author. Readers are permitted to alter, transform or build upon the Contribution as long as the resulting work is then distributed under this is a similar license. Readers are not permitted to use the Contribution for commercial purposes. Please read the full license for further details at - http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Hayakawa, T., Sakuma, S. & Arai, F. On-chip 3D rotation of oocyte based on a vibration-induced local whirling flow. Microsyst Nanoeng 1, 15001 (2015). https://doi.org/10.1038/micronano.2015.1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/micronano.2015.1
This article is cited by
-
Revolutionizing the female reproductive system research using microfluidic chip platform
Journal of Nanobiotechnology (2023)
-
Nanodroplet-mediated catheter-directed sonothrombolysis of retracted blood clots
Microsystems & Nanoengineering (2021)
-
A microfluidic approach for synchronous and nondestructive study of the permeability of multiple oocytes
Microsystems & Nanoengineering (2020)
-
An Open-Chip for Three-Dimensional Rotation and Translation of Particle Based on Dielectrophoresis
Multiscale Science and Engineering (2019)
-
Closed loop control of microscopic particles incorporating steady streaming and visual feedback
Biomedical Microdevices (2018)