Abstract
The C-type lectin-like receptor CD161 is expressed by lymphocytes found in human gut and liver, as well as blood, especially natural killer (NK) cells, T helper 17 (Th17) cells, and a population of unconventional T cells known as mucosal-associated invariant T (MAIT) cells. The association of high CD161 expression with innate T-cell populations including MAIT cells is established. Here we show that CD161 is also expressed, at intermediate levels, on a prominent subset of polyclonal CD8+ T cells, including antiviral populations that display a memory phenotype. These memory CD161intCD8+ T cells are enriched within the colon and express both CD103 and CD69, markers associated with tissue residence. Furthermore, this population was characterized by enhanced polyfunctionality, increased levels of cytotoxic mediators, and high expression of the transcription factors T-bet and eomesodermin (EOMES). Such populations were induced by novel vaccine strategies based on adenoviral vectors, currently in trial against hepatitis C virus. Thus, intermediate CD161 expression marks potent polyclonal, polyfunctional tissue-homing CD8+ T-cell populations in humans. As induction of such responses represents a major aim of T-cell prophylactic and therapeutic vaccines in viral disease and cancer, analysis of these populations could be of value in the future.
Similar content being viewed by others
INTRODUCTION
The phenotypic complexity contained within the CD8+ T-cell compartment, conventionally considered a relatively uniform “cytotoxic” population in contrast to multiple CD4+ T helper (Th) subsets, is being increasingly appreciated. In addition to the presence of unconventional T cells within the CD8+ pool,1 the identification of tissue-resident memory (TRM) cells, present within tissues such as skin and gut where they can provide rapid protection,2, 3, 4 is important in aiding the study of both pathophysiology and rational vaccine design. Understanding these functional distinctions, and identifying the markers that define them, is important in determining favorable immune outcomes in both disease and therapeutic/vaccination strategies.
CD161, or KLRB1, is a C-type lectin-like receptor expressed by the majority of natural killer (NK) cells5 and in a subset of T cells,6 and is particularly enriched in populations found in the gut and liver.7, 8, 9 Analysis of CD8+ T cells expressing high levels of this marker revealed a CD8+ T-cell subset analogous to CD4+ Th17 cells,9 subsequently identified to comprise a population of semi-invariant T cells known as mucosal-associated invariant T (MAIT) cells.10 MAIT cells express the T cell receptor (TCR) Vα7.2-Jα33/12/20, are restricted by the nonclassical class Ib molecule major histocompatibility complex (MHC)-related 1 (MR1), and are enriched at mucosal sites.11, 12, 13
Binding of CD161 to its ligand lectin-like transcript 1 (LLT1) has been described to have both inhibitory14, 15, 16 and costimulatory14, 16, 17 effects. Yet, independently of function, CD161 has been applied as a marker of type 17 phenotype across all T cells.18 More recently, we have also described this molecule to mark T cells with an enhanced innate ability to respond by a TCR-independent mechanism to interleukin (IL)-12 and IL-18.19 Although CD8+ T cells expressing high levels of CD161 (CD161hi/CD161++) predominantly comprise MAIT cells, a population of CD8+ T cells expressing lower, or intermediate, levels of CD161 (CD161int/CD161+) is also present in the circulation.6, 9, 10 The phenotypic characteristics and immunological function of this CD161intCD8+ T-cell population is yet to be defined. Although CD161 is associated with the ability to express IL-17,18 secretion of this cytokine among CD8+ T cells is restricted to the CD161hi subset, and was not evident from the CD161intCD8+ population.9 Here, we investigated the phenotype, function, and mucosal association of this novel CD8+ T-cell population.
RESULTS
CD161 is expressed by a subset of memory CD8+ T cells
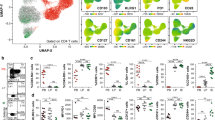
The NK cell marker CD161 is expressed by a subset of CD8+CD3+ lymphocytes,5 and defines three populations within the peripheral blood: CD161 high (CD161hi), CD161 intermediate (CD161int), and CD161 negative (CD161neg) (Figure 1a). Approximately equal percentages of CD161hi and CD161int were observed in healthy adults (mean 10.60% and 11.68%, respectively). Although these populations did express other NK cell receptors, expression of further NK-associated markers tested here were not specific to any of these populations (Supplementary Figure S1 online). The majority of CD161hiCD8+ T cells express the Vα7.2 TCR characteristic of MAIT cells, as described elsewhere,10, 13, 20 that display semi-invariant TCR expression of Vα7.2-Jα33/12/2013 with oligoclonal Vβ expression.1 However, unlike CD161hi MAIT cells the CD161intCD8+ T-cell subset was predominately Vα7.2− (Figure 1a). Furthermore, the CD161intCD8+ T-cell subset displayed polyclonal Vβ expression in a similar manner to CD161neg (Figure 1b).
CD161 expression defines a subset of memory CD8+ T cells in adults and infant peripheral blood. (a) Representative flow cytometry plots showing CD161 and CD8 (upper panel) or T cell receptor (TCR) Vα7.2 (lower panel) expression in CD8+CD3+ lymphocytes, and cumulative data for the percentage of each subset in healthy adult peripheral blood (n=28). Horizontal bars represent mean values. (b) Percentage of the CD161int and memory CD161neg CD8+ T-cell subsets expressing each TCR Vβ chain as determined by flow cytometry (n=10). Flow cytometric analysis of CCR7 and CD45RA expression was used to divide CD8+CD3+ lymphocytes into naive (CCR7+CD45RA+) and memory subsets in peripheral blood from adults (c,d) and 24 month olds (e,f). Memory cells were further subgated into central memory (TCM; CCR7+CD45RA−), effector memory (TEM; CCR7−CD45RA−), and terminal effector memory (TEMRA; CCR7−CD45RA+) subtypes (n=16). ****P<0.0001, NS, not significant by one-way analysis of variance (ANOVA) with Tukey’s multiple comparisons test. Data are represented as mean±s.e.m.
The differentiation phenotype of CD161intCD8+ T cells was investigated using the markers CCR7 and CD45RA to divide T cells into naive (TN: CCR7+CD45RA+), effector memory (TEM: CCR7−CD45RA−), central memory (TCM: CCR7+CD45RA−), and terminally differentiated memory (TEMRA: CCR7−CD45RA+) subsets.21 Whereas approximately a quarter of CD161neg cells displayed a naive phenotype, only a very small percentage (mean 3.59%) of CD161int were CCR7+CD45RA+, with the vast majority displaying a memory phenotype as for CD161hi cells (Figure 1c). As expected, CD161hiCD8+ T cells displayed an almost uniform TEM phenotype.22 Few CD8+ T cells with TCM phenotype were observed, as described elsewhere.23 On the other hand, TEM and TEMRA subsets were found in both the CD161int and CD161neg populations (Figure 1d). There was no significant difference in the percentage of either TEM or TEMRA subsets between the CD161int and CD161neg populations, suggesting that expression of CD161 (or loss of expression) is not associated with differentiation state.
CD161 expression was, therefore, largely restricted to memory cells within the adult circulation. To investigate the differentiation phenotype of the CD161intCD8+ population in early life, the same analysis was performed on peripheral blood mononuclear cells (PBMCs) from 24-month-old donors. Interestingly, although a larger percentage of the CD161int population displayed a naive (CCR7+CD45RA+) phenotype compared with adult blood, the majority (>89%) were already memory in phenotype (Figure 1e), as seen in the CD161hi population. In contrast, the majority of CD161negCD8+ T cells were naive. Again, CD161intCD8+ memory T cells were composed of both TEM and TEMRA subtypes (Figure 1f), with no difference in distribution between the two subsets, but a significantly (P<0.0001) greater proportion of the CD161neg population displayed a TCM phenotype compared with CD161int at this age.
CD161int CD8+ T cells are a resident memory population enriched within colon
CD161hi MAIT cells have previously been described to be enriched within the gut lamina propria12 and the liver9, 10 in humans. We similarly examined for the presence of the CD161intCD8+ T-cell population within healthy pretransplant liver and normal adjacent colonic tissue, derived from surgery for colorectal cancer. Due to the lack of an obvious CD161hi population by flow cytometry, tissue CD8+ T cells were divided based upon expression of CD161 and Vα7.2, and compared with peripheral blood (Figure 2a). As expected, CD161+Vα7.2+ cells, corresponding to MAIT cells, were a major population within the liver.24 However, CD161+Vα7.2− CD8+ T cells were also present in similar proportions to both MAIT (CD161+ Vα7.2+) and CD161− CD8+ T cells within the liver in the limited number of samples investigated here. In contrast, the majority of CD161-expressing CD8+ T cells in the colon were CD161+Vα7.2− T cells comprising approximately a third (mean 32.24%) of CD8+ T cells, with CD161+Vα7.2+ MAIT cells present in low numbers. As this CD161+ Vα7.2− population represents CD161int CD8+ T cells (Figure 1a), this demonstrated the CD161intCD8+ T-cell population to be enriched within tissues compared with blood, including the colon where they comprise the majority of CD161+ cells.
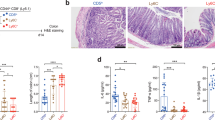
CD161intCD8+ T cells are a resident population enriched with the human colon. (a) Representative flow cytometry plot and cumulative data of CD8+CD3+ lymphocytes from blood (n=10), liver (n=4), and colon (n=11) divided based upon expression of T cell receptor (TCR) Vα7.2 and CD161. (b) Proportion of memory (CD45RO+) CD8+CD3+ lymphocytes from control (n=10), inflammatory bowel disease (IBD) noninflamed (n=15), or IBD (n=7) colon biopsies expressing CD161. **P<0.01 by one-way analysis of variance (ANOVA) with Tukey’s multiple comparisons test. Box and whiskers show min to max values. (c) Percentage of either CD161+ or CD161− CD8+ T cells expressing CD103 in control (n=6), IBD noninflamed (n=16), or IBD (n=13) tissue samples, with representative flow cytometry plots. ****P<0.001 or NS, not significant by paired t-test. (d) Percentage of memory CD8+ T-cell populations as defined by expression of both CD161 and CD103 in control (n=6), IBD noninflamed (n=12), or IBD (n=8) colon tissue samples. (e) Proportion of each CD8+ T-cell subset expressing CD69 in control colon samples (n=6). Box and whiskers show min to max values.
Analysis of the inflammatory infiltrate in patients with inflammatory bowel disease (IBD), however, revealed a significant (P<0.001) reduction in the percentage of CD161+ memory CD8+ T cells in biopsies of inflamed tissue compared with noninflamed and control tissue (Figure 2b). This is evident in comparison with both matched and nonmatched inflamed tissue, and in both Crohn’s disease and ulcerative colitis (Supplementary Figure S2). This CD161+CD8+ T-cell population was enriched for expression of CD103 (Figure 2c), an integrin preferentially expressed by human intestinal intraepithelial lymphocytes24 and a marker of residence in TRM cells of mucosal tissues including the intestine,4, 25 with expression thought to result from signals received within the gut.4, 26 Accordingly, IBD inflamed tissue was associated with an increase in the percentage of CD103− cells compared with noninflamed tissue, the majority of which were also CD161− (Figure 2d). A corresponding specific dilution of the CD161+, and in particular the CD161+CD103+, population was observed.
Together with CD103, CD69 has also been described as a marker of tissue retention and of TRM populations.23, 27 We therefore examined CD8+ T-cell subsets for CD69 expression within the gut and found the majority (mean 93%) of CD161int (CD161+ Vα7.2−) CD8+ T cells within the colon to express CD69 (Figure 2e).
CD161int CD8+ T cells display a preprogrammed phenotype
To further explore the phenotype of CD161intCD8+ T cells, mRNA microarray analysis was performed on sorted CD161int compared with CD161neg CD8+ T cells from healthy adult blood, with the exclusion of naive cells by the gating strategy shown (Figure 3a). Comparison of CD161int with CD161neg CD8+ T cells revealed a number of differentially expressed transcripts, including both upregulated and downregulated genes (Figure 3b), and with a 24.2-fold change in the expression of CD161 (KRLB1) (Supplementary Table S1). A total of 952 genes were identified to be significantly differentially expressed (P<0.01, >2-fold change), illustrated by the heatmap (Figure 3c and Supplementary Table S1). These transcripts included genes for both subunits of the IL18R (IL18R1 and IL18RAP), CXCR6, MDR1 (ABCB1), and PLZF (ZBTB16). The expression level of these markers was therefore examined by flow cytometry. This showed a higher percentage of the CD161int, when compared with the memory CD161neg, CD8+ T-cell population to be positive for each marker. However, as there appeared to be a gradient of expression levels, the average level of expression (geometric mean fluorescence intensity (geoMFI)), with background fluorescence minus one sample levels subtracted, was analyzed. Although differences observed were modest, this demonstrated significant increased expression of IL-18Rα (P<0.05), CXCR6 (P<0.001), MDR1 (P<0.01), and PLZF (P<0.01) within the CD161int CD8+ T-cell population that reflected microarray results for gene expression (Figure 3d).
CD161int CD8+ T cells display elevated expression of IL18Rα, CXCR6, MDR1, and PLZF in peripheral blood. (a) Gating strategy for sorting of CD161int and CD161neg subsets, and exclusion of naive cells, out of CD8+CD3+ lymphocytes from peripheral blood mononuclear cells (PBMCs) for microarray analysis. (b) Scatter plot of signal intensities of all mRNA probes. The signal intensities of each probe represented by a cross are shown in double logarithmic scale. Red diagonal lines define the areas of twofold differential signal intensities. Blue cross indicates unchanged genes, red cross indicates significantly upregulated genes (P<0.01), and green cross indicates significantly downregulated genes (P<0.01). (c) Heatmap illustrating the 952 significantly (P<0.01) differentially expressed transcripts between CD161int and CD161neg CD8+ T cells in 3 donors. Subsets clustered by one minus Pearson’s correlation. (d) Representative flow cytometry plots for CD161 expression vs. IL18Rα, MDR1, PLZF, and CXCR6. Percentage positive and geometric mean fluorescent intensity (geoMFI) for IL18Rα, MDR1, PLZF, and CXCR6 for CD161int and CD161neg subsets, as determined by flow cytometry, with background subtraction of geoMFI in fluorescence minus one samples, and naive (CCR7+CD45RA+) cells excluded (n=7). *P<0.05, **P<0.01, ***P<0.001, NS, not significant by paired t-test. Normalized signal intensities for expression of transcripts for IL18R1, CXCR6, ABCB1 (MDR1), and ZBTB16 (PLZF) and statistical significance from mRNA microarray expression analyses after normalization and correction for multiple testing, **P<0.01, ****P<0.0001. Floating bars show minimum and maximum values, with a line at the mean.
A CD161+Vα7.2− population was also evident among CD8+ T cells in the thymus and umbilical cord blood (UCB; Figure 4a). Although CD161 expression is associated with a memory phenotype, we confirmed that CD161intCD8+ T cells in UCB displayed a naive (CCR7+CD45RA+) phenotype (Figure 4b). Microarray analysis of naive UCB CD161int, compared with CD161neg, CD8+ T cells from four donors revealed a significant correlation in transcriptional profile with adult memory CD161intCD8+ T cells by Gene Set Enrichment Analysis (GSEA) that demonstrated significant (P<0.001) enrichment of those genes upregulated within adult CD161intCD8+ T cells (Figure 3) within the CD161int subset of UCB CD8+ T cells (Figure 4c). The naive CD161int population within UCB again displayed modestly higher expression of IL18Rα (P<0.05), MDR1 (P<0.05), and PLZF (P<0.05) than CD161neg CD8+ T cells, as measured by geoMFI, although there was no significant difference in expression of CXCR6 (Figure 4d). This indicates that, although naive, CD161intCD8+ T cells in UCB possess a preprogrammed phenotype reflective of that of CD161intCD8+ T cells in the adult circulation.
CD161intCD8+ T cells are present and preprogrammed early during development. (a) Representative flow cytometry plots showing CD161 expression by CD8+CD3+ lymphocytes in thymocytes and umbilical cord blood (UCB). (b) Representative flow cytometry plot showing naive (CCR7+CD45RA+) phenotype of UCB CD161intCD8+ T cells (blue) overlaid on bulk CD8+ T lymphocytes (red). (c) Gene set enrichment summary plot and data table for UCB CD161int vs. CD161neg CD8+ T-cell ranked genes, and enrichment of the adult CD161intCD8+ upregulated gene set. (d) Geometric mean fluorescent intensity (GeoMFI) for IL18Rα, MDR1, PLZF, and CXCR6 for UCB CD161int and CD161neg subsets, as determined by flow cytometry (n=3). *P<0.05, NS, not significant by paired t-test.
CD161int CD8+ T cells express functional MDR1
CD161intCD8+ T cells express higher levels of the multidrug efflux pump MDR1 than CD161neg cells in both UCB (Figure 4d) and adult blood (Figure 3d). Furthermore, a greater percentage of the CD161int population in adult blood expresses this pump compared with the CD161neg (mean 38.9% vs. 27.65%, respectively) within the memory CD8+ T-cell pool (Figure 5a).
CD161intCD8+ T cells express functional MDR1 in peripheral blood. (a) Representative flow cytometry plot and cumulative data for MDR1 expression by peripheral blood CD8+CD3+ lymphocytes, excluding CCR7+CD45RA+ naive cells (n=10). *P<0.05, ***P<0.001, by one-way analysis of variance (ANOVA) with Tukey’s multiple comparisons test. Data are represented as mean±s.e.m. (b,c) Representative flow cytometry plots of CD8+CD3+ lymphocytes, excluding CCR7+CD45RA+ naive cells, loaded with Rhodamine 123 (Rh123) on ice and either kept on ice (loading control) or incubated at 37 °C in media alone (efflux) (b) or in the presence of the ABCB1 inhibitors cyclosporine A (CsA) and verapamil (c). Cumulative data show Rh123 efflux from each subset calculated as per Methods, relative to the fluorescence of dye loaded (loading control). ****P<0.0001 by one-way ANOVA with Tukey’s multiple comparisons test (n=12) (b) or the percentage of each subset having effluxed Rh123. *P<0.05, ****P<0.0001, NS, not significant by two-way ANOVA with Dunnett’s multiple comparisons test, compared with the efflux sample (n=8) (c). Data are represented as mean±s.e.m.
CD161hiCD8+/MAIT cells have previously been described to express high levels of functional MDR1, enabling them to efflux xenobiotics.10, 28 Functional activity of MDR1 can be assayed by measuring efflux of the fluorescent substrate Rhodamine 123 (Rh123).29 Cells loaded with this cell-permeant dye are detected by flow cytometry (loading control; Figure 5b), with efflux determined by a loss in fluorescence (Efflux; Figure 5b). High levels of MDR1 activity were confirmed within the CD161hi population; however, the CD161intCD8+ T-cell population also displayed significant (P<0.0001) MDR1 activity. Similar frequencies of CD161hiCD8+ T cells were observed before and after efflux (data not shown), suggesting that this was not due to a downregulation of CD161 within the MAIT cell subset. Efflux was inhibited by the addition of MDR1 inhibitors cyclosporin A and verapamil (Figure 5c). Therefore, CD161intCD8+ T cells possess the ability to efflux through the expression of MDR1, which may enable survival within tissues with a high exposure to xenobiotics, such as the gut and liver.
CD161 expression marks polyfunctional antiviral cells equipped with cytotoxic mediators
To examine the specificity of the CD161intCD8+ T-cell population, MHC class I dextramers presenting epitopes from cytomegalovirus, Epstein–Barr virus (EBV), and influenza were utilized. These identified a subset of anti-viral CD8+ T cells specific for each virus to express CD161 in adult peripheral blood (Figure 6a). Dextramer-positive cells were rarely CD161hi, as previously described,10 corresponding to the restriction of CD161hi MAIT cells to MR1 presenting a bacterial ligand. Approximately 15% of antiviral populations were contained within the CD161intCD8+ T-cell population for all three viruses. Furthermore, although the majority of anti-viral CD8+ T cells were CD161neg, there was no significant difference in terms of frequency within the parent subsets, although there was a trend toward a higher percentage of influenza-specific cells in the CD161int population (Supplementary Figure S3).
CD161intCD8+ T cells are polyfunctional antiviral cells. (a) Representative flow cytometry plots showing CD161 expression by CD8+ T cells specific for epitopes from cytomegalovirus (CMV; NLVPMVATV), Epstein–Barr virus (EBV; GLCTVAML), and influenza virus (FLU; GILGFVFTL) as distinguished by dextramer staining of adult peripheral blood, and cumulative data for percentage of antiviral subset expressing CD161. Horizontal bars represent mean values (n=5). (b) Percentage of CD161neg or CD161int subsets expressing interferon-γ (IFNγ), interleukin-2 (IL-2), or a combination of both upon stimulation with phorbol 12-myristate 13-acetate (PMA)+ionomycin (n=18). Representative flow cytometry plots for cytokine expression in CD8+CD3+ peripheral blood mononuclear cells (PBMCs), excluding CCR7+CD45RA+ naive cells, are shown. (c) Scatter plot of signal intensities of all mRNA probes in stimulated (y axis) vs. unstimulated (x axis) CD161intCD8+ T cells from peripheral blood. The signal intensities of each probe represented by a cross are shown in double logarithmic scale. Red diagonal lines define the areas of twofold differential signal intensities. Blue cross indicates unchanged genes, red cross indicates significantly upregulated genes (P<0.01), and green cross indicates significantly downregulated genes (P<0.01). (d) Representative flow cytometry plots and cumulative data for percentage of CD8+CD3+ PBMCs, excluding CCR7+CD45RA+ naive cells, expressing cytotoxic effectors granzyme B and perforin. (e) Percentage of CD161int and CD161neg CD8+ T cells expressing cytokine (IL-2, IFNγ, tumor necrosis factor-α (TNFα), and macrophage inflammatory protein-1β (MIP1β) and cytotoxic mediators (granzyme B (GzmB), perforin, and CD107) singly or in combination as detected by cytometry by time-of-flight (CyTOF) (n=4). (f) Percentage of cells expressing EOMES (upper panel) or T-bet (lower panel) in memory CD8+CD3+ lymphocytes with representative flow cytometry plots. *P<0.05, **P<0.01, ***P<0.001, ****P<0.0001. NS, not significant by paired t-test. Data are represented as mean±s.e.m.
CD8+ T cells exert antiviral effects principally through cytokine release and cytotoxicity. The ability of the CD161intCD8+ T-cell population to perform these effector functions in comparison with the CD161negCD8+ T-cell population was investigated. As a variety of phenotypic and functional characteristics are associated with differentiation from a naive to memory phenotype,30 and the majority of CD161intCD8+ T cells display a memory phenotype whereas the CD161neg CD8+ T-cell population contained a mixture of naive and memory cells (Figure 1c), naive cells were excluded from all analyses.
Stimulation of CD8+ T cells with phorbol 12-myristate 13-acetate and ionomycin revealed a significantly greater proportion of the CD161intCD8+ T-cell population to express interferon-γ (IFNγ; P<0.0001) and IL-2 (P<0.05) than the memory CD161negCD8+ population, despite the fact that memory phenotypes are equally distributed between both populations (Figure 1d). Indeed, significantly (P<0.01) more CD161intCD8+ T cells were dual producers of both cytokines (Figure 6b). In a hypothesis-free approach, phorbol 12-myristate 13-acetate plus ionomcyin-stimulated CD161intCD8+ T-cell population were analyzed by mRNA microarray. This revealed the ability to express multiple cytokines and chemokines, including those associated with type 1, 2, and 17 responses (Figure 6c and Supplementary Table S2). However, comparison with stimulated CD161negCD8+ T cells did not identify expression of any specific cytokines uniquely produced by the CD161int population (Supplementary Figure S3 and Supplementary Table S3).
CD8+ T cells are considered cytotoxic lymphocytes, with cytotoxicity against target cells mediated through perforin and granzyme B. A significantly larger proportion (P<0.0001) of the CD161intCD8+ T-cell population expressed granzyme B (mean 60.74%) compared with the memory CD161negCD8+ T-cell population (mean 44.76%) (Figure 6d). Moreover, CD161intCD8+ T cells expressed significantly (P<0.001) higher levels of perforin (mean 48.83%) than CD161negCD8+ T cells (mean 27.26%), rendering these cells potently equipped for cytotoxic killing. This is in contrast to CD161hi MAIT cells that lack granzyme B and express low levels of perforin in the resting state (Figure 6d).31
The potential of this population to perform multiple functions was probed by cytometry by time-of-flight, which enables the simultaneous measurement of multiple parameters19 (Figure 6e). This demonstrated the ability of the CD161intCD8+ T-cell population to express multiple cytotoxic effectors simultaneously, together with the ability to degranulate as assayed by CD107 capture, and to a greater extent than observed within the CD161neg compartment which showed a greater propensity toward granzyme B expression only. These cytotoxic effectors were also expressed in addition to cytokine. Furthermore, the ability of CD161intCD8+ T cells to produce numerous lymphokines, including tumor necrosis factor-α and macrophage inflammatory protein-1β, with a trend toward greater a proportion of cytokine-producing cells than in the CD161neg compartment, was also observed. Together, this demonstrated the CD161intCD8+ T-cell population to be polyfunctional in terms of cytokine and cytotoxic potential.
The transcription factors T-bet and eomesodermin (EOMES) are critical for CD8+ T-cell effector function, regulating expression of IFNγ, granzyme B, and perforin.32, 33, 34 In line with their enhanced expression of cytokine and cytotoxic effectors, CD161intCD8+ T cells expressed significantly more of both EOMES and T-bet than the memory CD161neg population (Figure 6f).
CD161 expressing antiviral cells with cytotoxic phenotype can be induced by vaccination
The generation of potent T cells is a key aim of T-cell vaccine strategies. As intermediate CD161 expression marks highly functional antiviral CD8+ T cells, we wanted to investigate the potential for T-cell vaccines to induce CD161int antiviral T cells that reflect this phenotype. A novel recombinant adenoviral vector vaccination strategy was recently described to induce broad and sustained CD8+ T cells against hepatitis C virus.35 Examination of these antiviral CD8+ T cells showed that CD161 was expressed by ∼18% of these long-lasting cells (Figure 7a), compared with 15% of bulk CD8+ T cells. The perforin and granzyme B expression of these antiviral cells, as identified by MHC class I multimers, was analyzed by geoMFI due to small numbers of cells. This revealed that these CD161int cells expressed significantly (P<0.01) higher levels of granzyme B (Figure 7b) and perforin (Figure 7c) compared with antiviral CD161neg CD8+ T cells, demonstrating the potential to induce viral-specific CD161intCD8+ T cells that reflect the highly functional phenotype of the bulk CD8+ T-cell subset through vaccination.
CD161int-associated phenotype in antiviral CD8+ T cells induced by vaccination. CD8+ T cells targeting epitopes from hepatitis C virus, induced by a novel T-cell vaccine strategy based on adenoviral vectors35 were identified from peripheral blood mononuclear cells (PBMCs) using major histocompatibility complex (MHC) class I multimers. (a) Percentage of the antiviral and bulk CD8+ T-cell populations expressing CD161. Geometric mean fluorescent intensity (GeoMFI) for (b) granzyme B or (c) perforin of antiviral populations either CD161int or CD161neg, as determined by flow cytometry (n=10). **P<0.01 and NS, not significant by paired t-test. Data are represented as mean±s.e.m.
DISCUSSION
Delineating the phenotypic diversity contained within the T-cell compartment is of much importance in fully understanding immune mechanisms of both protection and pathology. Here, we describe CD161 to mark potent antiviral memory CD8+ T cells with enhanced effector function, including cytokine production and, in contrast to CD161hi/MAIT cells,31 high expression of cytotoxic mediators in the resting state, related to their elevated expression of T-bet and EOMES. These functional attributes have various implications for antiviral immunity,30, 36, 37 and thus CD161 may provide a useful marker in tracking CD8+ T-cell responses in viral disease and following vaccination. Indeed, analysis of antiviral cells induced in a T-cell vaccine strategy targeting hepatitis C virus35 revealed intermediate CD161 expression in vaccine-induced cells associated with elevated levels of granzyme B and perforin. These T-cell vaccine strategies are now also undergoing clinical trial for induction of protective immunity against Ebola virus.38
It is interesting to note that Vα7.2+ MAIT cells, although initially termed “mucosally-associated” according to a described enrichment within the gut lamina propria,12 were found here to be relatively rare within the colon, being instead much more predominant within the liver, in agreement with other studies.10, 23 However, MAIT cells have been detected at high frequencies by MR1-tetramer staining within jejunal mucosa,13 and this may reflect enrichment of MAIT cells at specific anatomical locations within the gut,39 with low frequencies in colon. On the other hand, CD161intCD8+ T cells were a major population of T cells within the colon and were also enriched, to a similar extent to MAIT cells, within the liver.
As a prominent population within the gut, expression of CD103 and CD69 by CD161intCD8+ T cells may function in their retention as TRM cells.27 Although CD103 and CD69 are considered markers of TRM,27 the expression of CD161 by this memory population has not yet been investigated. Here, an increase in the percentage of CD161neg(CD103−) cells was observed in inflamed colon tissue, potentially indicative of an influx of these cells, and resulted in the dilution of the CD161+ (CD103+) population. As CD161hi MAIT cells are rare within the colon, this CD161+ population largely corresponds to the CD161int CD8+ subset. Together, this could suggest a more stable residence of CD161intCD8+ T cells within the colon. This tissue location, together with their antiviral specificity, means that these cells are ideally placed to provide early protection at points of pathogen entry. Indeed, TRM cells are virus-specific cells that persist in tissues after viral infection, where they mediate enhanced protection against reinfection,2, 3 including through the recruitment of other effector cells.40 It has recently been shown that once activated, TRM can induce an antiviral state to provide immunity against unrelated viral infections, mediated through IFNγ.40, 41 As CD161intCD8+ T cells show an enhanced ability to produce IFNγ in response to cytokine stimulation, in the absence of TCR stimulation,19 this may provide a means by which CD161intCD8+ T cells could mediate protection against diverse viral infections. However, the presence of CD161intCD8+ T cells within the circulation suggests that this population may be more analogous to the CD4+ Th17 population described by Kleinschek et al.,42 found in blood and resident within gut, and also marked by CD161. Analysis of the phenotype of this gut-resident CD161intCD8+ T-cell population would provide clues as to their potential roles in homeostasis and pathology. For example, expression of functional MDR1 may facilitate survival of this population within such tissues with a high level of exposure to xenobiotics. Furthermore, as expression of this efflux pump has been associated with resistance to various drugs, including steroids,43 this has interesting implications for the treatment of conditions such as IBD.
Gene expression analysis of resting CD161intCD8+ T cells in peripheral blood revealed expression, at a lower level, of a number of MAIT cell-associated markers, in keeping with CD161 marking cells of a common transcriptional phenotype.19 Although the upregulation of these markers was modest, we have previously shown that they can still have a functional impact.19 Interestingly, these markers were also already upregulated in CD161intCD8+ T cells within UCB. Preprogramming of CD161-positive cells has previously been described for both CD161hiCD8+1 and CD161intCD4+44 T cells in UCB, representing precursors of their adult MAIT and Th17 counterparts, respectively. Similarly, there was a significant correlation by GSEA between UCB and adult CD161int genetic profiles, indicating that CD161intCD8+ T cells in UCB represent a naive yet precommitted population. This, together with the fact that CD161 expression is not readily induced on CD161negCD8+ T cells, either upon priming or restimulation45 (J.R. Fergusson, unpublished data), suggests that these cells represent a distinct population. Here we have shown this population to comprise highly potent memory T cells that reside in tissues including the colon, and which therefore warrant further investigation in the context of control and prevention of viral disease.
METHODS
Cells. All samples were in accordance with the ethical standards of the corresponding committee.
PBMCs were obtained from adults (whole blood leukocyte cones, NHS Blood and Transplant, Oxford, UK), 24 month olds (prospective birth cohort46), and UCB samples (Stem Cell Services, NHS Blood and Transplant).
Healthy liver-derived lymphocytes were obtained as previously described.23
Colonic tissue was either collected from surgical resections or from biopsies taken during routine colonoscopies. Control samples were taken from uninvolved tissue of patients undergoing surgery for colorectal cancer. Resection material was taken from IBD (both Crohn’s disease and ulcerative colitis) patients, undergoing surgery for severe disease, and biopsies from IBD patients were taken either from inflamed involved or normal noninflamed sites based on the judgment of the physicians performing the procedure. Noninflamed samples were also collected from IBD patients with quiescent disease. Details of lymphocyte isolation are detailed in the Supplementary Material. Ethical approval was obtained from the Oxfordshire Research Ethics Committee and informed written consent was given by all study patients.
Thymus samples were obtained after informed consent and ethics committee approval from the University of Oxford and Great Ormond Street Hospital, London.
Flow cytometry. Dead cells were excluded with the Near-IR Dead-Cell stain (Invitrogen, Paisley, UK). Antibodies used were: anti-CD3 PE-Cy7 or APC, anti-CD8 PerCP-Cy5.5 or eFluor 450, anti-IL18Rα PE, anti-CCR7 PE, anti-CD16 FITC, EOMES-eFluor660 (eBioscience, Hatfield, UK); anti-CD161 PE or APC, anti-CD8 VioGreen, anti-IFNγ FITC, anti-NKp80 PE (Miltenyi Biotec, Surrey, UK); anti-Vα7.2 PE or FITC or PE-Cy7, anti-Perforin Pacific Blue, anti-CXCR6 AlexaFluor647, anti-CCR7 PerCPCy5.5, anti-CD45RA PerCPCy5.5 or PECy7, anti-CD243/MDR1 PE, anti-CD56 BV421, anti-CD16 PECy7, anti-CD160 AlexaFluor647, anti-CD244 PerCPCy5.5 (Biolegend, London, UK); anti-GrB AlexaFluor700, anti-Perforin FITC, anti-RORγt PE, anti-CD103 FITC, anti-IFNγ AlexaFluor700, anti-IL-2 PerCPCy5.5 (BD Biosciences, Oxford, UK), anti-GrB APC (Invitrogen); anti-PLZF APC (R&D Systems, Abingdon, UK), anti-KLRG1 AlexaFluor488,47 and anti-T-bet FITC (Santa Cruz Biotechnology, Santa Cruz, TX). For transcription factor stainings, cells were stained with the Foxp3/Transcription Factor Staining Buffer Set (eBioscience). For staining of CXCR6, cells were incubated with CXCR6 antibody only for 10 min at 37 °C before further antibody staining.
For TCR typing, PBMCs were surfaced stained with Vβ antibodies: Vβ 1, 2, 4, 5.3, 7.2, 8, 9, 11, 13.2, 14, 16, 18, and 23-PE, and Vβ 3, 5.1, 5.2, 7.1, 12, 13.1, 13.6, 14, 16, 17, 20, 21.3, and 22-FITC (Beckman Coulter, High Wycombe, UK).
Antiviral responses were identified using MHC class I dextramers presenting NLVPMVATV (cytomegalovirus), GLCTLVAML (Epstein–Barr virus) or GILGFVFTL (influenza) (Immudex, Copenhagen, Denmark). Vaccine-induced responses were taken at the end of study (weeks 36 or 52) after vaccination, as described previously,35 and identified by pentamers loaded with hepatitis C virus NS3 1406 (KLSALGINAV) and hepatitis C virus NS3 (ATDALMTGY) (ProImmune, Oxford, UK).
Data were acquired on a MacsQuant (Miltenyi Biotec) or LSRII (BD Biosciences) and analyzed using FlowJo software (Treestar, Ashland, OR).
Cell stimulation. Phorbol 12-myristate 13-acetate and ionomycin stimulations were performed by adding 1 × cell stimulation cocktail (eBioscience; equivalent to 0.081 μM phorbol 12-myristate 13-acetate+1.34 μM ionomycin) to media containing 3 μg ml−1 Brefeldin A (eBioscience) and 2 μM monensin (eBioscience) for 3 h.
Rh123 efflux assay. PBMCs were loaded with 10 μg ml−1 Rh123 (Sigma, Dorset, UK) in RPMI+1% bovine serum albumin for 1 h on ice. Cells were resuspended in R10 with or without inhibitors, and either kept on ice (loading control) or cultured at 37 °C (efflux) for 30 min. MDR1 inhibitors cyclosporin A (Sigma) and verapamil hydrochloride (verapamil; Sigma) were added at the lowest inhibitory concentration; 0.5 and 50 μM, respectively. After efflux, cells were returned to ice and surface stained for analysis. Rh123 fluorescence was collected at 585/40 nm. Efflux was calculated as:

Microarray. CD8+ T cells were enriched from PBMCs using CD8 Microbeads (Miltenyi Biotec) and sorted on a MoFlo Legacy or MoFlo XDP (Beckman Coulter). Sorted cells were either snap frozen immediately (unstimulated array) or stimulated for 4 h (determined optimal, see Supplementary Figure S3) with cell stimulation cocktail (eBioscience) before snap freezing (stimulated array). Frozen cell pellets were sent to Miltenyi Genomic Services (Bergisch Gladbach, Germany) for RNA extraction and hybridization to Agilent Whole Human Genome Oligo Microarray (Agilent Technologies, Santa Clara, CA). The Agilent Feature Extraction Software was used to read out and process the microarray image files that were further analyzed using the Rosetta Resolver gene expression data analysis system (Rosetta Biosoftware, Seattle, WA). Genes considered to be significantly differentially expressed were those genes with a fold change >2 and P-value <0.01. Cord blood microarray analyses were performed as previously described.1 Heatmaps were generated using GENE-E (http://www.broadinstitute.org/cancer/software/GENE-E/index.html). GSEA was performed using GSEA version 2.0.14,48 with CD161hi/MAIT cell gene sets obtained from GSE62099.
Cytometry by-time-of flight. PBMCs were stimulated as above in the presence of 2.5 μg ml−1 anti-CD107α, 1.25 μg ml−1 anti-CD107b (BD Bioscience), and 10 μM TAPI-2 (VWR International, West Sussex, UK). Following stimulation, cells were resuspended in cytometry buffer (phosphate-buffered saline+0.05% sodium azide+2 mM EDTA+2% fetal calf serum) and stained with isotope-tagged antibodies before acquisition on the cytometry by time-of-flight as previously described.19
Statistical analysis. Statistical analysis was performed using Prism version 6 software (GraphPad, La Jolla, CA). Data are represented as mean±s.e.m. *P<0.05, **P<0.01, ***P<0.001, ****P<0.0001, NS, not significant P>0.05, as stated in figure legends.
References
Walker, L.J. et al. Human MAIT and CD8 alpha alpha cells develop from a pool of type-17 precommitted CD8+ T cells. Blood 119, 422–433 (2012).
Gebhardt, T. et al. Memory T cells in nonlymphoid tissue that provide enhanced local immunity during infection with herpes simplex virus. Nat. Immunol. 10, 524–530 (2009).
Jiang, X. et al. Skin infection generates non-migratory memory CD8+ T(RM) cells providing global skin immunity. Nature 483, 227–231 (2012).
Masopust, D., Vezys, V., Wherry, E.J., Barber, D.L. & Ahmed, R. Cutting edge: gut microenvironment promotes differentiation of a unique memory CD8 T cell population. J. Immunol. 176, 2079–2083 (2006).
Lanier, L.L., Chang, C. & Phillips, J.H. Human NKRP1A. A disulfide-linked homodimer of the C-type lectin superfamily expressed by a subset of NK and T lymphocytes. J. Immunol. 153, 2417–2428 (1994).
Takahashi, T., Dejbakhsh-Jones, S. & Strober, S. Expression of CD161 (NKR-P1A) defines subsets of human CD4 and CD8 T cells with different functional activities. J. Immunol. 176, 211–216 (2006).
Ishihara, S. et al. CD8+NKR-P1A+ T cells preferentially accumulate in human liver. Eur. J. Immunol. 29, 2406–2413 (1999).
O’Keeffe, J. et al. Diverse populations of T cells with NK cell receptors accumulate in the human intestine in health and in colorectal cancer. Eur. J. Immunol. 34, 2110–2119 (2004).
Billerbeck, E. et al. Analysis of CD161 expression on human CD8+ T cells defines a distinct functional subset with tissue-homing properties. Proc. Natl. Acad. Sci. USA 107, 3006–3011 (2010).
Dusseaux, M. et al. Human MAIT cells are xenobiotic-resistant, tissue-targeted, CD161hi IL-17-secreting T cells. Blood 117, 1250–1259 (2011).
Tilloy, F. et al. An invariant T cell receptor alpha chain defines a novel TAP-independent major histocompatibility complex class Ib-restricted alpha/beta T cell subpopulation in mammals. J. Exp. Med. 189, 1907–1921 (1999).
Treiner, E. et al. Selection of evolutionarily conserved mucosal-associated invariant T cells by MR1. Nature 422, 164–169 (2003).
Reantragoon, R. et al. Antigen-loaded MR1 tetramers define T cell receptor heterogeneity in mucosal-associated invariant T cells. J. Exp. Med. 210, 2305–2320 (2013).
Aldemir, H. et al. Cutting Edge: lectin-like transcript 1 is a ligand for the CD161 receptor. J. Immunol. 175, 7791–7795 (2005).
Rosen, D.B. et al. Functional consequences of interactions between human NKR-P1A and its ligand LLT1 expressed on activated dendritic cells and B cells. J. Immunol. 180, 6508–6517 (2008).
Germain, C. et al. Induction of lectin-like transcript 1 (LLT1) protein cell surface expression by pathogens and interferon-γ contributes to modulate immune responses. J. Biol. Chem. 286, 37964–37975 (2011).
Exley, M., Porcelli, S., Furman, M., Garcia, J. & Balk, S. CD161 (NKR-P1A) costimulation of CD1d-dependent activation of human T cells expressing invariant Vα24JαQ T cell receptor α chains. J. Exp. Med. 188, 867–876 (1998).
Maggi, L. et al. CD161 is a marker of all human IL-17-producing T-cell subsets and is induced by RORC. Eur. J. Immunol. 40, 2174–2181 (2010).
Fergusson, J.R. et al. CD161 defines a transcriptional and functional phenotype across distinct human T cell lineages. Cell Rep. 9, 1–14 (2014).
Ussher, J.E. et al. CD161++CD8+ T cells, including the MAIT cell subset, are specifically activated by IL-12+IL-18 in a TCR-independent manner. Eur. J. Immunol. 44, 195–203 (2014).
Sallusto, F., Lenig, D., Forster, R., Lipp, M. & Lanzavecchia, A. Two subsets of memory T lymphocytes with distinct homing potentials. Nature 401, 708–712 (1999).
Martin, E. et al. Stepwise development of MAIT cells in mouse and human. PLoS Biol. 7, e54 (2009).
Sathaliyawala, T. et al. Distribution and compartmentalization of human circulating and tissue-resident memory T cell subsets. Immunity 38, 187–197 (2013).
Tang, X.-Z. et al. IL-7 licenses activation of human liver intrasinusoidal mucosal-associated invariant T cells. J. Immunol. 190, 3142–3152 (2013).
Cepek, K.L. et al. Adhesion between epithelial cells and T lymphocytes mediated by E-cadherin and the alphaEbeta7 integrin. Nature 372, 190–193 (1994).
Mackay, L.K. et al. The developmental pathway for CD103(+)CD8+ tissue-resident memory T cells of skin. Nat. Immunol. 14, 1294–1301 (2013).
Schenkel, J.M. & Masopust, D. Tissue-resident memory T cells. Immunity 41, 886–897 (2014).
Turtle, C.J., Swanson, H.M., Fujii, N., Estey, E.H. & Riddell, S.R. A distinct subset of self-renewing human memory CD8+ T cells survives cytotoxic chemotherapy. Immunity 31, 834–844 (2009).
Neyfakh, A.A., Serpinskaya, A.S., Chervonsky, A.V., Apasov, S.G. & Kazarov, A.R. Multidrug-resistance phenotype of a subpopulation of T-lymphocytes without drug selection. Exp. Cell. Res. 185, 496–505 (1989).
Appay, V., van Lier, R.a.W., Sallusto, F. & Roederer, M. Phenotype and function of human T lymphocyte subsets: consensus and issues. Cytometry A 73, 975–983 (2008).
Kurioka, A. et al. MAIT cells are licensed through granzyme exchange to kill bacterially sensitized targets. Mucosal Immunol. 8, 429–440 (2015).
Sullivan, B.M., Juedes, A., Szabo, S.J., von Herrath, M. & Glimcher, L.H. Antigen-driven effector CD8 T cell function regulated by T-bet. Proc. Natl. Acad. Sci. USA 100, 15818–15823 (2003).
Pearce, E.L. et al. Control of effector CD8+ T cell function by the transcription factor Eomesodermin. Science 302, 1041–1043 (2003).
Cruz-Guilloty, F. et al. Runx3 and T-box proteins cooperate to establish the transcriptional program of effector CTLs. J. Exp. Med. 206, 51–59 (2009).
Barnes, E. et al. Novel adenovirus-based vaccines induce broad and sustained T cell responses to HCV in man. Sci. Transl. Med. 4, 1–11 (2012).
Zimmerli, S.C. et al. HIV-specific IFN-gamma/IL-2-secreting CD8 T cells support CD4-independent proliferation of HIV-1-specific CD8 T cells. Proc. Natl. Acad. Sci. USA 102, 7239–7244 (2005).
Harari, A. et al. Functional signatures of protective antiviral T-cell immunity in human virus infections. Immunol. Rev. 211, 236–254 (2006).
Ledgerwood, J.E. et al. Chimpanzee adenovirus vector ebola vaccine - preliminary report. N. Engl. J. Med. advance online publication, 26 November 2014 doi:10.1056/NEJMoa1410863.
Ussher, J.E., Klenerman, P. & Willberg, C.B. Mucosal-associated invariant T-cells: new players in anti-bacterial immunity. Front. Immunol. 5, 450 (2014).
Schenkel, J.M. et al. Resident memory CD8 T cells trigger protective innate and adaptive immune responses. Science 346, 98–101 (2014).
Ariotti, S. et al. Skin-resident memory CD8+ T cells trigger a state of tissue-wide pathogen alert. Science 346, 101–105 doi:10.1126/science.1254803 (2014).
Kleinschek, M.A. et al. Circulating and gut-resident human Th17 cells express CD161 and promote intestinal inflammation. J. Exp. Med. 206, 525–534 (2009).
Ramesh, R. et al. Pro-inflammatory human Th17 cells selectively express P-glycoprotein and are refractory to glucocorticoids. J. Exp. Med. 211, 89–104 (2014).
Cosmi, L. et al. Human interleukin 17–producing cells originate from a CD161+CD4+ T cell precursor. J. Exp. Med. 205, 1903–1916 (2008).
Northfield, J.W. et al. CD161 expression on hepatitis C virus-specific CD8+ T cells suggests a distinct pathway of T cell differentiation. Hepatology 47, 396–406 (2008).
Saghafian-Hedengren, S., Sverremark-Ekström, E., Linde, A., Lilja, G. & Nilsson, C. Early-life EBV infection protects against persistent IgE sensitization. J. Allergy Clin. Immunol. 125, 433–438 (2010).
Marcolino, I. et al. Frequent expression of the natural killer cell receptor KLRG1 in human cord blood T cells: correlation with replicative history. Eur. J. Immunol. 34, 2672–2680 (2004).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Acknowledgements
J.R.F is supported by the Wellcome Trust IITM Programme (092871/Z/10/Z). This work was also supported by the Wellcome Trust (WT091663MA), the Medical Research Council, the NIHR Biomedical Research Centre (Oxford), the Nuffield Department of Clinical Medicine (Oxford), the James Martin School for the 21st Century (Oxford), the NIH (NIAD U19AI 082630), and the Oxford Dominions Trust. G.H. is supported by the NIHR Biomedical Research Centre.
Author contributions
J.R.F. designed the study, analyzed data and drafted the manuscript; M.H.H., L.S., L.J.W., A.K., E.W.N., and J.E.U. contributed and analyzed critical data to this study; A.B., G.H., M.M.D., E.S-E., F.P., S.C., A.F., and E.B. provided essential tissue and cell samples; A.L. provided technical assistance; C.B.W. and J.E.U. provided technical and supervisory support; P.K. designed and supervised the study and manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declared no conflict of interest.
Additional information
SUPPLEMENTARY MATERIAL is linked to the online version of the paper
Rights and permissions
About this article
Cite this article
Fergusson, J., Hühn, M., Swadling, L. et al. CD161intCD8+ T cells: a novel population of highly functional, memory CD8+ T cells enriched within the gut. Mucosal Immunol 9, 401–413 (2016). https://doi.org/10.1038/mi.2015.69
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mi.2015.69
This article is cited by
-
CD161 expression defines new human γδ T cell subsets
Immunity & Ageing (2022)
-
High-dimensional profiling reveals Tc17 cell enrichment in active Crohn’s disease and identifies a potentially targetable signature
Nature Communications (2022)
-
Systems analysis and controlled malaria infection in Europeans and Africans elucidate naturally acquired immunity
Nature Immunology (2021)
-
Local heroes or villains: tissue-resident memory T cells in human health and disease
Cellular & Molecular Immunology (2020)
-
Mucosal-associated invariant T cells and disease
Nature Reviews Immunology (2019)