Abstract
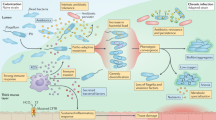
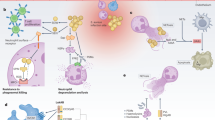
Methicillin-resistant Staphylococcus aureus (MRSA) causes chronic pulmonary infections in patients with cystic fibrosis (CF). This study tracks the 13-year evolution (1996–2009) of a single MRSA clone in a male patient with CF, evaluating both the host immunogenic response and the microbial variations. Whole-genome sequencing was performed for the initial (CF-96) and evolved (CF-09) isolates. The immunogenicity of CF-96 and CF-09 was evaluated by incubation with innate immune cells from healthy volunteers. We also studied the patient’s innate immune response profile, cytokine production, expression of triggering receptor expressed on myeloid cells-1 (TREM-1), and phagocytosis. A total of 30 MRSA ST247-SCCmecI-pvl− isolates were collected, which evidenced a genome size reduction from the CF-96 ancestor to the evolved CF-09 strain. Up to six changes in the spa-type were observed over the course of the 13-year evolution. Cytokine production, TREM-1 expression, and phagocytosis were significantly lower for the healthy volunteer monocytes exposed to CF-09, compared with those exposed to CF-96. Patient monocytes exhibited a reduced inflammatory response when challenged with CF-09. Genetic changes in MRSA, leading to reduced immunogenicity and entry into the refractory state, may contribute to the attenuation of virulence and efficient persistence of the bacteria in the CF lung.
Similar content being viewed by others
Introduction
Methicillin-resistant Staphylococcus aureus (MRSA) commonly causes chronic pulmonary infections in patients with cystic fibrosis (CF). The prevalence of MRSA varies, and the incidence of MRSA has increased significantly within our institution and other institutes over the past 15 years.1, 2, 3
CF-MRSA isolates frequently exhibit a hypermutational phenotype, an increased ability to form biofilm, production of small colony variants, and high antimicrobial resistance rates.1, 2, 4 These features are further facilitated by CF-lung conditions and bacterial adaptive processes.
Surprisingly, CF pathogens have low invasive capability, which is probably due to gene mutations in the host’s CF transmembrane conductance regulator that prevent the internalization of bacteria by epithelial cells. In contrast, their complete eradication is almost impossible in CF once the colonization is established.4, 5 It is difficult to distinguish between colonization and infection, a situation referred to as “pathogenic colonization”.6 MRSA-positive cultures are commonly seen in sputum samples, whereas respiratory acute infection and bacteremia are unusual, a phenomenon known as attenuation.7 During attenuation, microorganisms and the innate immune system fight a battle in which bacterial virulence traits and the innate immune response must be downregulated to ensure their coexistence.
Several studies have suggested that the innate immune response in CF is compromised.8 We have published a number of studies that agree with this hypothesis, demonstrating patent endotoxin tolerance in circulating immune cells obtained from patients with CF.9, 10, 11 This phenomenon, defined as the inability of innate immune cells to mount a standard inflammatory response after bacterial colonization,12, 13 is not restricted to patients with CF.14, 15, 16 This response is “locked” into an endotoxin tolerance state, owing to endotoxin diffusion into the bloodstream and a notable downregulation of the transmembrane receptor known as the triggering receptor expressed on myeloid cells-1 (TREM-1).9, 10, 11 This response could be beneficial for the lifespan of patients with CF, because it allows patients to orchestrate a moderate innate response against pathogen colonization. Patients with CF could therefore avoid a permanent “cytokine storm” that may endanger their lives.
Considering the particularities of MRSA isolates colonizing CF lungs and the tolerance of the innate immune system of these patients, the purpose of our study was to analyze the long-term evolution of MRSA in vivo for a single patient with CF, evaluating the microbiological aspects, and the immunogenicity characteristics.
Results
Clinical and microbiological features
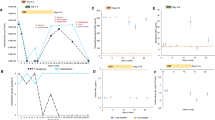
The clinical management of this patient during the 13-year study period was challenging. The forced expiratory volume in 1-s index decreased from 53 to 23% and, consequently, affected his quality of life (Figure 1). He had numerous lung exacerbations (3–4 per year, range 1–9), which were treated with oral cycles of trimethoprim/sulfamethoxazole, ciprofloxacin, fosfomycin, and rifampin. The inhaled antibiotic therapy was based on vancomycin,17 which was maintained almost without interruption during the 13 years of the study, combined with tobramycin or colistin to ensure protection against Pseudomonas aeruginosa coinfection (Figure 1).
Clinical evolution and antibiotherapy of the patient during the study period. Progression of the FEV1 values related to antibiotic consumption and clinical incidences. AMX/CLA, amoxicillin-clavulanate; AZM, azithromycin; AZT, aztreonam; CIP, ciprofloxacin; COL, colistin; FEV1, forced expiratory volume in 1 s; FOS, fosfomycin; LEV, levofloxacin; MOX, moxifloxacin; RIF, rifampin; SxT, cotrimoxazole; TOB, tobramycin; VAN, vancomycin. The number in the brackets corresponds to the number of cycles. 1The inhaled preparation was the intravenous solution. 2Uninterrupted oral treatment with 500 mg of azithromycin per 48 h.
Sputum cultures showed chronic MRSA lung colonization, with the sporadic presence of nonpersistent P. aeruginosa, methicillin-susceptible S. aureus, Haemophilus influenzae and Aspergillus fumigatus. A total of 30 MRSA isolates were sequentially recovered, showing a closely related pulsed-field gel electrophoresis pattern (Figure 2). Most of the isolates exhibited an antibiotic multiresistant phenotype, characterized by aminoglycoside, quinolone, and macrolide nonsusceptibility. A small reduction in glycopeptide susceptibility was observed from the initial CF-96 isolate (Minimum inhibitor concentration 2 μg ml–1 for vancomycin and 0.5 μg ml–1 for teicoplanin) to the final CF-09 isolate (3 μg ml–1 for vancomycin and 1 μg ml–1 for teicoplanin). Both CF-06 and CF-09 were able to grow in the presence of 8 μg ml–1 of both glycopeptides after in vitro progressive induction. Molecular typing experiments revealed that all 30 isolates corresponded to ST247-SCCmecI-pvl−.
Clonality of the isolates and spa-type evolution. PFGE-SmaI of 28 MRSA isolates obtained from the same patient with CF over 13 years. The first and last lines correspond to the molecular marker. The year of isolation and the corresponding spa-type detected phenotype are marked for each isolate. CF, cystic fibrosis; MRSA, methicillin-resistant S. aureus; PFGE, pulsed-field gel electrophoresis.
Evolution of the MRSA strain within the 13-year timeframe
The spa-type changed up to six times during the study (Figure 2), and five of the six spa-types were grouped in the same spa-clonal complex (spa-CC051). All isolates showed slow growth compared with the growth of their genetic counterpart, the HARMONY control strain. The growth of a colony variant with a high degree of antibiotic dependence was detected in 1997 (Figure 3). Whole-genome sequencing of MRSA CF-96 (5,967 genes) and CF-09 (5,901 genes) revealed the total loss of 66 genes for CF-09, with changes in the chromosomal disposition and an overall reduction in genome size by 200 kb (Figure 4; Supplementary Tables 1–3 online; and Supplementary Figure 1).
Bacterial growth deficiencies. (a) Poor growth of the CF-96 isolate in comparison with the HARMONY MRSA-ST247 control strain after 24 h of incubation at 37 °C. (b) Antibiogram in Mueller–Hinton of this growth-deficient isolate. CF, cystic fibrosis; MRSA, methicillin-resistant S. aureus.
Genomes differences. (a) Comparison of the CF-96 and CF-09 MRSA genomes, which were sequenced with the S. aureus MSHR1132 strain using BRIG software. (b) Comparison of the pST75 plasmid with plasmid sequences found in our isolates. CF, cystic fibrosis; MRSA, methicillin-resistant S. aureus.
The draft genome sequence of the MRSA CF-96 contains a total of 29 contigs and 4 scaffolds that correspond to 93% of its chromosome. The genome of the MRSA CF-09 isolate was mapped as a function of its ancestor, the CF-96 isolate. A single plasmid was detected for both strains and was only represented in two scaffolds, one of which contained 95% of the plasmid. The first CF-96 isolate carried three prophages, whereas only two were detected for the last CF-09 isolate (Figure 4 and Supplementary Tables 1–3).
There were numerous differences between CF-96 and CF-09. The most relevant attenuation-related changes were amino-acid alterations in the cardiolipin synthase protein codified by cls1 (Phe→Leu),18 the protein coded by purD involved in purine biosynthesis and SaeSR regulation (Gly→Asp),19 and the staphylococcal secretory antigen (ssaA1; Gln→Lys). In addition, a number of virulence genes were deleted in the evolved isolate, such as oppF2 belonging to the nickel/peptide/opine PepT subfamily of ABC-transporters and the YMC/09/04/R1988 SPP1 family prophage.20
CF-96 and CF-09 provoked various innate responses by human monocytes
The genetic differences observed between the initial and evolved MRSA strains suggest an adaptation process by which the MRSA strains became “invisible” to the innate immune response. When human circulating monocytes from healthy volunteers were exposed to both isolates (CF-96 and CF-09) in vitro, the inflammation was lower for those cells treated with the evolved strain (Figures 5 and 6). The interleukin (IL)-6, IL-12, extracellular tumor necrosis factor α (TNFα), and intracellular TNFα levels were significantly lower in the presence of CF-09. A similar tendency was observed for IL-1β but not for IL-10. Curiously, the inflammation generated after exposure to the genetically similar HARMONY strains was even higher in comparison with the response mounted by the first MRSA isolate (CF-96).
Differential inflammatory response against CF-96 and CF-09. Human monocyte cultures from healthy volunteers were challenged with a freshly prepared sonicated and homogenized crude extract of CF-96, CF-09, and the reference strain HARMONY ST247-MRSA (Harmony) for the indicated time; final protein concentration: 100 ng ml–1. Levels of (a) IL-1β, (b) IL-6, (c) IL-10, and (d) IL-12p70 were quantified in the supernatant from monocyte cultures using CBA and flow cytometry (n=3, ***P<0.005, **P<0.01, *P<0.05). CBA, cytometric bead array; CF, cystic fibrosis; IL, interleukin; MRSA, methicillin-resistant S. aureus; NS, not significant.
Extracellular and intracellular levels of TNFα. Human monocytes were treated with CF-96, CF-09, and HARMONY as in Figure 4. (a) Levels of TNFα in the supernatant were analyzed by CBA (n=3, ***P<0.005, *P<0.05). (b) After 16 h of incubation, cells were harvested, fixed, permeabilized, stained extracellularly with CD14-FITC and intracellularly with TNFα-APC and analyzed by flow cytometry. The TNFα g-mean of CD14-positive cells is shown (n=2, *P<0.05). APC, allophycocyanin; CBA, cytometric bead array; CF, cystic fibrosis; FITC, fluorescein isothiocyanate; NS, not significant; TNFα, tumor necrosis factor α.
The mRNA expression levels of TNFα and IL-12 were further investigated. The findings confirmed our observations made at the protein level (Figure 7a and b). Interestingly, both Toll-like receptor 2 (TLR2) and TLR4, the main receptors for Gram-positive and Gram-negative bacteria, were not affected (Supplementary Figure 2). In contrast, mRNA expression of the pseudokinase IL-1 receptor-associated kinase-M, a negative regulator of inflammation, was higher in monocytes exposed to the MRSA CF-09 than in those exposed to CF-96 (Figure 7c). These data were also confirmed at the protein level (data not shown). In line with these findings, there was less TREM-1 mRNA in monocytes challenged with CF-09 than in those exposed to CF-96 (Figure 7d).
Transcriptional analysis. Human monocytes were treated with CF-96, CF-09, and HARMONY for 3 h; final protein concentration: 100 ng ml–1. Cells were then harvested, total RNA was purified, and complementary DNA was synthesized. mRNA levels of (a) TNFα, (b) IL-12, (c) IRAK-M, and (d) TREM-1) were analyzed by real-time quantitative PCR (n=3, ***P<0.005, **P<0.01, *P<0.05). CF, cystic fibrosis; IL, interleukin; IRAK-M, IL-1 receptor-associated kinase-M; NS, not significant; TNFα, tumor necrosis factor α; TREM-1, triggering receptor expressed on myeloid cells-1.
TREM-1 is an important immune response element that strongly enhances leukocyte inflammation in the presence of microbial products.21 We therefore performed an in-depth analysis of its expression using monocytes from healthy volunteers after a challenge with HARMONY, CF-96, or CF-09. The reference HARMONY strain and CF-96 induced strong TREM-1 expression on the cell surface after 5 h of incubation. However, incubation with the CF-09 strain did not lead to an increase in TREM-1 basal levels (Figure 8a and b).
TREM-1 and sTREM-1 expression. Human monocytes were treated with CF-96, CF-09, and HARMONY for the indicated time; final protein concentration: 100 ng ml–1. (a, b) Cells were harvested and stained with CD14-APC and TREM-1-FITC, followed by flow cytometry analysis. (a) The percentages of TREM-1/CD14-positive cells are shown (n=3). (b) TREM-1 MFI of CD14-positive cells are shown (n=3). (c) sTREM-1 was quantified in the culture supernatants by ELISA (n=3). APC, allophycocyanin; CF, cystic fibrosis; ELISA, enzyme-linked immunosorbent assay; FITC, fluorescein isothiocyanate; MFI, median fluorescence intensity; soluble TREM-1; sTREM-1, TREM-1, triggering receptor expressed on myeloid cells-1.
TREM-1 undergoes a proteolytic cleavage (shedding) of its mature cell surface-anchored form after 5–7 h of bacterial exposure, generating a soluble form known as soluble TREM-1 (sTREM-1).21 As expected, 16 h of HARMONY or CF-96 challenge provoked a significant reduction in the membrane-anchored TREM-1, resulting in high levels of sTREM-1 (Figure 8c). The same challenge with CF-09, however, affected neither TREM-1 nor sTREM-1 levels. These data demonstrate that the evolved CF-09 MRSA strain gives rise to a significantly reduced inflammatory response.
Differential phagocytosis of CF-96 and CF-09 strains by “healthy” monocytes
The most important role of monocytes is pathogen phagocytosis. Given that our data indicated a patent differential innate response against CF-96 and CF-09, we proceeded to study how these strains affect monocyte phagocytosis. When human monocyte cultures from healthy volunteers were exposed to CF-96, CF-09, and HARMONY, a marked decrease in phagocytosis was observed (Figure 9). CF-96 was less phagocytized than HARMONY, while CF-09 was the least phagocytosed.
The phagocytosis of CF-09 is impaired. Monocytes-derived macrophages from healthy volunteers were exposed to CF-96, CF-09, and HARMONY (MOI=5) for 2 h. Next, the cells were washed and treated with gentamicin. Then, the cells were lysed, their cytosols were spread on agar dishes, and incubated overnight. CFUs were counted, and the number of CFUs is shown (n=8, *P<0.03). CF, cystic fibrosis; CFU, colony-forming unit; MOI, multiplicity of infection.
The patient’s innate immune system differentially modulates its response to CF-09 and CF-96
In addition to the evident genetic “evolution” of the MRSA isolates, which correlated with a downregulation of immunogenicity, the patient’s own innate immune system also showed a reduced response to bacterial stimuli. Monocytes isolated from the patient in 2009 were exposed to HARMONY, CF-96, and CF-09. The analysis of cytokine production revealed a low level of inflammation after CF-09 exposure (Figure 10). Compared with cells challenged with CF-96 or HARMONY, monocytes challenged with CF-09 had significantly lower protein levels of IL-1β, IL-6, and TNFα. CF-09-challenged monocytes also showed significantly lower IL-12 protein levels when compared with HARMONY-challenged cells. These data were further confirmed at the mRNA level (data not shown).
Modulation of the patient innate response. Monocytes isolated from the patient in 2009 were challenged with CF-96, CF-09, and HARMONY (H) for 16 h; final concentration: 100 ng ml–1. Supernatant levels of (a) IL-1β, (b) IL-6, (c) IL-12p70, and (d) TNFα were quantified using CBA. The mean of two replicas is shown. CBA, cytometric bead array; CF, cystic fibrosis; IL, interleukin; TNFα, tumor necrosis factor α.
Curiously, cytokine production was low even in the presence of HARMONY ST247-MRSA. In line with the endotoxin tolerance status, when we compared the inflammatory response provoked by HARMONY on either monocytes from the patient or those from healthy controls, we confirmed that monocytes from the patient were “locked” into a refractory state (Supplementary Figure 3A–D). In agreement with our previous data, TREM-1 expression was also impaired in monocytes from patients (Supplementary Figure 3E and F).
Discussion
The molecular epidemiology of MRSA isolates has been defined worldwide by various techniques such as pulsed-field gel electrophoresis, multilocus sequence typing, spa typing and Staphylococcal cassette chromosome mec (SCCmec) typing. These techniques have demonstrated that only a few MRSA lineages are related to human infections.22 In recent years, the prevalence of MRSA CF-lung colonization has increased. MRSA CF-lung colonization is associated with significant tissue damage and a poorer disease prognosis.1, 2, 3, 4 Aggressive antimicrobial therapies (mostly delivered by inhalation) are prescribed at the MRSA diagnosis or during exacerbations but have little effect on bacterial eradication. Bacterial colonization from the same genetic lineage usually persists for years,4, 5 and various MRSA lineages are reported for patients with CF. The most frequent clone in our unit is ST228-SCCmecI-pvl−, also known as the German clone.3
Despite the aggressive antibiotic strategy used for the patient described in this study, clinicians alerted us to his poor clinical management. Antibiotic susceptibility tests showed an antibiotic multiresistant phenotype with possible heteroresistance to glycopeptides. The administration of inhaled vancomycin remained uninterrupted during the 13 years of treatment. All 30 recovered MRSA isolates corresponded to the Iberian clone ST247-SCCmecIV-pvl−, with high nucleotide sequence conservation in their multilocus sequence typing alleles. However, the emergence of new sequence types has been reported for CF-associated P. aeruginosa isolates under similar conditions.23 We were able to detect unusual transient S. aureus variants presenting tetracycline-dependent growth. This phenotype has been previously described in Staphylococcus epidermidis.24
During the adaptation to CF lungs, microbial virulence attenuation is necessary for successful persistence. This attenuation has been previously reported for P. aeruginosa.25 S. aureus evolves in chronically colonized CF patients by modulating factors necessary for persistence rather than virulence.26 Nucleotide mutations in the spa gene have been an effective evasion strategy and are also able to condition the host’s adaptive immune response.27, 28 In our case, up to six changes in the spa-type were observed over 13 years. The observed chromosomal reduction with the loss of 66 genes indicates niche adaptation. The attenuation process is also evidenced by the changes in cardiolipin synthase CL18 and the regulation of SaeSR,19 the staphylococcal secretory antigen ssaA1 and the oppF2-ABC-transporter.20
Both proinflammatory cytokine production and phagocytosis were significantly impaired when monocytes were incubated with the MRSA-evolved isolate CF-09. Immunology studies indicated a patent reduction of the innate response against CF-09, possibly due to IL-1 receptor-associated kinase-M upregulation and an impaired expression of TREM-1. These findings indicate that the innate immune cells from the patient were in a refractory state. The loss of MRSA immunogenicity combined with the refractory state of the monocytes from the patient may contribute to the “coexistence” of the host and the pathogen. As has been previously reported, patients with CF show a refractory state known as endotoxin tolerance.9, 10, 11
In summary, we have described a long-term evolutionary scenario that illustrates the in vivo “battle for coexistence” between MRSA and the long-term colonized/infected host, who is continuously exposed to antibiotics. MRSA adapts with genetic reductions and mutations of surface proteins, while monocytes adapt their response by becoming more tolerant. The adaptations by MRSA and monocytes work toward the final objective of survival under equilibrium conditions. With these findings in mind, the patient’s immune status should also be considered for MRSA treatment, especially when dealing with eradication protocols.
Methods
Patient and healthy volunteers. A male patient with CF was followed-up from 1996 to 2009 at the Cystic Fibrosis Unit of the Ramón y Cajal University Hospital. This patient was born in 1987 and was diagnosed at 30 months of age with a homozygotic ΔF508 genotype. Chronic MRSA colonization was established at 9 years of age, with 2–3 exacerbations per year. He was also diagnosed with diabetes mellitus and hepatic fibrosis. Eight age-matched healthy volunteers with no personal history of CF or other significant disease were included as controls. Written informed consent for the study was obtained from all enrolled participants. This study was approved by the local ethics committee.
Bacterial isolates and antibiotic susceptibility. Regular sputum samples were obtained over a period of 13 years (1996–2009) at routine check-ups (3–4 per year) or during exacerbations. Bacterial identification was performed using the semiautomatic Wider system (Fco. Soria Melguizo, Madrid, Spain) and the matrix-assisted laser desorption/ionization time-of-flight mass spectrometer (Bruker, Leizpig, Germany). Antibiotic susceptibility was performed using the agar dilution method. The HARMONY ST247-MRSA strain corresponded to the same genetic lineage as that of our strain and was used as the reference for all experiments.
Biofilm formation. Dense, population-size biofilms were initially established over 24 h at 37 °C in a static model within a nitrocellulose filter disk (25 mm in diameter; Millipore, Billerica, MA). These biofilms were then deposited onto 7% sheep blood agar commercial plates (Oxoid, Hampshire, England) and inoculated overnight with 100 μl of brain–heart infusion broth culture. After incubating for 72 h, the filter containing the biofilm-grown bacteria was suspended in saline solution, sonicated, homogenized with a vortex mixer, and passed 15 times through a 30-gauge needle to prepare suitable inoculums.29, 30 Samples were then analyzed using microscopy, and the size of the bacteria was determined by a Leica microscope (software version 2.6.0; Wetzlar, Germany). The concentration of the bacteria was adjusted by densitometry, and multiplicity of infection was calculated for phagocytosis assays. The protein concentrations of the crude extracts were obtained using a commercially available kit (Bio-Rad protein assay, Bio-Rad, Birmingham, UK).
SCC mec typing and panton-valentine leukocidin ( pvl ) detection. A multiplex polymerase chain reaction (PCR) scheme was used to determine the SCCmec, according to a previously described method.31 The presence of pvl genes was confirmed by a specific PCR using the MRSA ATCC 49775 strain as the positive control.
DNA sequencing of the polymorphic X region of the protein A gene ( spa ) gene. The spa gene was amplified from all isolates, as previously described.32 Nucleotide sequences were analyzed using the Ridom StaphType software (Ridom GmbH, Würzburg, Germany).33 The Based Upon Repeat Pattern algorithms,34 as implemented in the Ridom StaphType software, were used to cluster spa-types (spa-CC) in MRSA and methicillin-sensitive S. aureus isolates.24
Clonal relatedness. Pulsed-field gel electrophoresis-SmaI was applied to analyze the genetic relatedness of all isolates using a CHEF DR-III apparatus (Bio-Rad) with the Lambda Ladder PFGE Marker (New England Biolabs, Beverly, MA), under the HARMONY protocol.35 Isolates were later typed by multilocus sequence typing using the scheme developed by Enright et al.36 Alleles of each locus were compared, and sequence types were assigned based on the S. aureus multilocus sequence typing database (http://saureus.mlst.net).
Genome sequencing. The first (1996) MRSA isolate (CF-96) was pyrosequenced de novo using Illumina technology (GACT, Constance, Germany, Switzerland) in a unique step following the standard BG7 pipeline, annotated using the BG7 system we designed especially to handle NGS data 454 paired end reads, circled by Era7 Bioinformatics (http://era7bioinformatics.com/), and registered in the NCBI database under the accession number PRJNA124067. The evolved (2009) MRSA isolate (CF-09) was also pyrosequenced and circularized in relation to CF-96, using the Newbler software (Roche, Branford, CT). CF genomes were compared with the most similar types found in the databases (S. aureus MSHR1132 with the plasmid ST75) using the BLAST Ring Image Generator (BRIG) software (Brisbane, Australia). Assignation of the clusters of orthologous groups (COGs) was undertaken with the BLAST algorithm using the NCBI database (http://www.ncbi.nlm.nih.gov/COG/).
Human monocytes isolation and culture. Peripheral blood mononucleated cells were isolated from the blood of healthy volunteers and the patient with CF by centrifugation on Ficoll-Hypaque Plus (Amersham Biosciences, Eindhoven, The Netherlands), following previously published protocols.10, 29 The purity of all cultures was verified based on staining for CD14 (on average 90% CD14+ purity), CD89, CD1a, and CD16b (Supplementary Figure 1A). Cells were cultured in the presence or absence of CF-96, CF-09, and the HARMONY ST247-MRSA reference strain previously grown as biofilm, sonicated, and homogenized, as described previously.29
Fluorescent-activated cell sorting analysis of cell surface expression of CD14, TREM-1, and TLR4/MD2. Monocytes from healthy volunteers and patients were washed with phosphate-buffered saline and incubated with anti-CD14, anti-TLR2, or anti-TLR4/MD2 antibodies, followed by an anti-goat fluorescein isothiocyanate-conjugated secondary polyclonal antibody (Jackson ImmunoResearch, Baltimore, MD). To correct nonspecific binding, appropriate isotype control antibodies were used. For double staining (CD14 and TREM-1), cells were incubated with a CD14-allophycocyanin conjugate (Miltenyi, Bergisch Gladbach, Germany). Samples were analyzed by flow cytometry using a BD FACSCalibur flow cytometer (BD Biosciences, San Diego, CA), and the data were analyzed with FlowJo software (Ashland, OR).
Intracellular staining of TNFα. Cells were fixed, permeabilized with 100 μl of fluorescent-activated cell sorting lysis solution (10x; Becton Dickinson, San Jose, CA), washed and extracellularly stained (CD14-FITC Inmunostep, Salamanca, Spain). Cells were then washed twice with phosphate-buffered saline, incubated with the intracellular antibody TNFα-allophycocyanin (Becton Dickinson) for 30 min in the dark at room temperature, and analyzed by flow cytometry.
Cytometric bead array. Cytokine levels in the culture supernatants were determined using the CBA Flex Set (BD Biosciences), following the manufacturer’s protocol.
Phagocytosis assay. The phagocytosis assay was performed as previously described.37, 38 The macrophages, obtained from human monocytes after 7 days of differentiation by plastic attachment, were exposed to bacteria for 2 h (multiplicity of infection=5). The cells were then washed and kept with 300 μg ml–1 of gentamicin (Normon SA; Tres Cantos, Madrid, Spain) for 30 min. Phagocytosis was then analyzed by counting the colony-forming units generated when the cytosol of these cells was spread on blood agar plates.
mRNA isolation and quantification. After the cells and bacteria were incubated, the mix was washed once with phosphate-buffered saline, and RNA was isolated using TRI-Reagent (IMICO, Cincinnati, OH). Purified RNA was treated with RNase-free DNase I (Amersham Biosciences), and complementary DNA was synthesized by reverse transcription of 1-μg RNA using a poly(dT) oligonucleotide primer (Roche, Palo Alto, CA). The expression levels of TNFα, IL-12, IL-1 receptor-associated kinase-M, and TREM-1 were analyzed by real-time quantitative PCR (LightCycler; Roche Diagnostics, Indianapolis, IN) using a Fast-Start DNA master SYBR Green system (Roche Diagnostics) and specific primers, as described previously.10
Enzyme-linked immunosorbent assay quantitation of sTREM-1. Concentrations of sTREM-1 in the supernatants of monocyte cultures were determined using a commercially available ELISA kit (DuoSet; R&D Systems, Minneapolis, MN), following the manufacturer’s instructions (lower limit of detection: 15 pg ml–1).
Statistical analysis. The number of experiments analyzed is indicated in each figure legend. Data were collected from a minimum of three experiments and expressed as mean±s.d., except in Figure 10. The statistical significance was calculated using an unpaired t-test. Differences were considered significant at P-values ≤0.05 using Prism 6.0 software (GraphPad, La Jolla, CA).
References
Dasenbrook, E.C. Update on methicillin-resistant Staphylococcus aureus in cystic fibrosis. Curr. Opin. Pulm. Med. 17, 437–441 (2011).
Goss, C.H. & Muhlebach, M.S. Staphylococcus aureus and MRSA in cystic fibrosis. J. Cyst. Fibros. 10, 298–306 (2011).
Molina, A. et al. High prevalence in cystic fibrosis patients of multiresistant hospital-acquired methicillin-resistant Staphylococcus aureus ST228-SCCmecI capable of biofilm formation. J. Antimicrob. Chemother. 62, 961–967 (2008).
Stone, A. & Saiman, L. Update on the epidemiology and management of Staphylococcus aureus, including methicillin-resistant Staphylococcus aureus, in patients with cystic fibrosis. Curr. Opin. Pulm. Med. 13, 515–521 (2007).
Lo, D.K., Hurley, M.N., Muhlebach, M.S. & Smyth, A.R. Interventions for the eradication of methicillin-resistant Staphylococcus aureus (MRSA) in people with cystic fibrosis. Cochrane Database Syst. Rev. 2, CD009650 (2013).
Cantón, R. et al. Antimicrobial therapy for pulmonary pathogenic colonisation and infection by Pseudomonas aeruginosa in cystic fibrosis patients. Clin. Microbiol. Infect. 11, 690–703 (2005).
Gill, S.R. et al. Potential associations between severity of infection and the presence of virulence-associated genes in clinical strains of Staphylococcus aureus. PLoS One 6, e18673 (2011).
John, G., Chillappagari, S., Rubin, B.K., Gruenert, D.C. & Henke, M.O. Reduced surface toll-like receptor-4 expression and absent interferon-γ-inducible protein-10 induction in cystic fibrosis airway cells. Exp. Lung Res. 37, 319–326 (2011).
del Fresno, C. et al. Monocytes from cystic fibrosis patients are locked in an LPS tolerance state: down-regulation of TREM-1 as putative underlying mechanism. PLoS One 3, e2667 (2008).
del Fresno, C. et al. Potent phagocytic activity with impaired antigen presentation identifying lipopolysaccharide-tolerant human monocytes: demonstration in isolated monocytes from cystic fibrosis patients. J. Immunol. 182, 6494–6507 (2009).
del Campo, R. et al. Translocated LPS might cause endotoxin tolerance in circulating monocytes of cystic fibrosis patients. PLoS One 6, e29577 (2011).
Biswas, S.K. & Lopez-Collazo, E. Endotoxin tolerance: new mechanisms, molecules and clinical significance. Trends Immunol. 30, 475–487 (2009).
Arnalich, F. et al. Plasma levels of mitochondrial and nuclear DNA in patients with massive pulmonary embolism in the emergency department: a prospective cohort study. Crit. Care 17, R90 (2013).
Escoll, P. et al. Rapid up-regulation of IRAK-M expression following a second endotoxin challenge in human monocytes and in monocytes isolated from septic patients. Biochem. Biophys. Res. Commun. 311, 465–472 (2003).
del Fresno, C. et al. Inflammatory responses associated with acute coronary syndrome up-regulate IRAK-M and induce endotoxin tolerance in circulating monocytes. J. Endotoxin Res. 13, 39–52 (2007).
López-Collazo, E. & del Fresno, C. Pathophysiology of endotoxin tolerance: mechanisms and clinical consequences. Crit. Care 17, 242 (2013).
Máiz, L., Cantón, R., Mir, N., Baquero, F. & Escobar, H. Aerosolized vancomycin for the treatment of methicillin-resistant Staphylococcus aureus infection in cystic fibrosis. Pediatr. Pulmonol. 26, 287–289 (1998).
Ohniwa, R.L., Kitabayashi, K. & Morikawa, K. Alternative cardiolipin synthase Cls1 compensates for stalled Cls2 function in Staphylococcus aureus under conditions of acute acid stress. FEMS Microbiol. Lett. 338, 141–146 (2013).
Voyich, J.M. et al. The SaeR/S gene regulatory system is essential for innate immune evasion by Staphylococcus aureus. J. Infect. Dis. 199, 1698–1706 (2009).
Hiron, A. et al. A nickel ABC-transporter of Staphylococcus aureus is involved in urinary tract infection. Mol. Microbiol. 77, 1246–1260 (2010).
Gómez-Piña, V. et al. Metalloproteinases shed TREM-1 ectodomain from lipopolysaccharide-stimulated human monocytes. J. Immunol. 179, 4065–4073 (2007).
Deurenberg, R.H. & Stobberingh, E.E. The molecular evolution of hospital- and community-associated methicillin-resistant Staphylococcus aureus. Curr. Mol. Med. 9, 100–115 (2009).
García-Castillo, M. et al. Emergence of a mutL mutation causing multilocus sequence typing-pulsed-field gel electrophoresis discrepancy among Pseudomonas aeruginosa isolates from a cystic fibrosis patient. J. Clin. Microbiol. 50, 1777–1778 (2012).
Ruhen, R.W. Tetracycline dependence in a strain of Staphylococcus epidermidis. Pathology 8, 105–108 (1976).
Hoboth, C. et al. Dynamics of adaptive microevolution of hypermutable Pseudomonas aeruginosa during chronic pulmonary infection in patients with cystic fibrosis. J. Infect. Dis. 200, 118–130 (2009).
Goerke, C. & Wolz, C. Adaptation of Staphylococcus aureus to the cystic fibrosis lung. Int. J. Med. Microbiol. 300, 520–525 (2010).
Hirschhausen, N. et al. Extended Staphylococcus aureus persistence in cystic fibrosis is associated with bacterial adaptation. Int. J. Med. Microbiol. 303, 685–692 (2013).
Becker, S., Frankel, M.B., Schneewind, O. & Missiakas, D. Release of protein A from the cell wall of Staphylococcus aureus. Proc. Natl. Acad, Sci. USA 111, 1574–1579 (2014).
Hernández-Jiménez, E. et al. Biofilm vs. planktonic bacterial mode of growth: which do human macrophages prefer? Biochem. Biophys. Res. Commun. 441, 947–952 (2013).
Waite, G.N., Waite, L.R., Hughes, E.F. & Balcavage, W.X. Biophotonic hydrogen peroxide production by antibodies, T cells, and T-cell membranes. Biochem. Biophys. Res. Commun. 338, 1110–1117 (2005).
Oliveira, D.C. & de Lencastre, H. Multiplex PCR strategy for rapid identification of structural types and variants of the mec element in methicillin-resistant Staphylococcus aureus. Antimicrob. Agents Chemother. 46, 2155–2161 (2002).
Strommenger, B. et al. Assignment of Staphylococcus isolates to groups by spa typing, SmaI macrorestriction analysis, and multilocus sequence typing. J. Clin. Microbiol. 44, 2533–2540 (2006).
Harmsen, D. et al. Typing of methicillin-resistant Staphylococcus aureus in a university hospital setting by using novel software for spa repeat determination and database management. J. Clin. Microbiol. 41, 5442–5448 (2003).
Mellmann, A. et al. Based upon repeat pattern (BURP): an algorithm to characterize the long-term evolution of Staphylococcus aureus populations based on spa polymorphisms. BMC Microbiol. 7, 98 (2007).
Murchan, S. et al. Harmonization of pulsed-field gel electrophoresis protocols for epidemiological typing of strains of methicillin-resistant Staphylococcus aureus: a single approach developed by consensus in 10 European laboratories and its application for tracing the spread of related strains. J. Clin. Microbiol. 41, 1574–1585 (2003).
Enright, M.C., Day, N.P., Davies, C.E., Peacock, S.J. & Spratt, B.G. Multilocus sequence typing for characterization of methicillin-resistant and methicillin-susceptible clones of Staphylococcus aureus. J. Clin. Microbiol. 38, 1008–1015 (2000).
de las Heras, B. et al. Kaurane diterpenes protect against apoptosis and inhibition of phagocytosis in activated macrophages. Br. J. Pharmacol. 152, 249–255 (2007).
Pinheiro da Silva, F. et al. CD16 promotes Escherichia coli sepsis through an FcR gamma inhibitory pathway that prevents phagocytosis and facilitates inflammation. Nat. Med 13, 1368–1374 (2007).
Acknowledgements
We thank the participants (the patient and healthy volunteers) for their invaluable contributions, as well as Mercedes Rodríguez-Baños, Aurora Muñoz, and Victor Toledano for their technical assistance. We are also grateful to Auxiliadora Molina for the early work with microbiological cultures of this patient, as well as to ServingMed.com for editing the manuscript. This work was supported by grants from the Instituto Carlos III-FIS (Ministry of Economy and Competitiveness) to R.C., R.d.C., and E.L.-C. This study was supported by the Ministry of Economy and Competitiveness, Instituto de Salud Carlos III and cofinanced by the European Development Regional Fund “A way to achieve Europe” (ERDF) and the Spanish Network for Research in Infectious Diseases (RD12/0015/0004).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declared no conflict of interest.
Additional information
SUPPLEMENTARY MATERIAL is linked to the online version of the paper
Supplementary information
Rights and permissions
About this article
Cite this article
López-Collazo, E., Jurado, T., de Dios Caballero, J. et al. In vivo attenuation and genetic evolution of a ST247-SCCmecI MRSA clone after 13 years of pathogenic bronchopulmonary colonization in a patient with cystic fibrosis: implications of the innate immune response. Mucosal Immunol 8, 362–371 (2015). https://doi.org/10.1038/mi.2014.73
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mi.2014.73
This article is cited by
-
Identification of proteins regulated by acid adaptation related two component system HPK1/RR1 in Lactobacillus delbrueckii subsp. bulgaricus
Archives of Microbiology (2018)
-
Response to “In vivo attenuation and genetic evolution of a ST247-SCCmecI MRSA clone after 13 years of pathogenic bronchopulmonary colonization in a patient with cystic fibrosis: implications of the innate immune response”
Mucosal Immunology (2015)