Abstract
Intracellular parasites reprogram host functions for their survival and reproduction. The extent and relevance of parasite-mediated host responses in vivo remains poorly studied, however. We utilized Eimeria falciformis, a parasite infecting the mouse intestinal epithelium, to identify and validate host determinants of parasite infection. Most prominent mouse genes induced during the onset of asexual and sexual growth of parasite comprise interferon γ (IFNγ)-regulated factors, e.g., immunity-related GTPases (IRGA6/B6/D/M2/M3), guanylate-binding proteins (GBP2/3/5/6/8), chemokines (CxCL9-11), and several enzymes of the kynurenine pathway including indoleamine 2,3-dioxygenase 1 (IDO1). These results indicated a multifarious innate defense (tryptophan catabolism, IRG, GBP, and chemokine signaling), and a consequential adaptive immune response (chemokine-cytokine signaling and lymphocyte recruitment). The inflammation- and immunity-associated transcripts were increased during the course of infection, following influx of B cells, T cells, and macrophages to the parasitized caecum tissue. Consistently, parasite growth was enhanced in animals inhibited for CxCr3, a major receptor for CxCL9-11 present on immune cells. Interestingly, despite a prominent induction, mouse IRGB6 failed to bind and disrupt the parasitophorous vacuole, implying an immune evasion by E. falciformis. Furthermore, oocyst output was impaired in IFNγ-R−/− and IDO1−/− mice, both of which suggest a subversion of IFNγ signaling by the parasite to promote its growth.
Similar content being viewed by others
INTRODUCTION
The protozoan phylum Apicomplexa consists of about 5,000 parasite species of medical and veterinary importance, nearly all of which are obligate intracellular pathogens of diverse host cell types.1 The most prominent genera include Toxoplasma, Plasmodium, Eimeria, Cryptosporidium, and Theileria. A successful parasite infection necessitates a reprogramming of host-cell processes, such as cytoskeleton remodeling, metabolism, anti-microbial defense, inflammation, apoptosis, and cell cycle. Such changes are often preceded by an adaptive and subversive modulation of the host transcriptome in response to infection. DNA microarrays have proven excellent tools to examine gene expression on a genomic scale.2 Several studies employing this method have shaped our knowledge of host manipulation by apicomplexan parasites.3, 4, 5, 6, 7 Most of the research thus far has been performed using parasite cultures, however.3, 6, 8 There are few reports on ex vivo transcriptomics, but such work has used whole organs, tissues, or cells not serving as host to the parasite.5, 7, 9, 10 Others have made use of a non-model host organism, namely chickens.11, 12, 13 Therefore, identification of host-cell pathways and proteins modulated by infection in vivo and their physiological significance for parasite development remain very much underappreciated.
The outcome of an in vivo infection by an intracellular parasite depends on a sequence of events, such as pathogen–host interactions within the parasitized cell, cell-to-cell signaling, and the ensuing immune response. Infection-mediated signaling initiates the production of several chemokines and cytokines, which leads to a coordinated immune response against the pathogen. Interferon γ (IFNγ) is a key protective cytokine secreted by diverse cell types, which overarches innate and adaptive immune responses. For example, IFNγ stimulates the expression of IRG and GBP, defending against various microorganisms including parasites.14, 15, 16, 17, 18, 19, 20, 21 Likewise, IFNγ induces degradation of tryptophan via the kynurenine pathway to inhibit the growth of auxotrophic pathogens.22, 23, 24, 25 Further, IFNγ activates the transcription of various genes via JAK-STAT signaling including, but not limited to, the production of chemokines, cytokines, and reactive oxygen/nitrogen species.26 Consequently, the production of IFNγ is considered as pivotal for triggering effector mechanisms against Toxoplasma gondii,27, 28 Cryptosporidium,29, 30 Plasmodium,31, 32 and Eimeria33, 34, 35 infections.
The above-mentioned findings advocate a defensive role of IFNγ, which is largely in the favor of the host. It is not surprising, however, that apicomplexan parasites have instrumented multiple strategies to co-opt, exploit, or even subvert host signaling for their own benefits in the process of coevolution with their respective hosts. For instance, a number of different laboratories have demonstrated the ability of certain T. gondii strains to subvert and inactivate the IFNγ-inducible IRG and GBP system in the mouse.20, 36, 37, 38, 39 We have recently shown that E. falciformis co-opts the expression of a rate-limiting enzyme (IDO1) of tryptophan catabolism in the mouse.40 Notably, the use of a rodent host has been indispensable to examine the two facets of parasite–host interplay. In particular, E. falciformis, a host, tissue, and cell-specific monoxenous parasite of the murine caecum epithelial cells has proven valuable to explore immune and metabolic responses in the parasitized tissue.40, 41, 42 This study aimed at identifying and validating host determinants of the parasite infection at various cellular levels using the Eimeria-mouse mode.
RESULTS
Eimeria induces a significant modulation of mouse genes during early and late infection
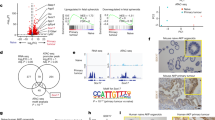
To perform an ex vivo gene expression analysis, we first isolated parasitized caecum epithelial cells from mice infected with E. falciformis at two distinct time points of 24 and 144 h, which coincide with the onset of asexual and sexual development, respectively. An assessment of the sample purity by flow cytometry showed a substantial proportion of epithelial cells (UEA+) accounting for 80–90% of the total cell number (Supplementary Figure S1A online). Immune cells (CD45+) comprised a minor 10–20% fraction. Similar results were achieved by qPCR, where EPCAM transcript appeared about five cycles earlier than CD45, which indicated a much higher abundance of epithelial cells over immune cells (Supplementary Figure S1B). Total RNA was isolated and subjected to expression analyses using whole-genome mouse microarray chips (Supplementary Figure S1C). Three independent infections with different oocyst batches were performed to obtain a set of most reliable genes modulated during in vivo infection. Normalized expression levels and expression differences for individual probes were consistent over a dynamic range in the three arrays as apparent by their symmetrical distribution along the base line expression (not shown). Heat maps of the transcriptomes confirmed reproducible differential expression of the majority of genes in all parasitized samples (Figure 1a). They also indicated a predominant upregulation over downregulation of host genes, particularly during late infection.
A notable number of the mouse genes are modulated in response to early and late infection with Eimeria falciformis. (a) Heat maps illustrating fold changes of mouse genes induced (red gradient) or repressed (green gradient) following the parasite infection in three biologically independent assays. Genes are ordered based on their hierarchical clustering (dendrogram not shown); columns as shown are not clustered. Significance of the observed difference over the three biological replicates for early (24 h p.i.) and late (144 h p.i.) time points are given in the right panel. Total RNA isolated from Eimeria-infected or uninfected mice was subjected to gene expression analyses. (b) Venn diagram depicting the number of aggregate consensus mouse transcripts from the three assays shown in (a). Significance of regulation was determined by two independent statistical analyses (see Methods).
Only genes modulated ≥1.5-fold and qualifying for significance in two different procedures were considered for downstream analyses. An aggregate consensus of 149 and 1,932 transcripts was regulated during early and late infection (Figure 1b). Further analyses revealed distinct modulation of mouse genes during the two infection phases. A set of 17 and 1,062 mouse genes was upregulated exclusively at 24 and 144 h post infection, respectively (Figure 1b). The numbers of distinctly downregulated transcripts were also equally noteworthy (9 vs. 747 genes in early and late infection). In total, 123 genes were differentially regulated during the entire infection period, of which 117 were induced and 6 were repressed. As apparent, the host response was more prominent 144 h post infection and the ratio of induced vs. repressed genes changed from 9 to 1.6 from early to late infection. Taken together, these results demonstrate that a notable fraction of the mouse genome (up to 10%) is modulated in response to intestinal infection with an intracellular pathogen.
IFNγ signaling is the most prominently induced pathway during Eimeria infection
Ingenuity Pathway Analyses (Ingenuity Systems, Redwood City, CA) identified the major processes significantly altered during the parasite infection, which included cellular development, growth, proliferation, movement, maintenance, morphology, death, and signaling (Supplementary Figure S2A). The gene regulation was more prominent 144 h post infection. Likewise, we also noted an appreciably high transcriptional response associated with metabolism (Supplementary Figure S2B) and immunity (see below, Figure 5c) during advanced infection. The major metabolic processes affected belong to molecular transport, as well as lipid, amino acid, vitamin, and mineral metabolism. It is noticeable that most gene categories, albeit amplified from the early to late infection, were conserved throughout infection. Importantly, a majority of the genes highly induced in response to parasite infection are regulated by IFNγ (Table 1; Supplementary Figures S3 and S4).
IFNγ signaling was apparent within 24 h of infection, and nearly all genes in the network showed an induction (Supplementary Figure S3). The cytokine-regulated network was more notably induced during late infection, and a few genes also exhibited a downregulation (Supplementary Figure S4). Table 1 summarizes the most upregulated genes after 24 and 144 h of infection. Interestingly, 17 of the 20 most upregulated genes early during infection are regulated by IFNγ. The number is equally impressive at the later time point (15 of the 20 genes). Some of these genes are IRGs (IRGA6, B6, D, M2, M3), GBPs (GBP2, 3, 5, 6, 8), chemokines (CxCL9, 10, CCL4), and IDO1. In contrast, only one IFNγ-regulated gene (FHL1) was downregulated (Table 2). Lipid metabolism and molecular transport categories were particularly enriched among the most repressed genes (Table 2; Supplementary Figure S2B). In brief, the above data indicated a host response rising in intensity during the course of infection, and suggested a key role of IFNγ-mediated signaling in Eimeria-infected mice.
Eimeria development is impaired in IFNγ-R−/− and IDO1−/− mice
IDO1 is the first and rate-limiting enzyme of tryptophan catabolism and it is potently induced by IFNγ.43, 44 We have shown that an induced expression of mouse IDO1 promotes the development of E. falciformis.40 To examine whether IFNγ signaling is required for induction of IDO1 in Eimeria-infected animals, we performed immunohistochemical staining using wild-type and IFNγ-R−/− mice (Figure 2a). The parental mice showed a marked IDO1 expression in the epithelial cells of the caecum of parasitized host, whereas no staining was detectable in the uninfected controls. The protein expression was confined to the epithelial cells serving as host cells for the parasite. IDO1 staining was completely absent in infected or naïve transgenic animals lacking their IFNγ receptor. Next, we tested the parasite’s growth in IFNγ-R−/− and IDO1−/− mice. The IFNγ-R-ablated animals shed a reduced amount of oocysts throughout the patent period from day 7 to 14, when compared with the parental mice (Figure 2b). Total oocyst output was decreased by 40% in mice disrupted for IFNγ-R signaling, which correlated well with the impaired development in IDO1−/− animals (Figure 2c). In brief, these results show that mouse IDO1 is induced by Eimeria infection in an IFNγ-dependent manner, and that the absence of IDO1 or of IFNγ signaling impairs parasite growth in vivo.
Mouse interferon γ (IFNγ) signaling is required for induction of indoleamine 2,3-dioxygenase 1 (IDO1) and an efficient parasite development. (a) Immunohistochemical staining of IDO1 in the caecum of parental and IFNγ-R−/− mice (naïve or d6 post infection). A parasite-induced IDO1 expression is present in epithelial cells (ECs) of wild type (WT) but not in the IFNγ-R−/− and naïve animals. Lumen (LU), lamina propria (LP), and submucosa (SM) are not stained. (b-c) Oocyst kinetics and total yield representing the parasite development in wild-type, IFNγ-R−/−, and IDO1−/− mice. The mice were infected with 50 oocysts and monitored for the oocyst output on the indicated days. The bars show the means±s.e. from two independent experiments with five mice per group (n=10 animals; *P≤0.05; **P≤0.01). The two-tailed non-parametric Mann–Whitney test was used for determination of statistical significance.
Eimeria induces tryptophan catabolism in the parasitized mouse
The aforesaid results motivated us to examine the expression of downstream enzymes of tryptophan catabolism in Eimeria-infected mice. To this end, we isolated parasitized samples throughout the course of infection (d1–d7), and quantified the transcript expression of all major enzyme isoforms (Figure 3a). Interestingly, the second isoform of IDO (IDO2) was upregulated by about 35-fold on day 1, and sustained its induced expression until the end of infection on day 7 (Figure 3a). Likewise, transcripts of other pathway enzymes, e.g., kynurenine aminotransferase (KAT), kynureninase (KU), kynurenine 3-hydroxylase (K3H), glutamate oxaloacetate transaminase (GOT), and 3-hydroxyanthranilate dioxygenase (HAD) were upregulated within the first day of infection and remained evidently induced throughout the parasite infection. Some of these enzymes (IDO2, KU, K3H, and GOT) were also induced in microarray studies (Table 3). These results are indicative of an enhanced tryptophan catabolism in the parasitized animals.
Several enzymes of tryptophan catabolism are upregulated throughout the life cycle of Eimeria. (a) qPCR data showing modulation of transcripts associated with tryptophan catabolism in the mouse infected with Eimeria for the indicated periods. The fold induction was normalized to Mm18S rRNA. The bars show the means±s.e. of two assays. ND, not detectable. (b) Parasite development in the mice inhibited for their IDO1/2 (1-MT), kynurenine 3-hydroxylase (K3H) (Ro61-8048), and kynurenine aminotransferase (KAT) (S-ESBA) functions. The panel b has been adapted from previous work.40 The animals were treated with the inhibitors in the drinking water (1-MT) or by oral tube (Ro61-8048, S-ESBA) and examined for the oocyst shedding from d6 to d14. Statistical significance was evaluated by the two-tailed non-parametric Mann–Whitney test (***P≤0.001). IDO, indoleamine 2,3-dioxygenase.
To test the importance of the kynurenine pathway for Eimeria growth, we used a variety of inhibitors (1-MT, Ro61-8048, and S-ESBA) with proven efficacy against the enzymes of the kynurenine pathway (IDO, K3H, and KAT) in the mouse.45, 46, 47 Interestingly, we found a significant impairment in the oocyst output of animals receiving 1-MT, Ro61-8048, and S-ESBA, which inhibit IDO, K3H, and KAT activity, respectively (Figure 3b).40 In conclusion, the data show that tryptophan catabolism is required for an efficient development of E. falciformis in the mouse.
Eimeria evades the recruitment of host IRGB6 onto its vacuole
The IFNγ-regulated signaling network identified a set of IRGs and GBPs (Table 1; Supplementary Figures S3 and S4). IRG proteins are induced by IFNγ and constitute an important part of murine innate defense mechanisms against intracellular pathogens including T. gondii.14, 15, 16, 17, 18, 19, 20, 21, 48 Expression levels of several members of the IRG family, such as IRGM2-M3, IRGA6, IRGB6, and IRGD, were highly induced in Eimeria-infected animals. The upregulated GBP family included GBP2, GBP3, GBP5, GBP6, and GBP8. To deduce the impact of these proteins during Eimeria infection, we first quantified the expression of selected members by qPCR and microarray methods (Table 3). Both GBP2 and GBP3 were induced upon 24 h as well as 144 h post infection. Likewise, IRGM3 and IRGB6 were substantially upregulated at both time points of infection.
It has been shown that members of the IRG family including IRGB6 are recruited onto the vacuole of type II strain of T. gondii, and participate in subsequent disruption of the vacuolar membrane.14 In contrast, type I strain of T. gondii can phosphorylate and inactivate the host IRG proteins resulting in a failure to recognize the vacuole and increased parasite survival.38 To deduce whether IRGB6 has a role in E. falciformis infection, we infected young adult mouse colon (YAMC) cells with Eimeria, and immunostained host cell and the parasite with anti-IRGB6 and anti-actin antibodies, respectively (Figure 4a). In an identical set-up, we included type I and II strains of T. gondii for comparative analyses. As expected, the vacuole of type II strain was coated with host IRGB6 protein, whereas type I strain successfully prevented coating of its vacuolar membrane. Similarly to type I strain, the Eimeria vacuole was not stained by mouse IRGB6. A quantification of the IRGB6-stained vacuoles showed coating of about 45% vacuoles for type II strain, 6% for type I strain, and <2% for E. falciformis (Figure 4b) within 1–2 h of infection. In summary, these data demonstrate that despite its conspicuous induction IRGB6 fails to recognize the vacuolar membrane of E. falciformis similar to type I strain of T. gondii.
Immunity-related GTPaseB6 (IRGB6) fails to recognize the Eimeria vacuole despite a notable induction. (a) Immunostaining of young adult mouse colon (YAMC) cells parasitized with sporozoites of E. falciformis or tachyzoites of T. gondii (multiplicity of infection, 0.5; 1–2 h post infection) using anti-TgActin/Alexa488 and anti-MmIRGB6/Alexa594 antibodies. Type I and II strains of T. gondii were included as controls for recruitment of host IRGB6 protein onto the parasitophorous vacuole. (b) Quantification of IRGB6 loading onto the vacuoles of indicated parasites after 1 and 2 h of infection. Parasite vacuoles coated with IRGB6 staining in panel a were estimated, and plotted as the percentage of total infected cells. The error bars indicate the means±s.e. of three assays. The two-tailed t-test was used for determination of statistical significance (**P≤0.01; ***P≤0.001).
Inhibition of CxCr3 signaling in the mouse promotes the parasite development
The aforementioned results also showed several chemokines, in particular of the CxCL family, to be highly upregulated during Eimeria infection. These chemokines are secreted by intestinal epithelial cells, and are considered as critical to recruit immune cells. To assess their relative expression levels, we first validated our microarray data by qPCR method (Table 3). We observed an early upregulation of all tested chemokines, which was sustained until late infection. The transcripts of CxCL9, CxCL10, as well as CxCL11 were induced approximately to the same order of magnitude. It is known that CxCr3 receptor present primarily on immune cells serves as a ligand to these three chemokines.49 Hence, we inhibited this receptor with a known inhibitor (AMG487) in Eimeria-infected animals, and monitored the parasite development (Figure 5a). We observed a notably higher oocyst shedding from d7 until d10 by the drug-treated animals. Total oocyst yield was increased by about 31%, compared with the control group (Figure 5b).
Inhibition of the mouse CxCr3 increases Eimeria development. (a, b) Parasite development in the mice inhibited for CxCr3 function. The animals were infected with 50 oocysts, and AMG487 (CxCr3 inhibitor) was administered subcutaneously twice a day (5 mg kg−1 bodyweight) from d1 until the end of experiment. Parasite yield was scored from d6 to d14 from two independent assays (n=12 animals). (c) Ingenuity Pathway Analysis of the induced (smooth bar) and repressed (hatched bar) immunity-related processes during early (24 h; dark gray) and late (144 h; light gray) infection of mouse with Eimeria. WT, wild type.
Consistent with chemokine expression and CxCr3 signaling, the gene categories significantly enriched during the advanced-stage infection included inflammation, immune cell trafficking, cell-mediated immunity, antigen presentation, and antimicrobial response (Figure 5c; Table 1). To corroborate these data, we tested the recruitment of selected immune cells at the site of infection (Figure 6). As expected, we detected a basal staining for B cells (B220+), T cells (CD3+), and macrophages (F4/80+) in naïve animals (Figure 6a). Expression of a macrophage-specific marker, but not of B- and T-cell markers was modestly higher 24 h post infection. During the late infection, however, we observed a much greater influx of all cell types to the site of parasite infection (Figure 6a). An arbitrary quantification of histological samples confirmed a significant recruitment of the indicated cell types (Figure 6b). Collectively, these results demonstrate the requirement of chemokine signaling and cell-mediated immune response in limiting parasite infection in the mouse.
An elevated transcript expression correlates with an amplified immune response in the Eimeria-infected mice. (a–d) Immunohistochemical staining of B cells (B220+), macrophages (F4/80+), and T cells (CD3+) in the caecum of naïve and infected animals. EC, epithelial cells; HPF, high power field; IEL, intraepithelial lymphocyte; LP, lamina propria; LU, lumen; SM, submucosa. Bar graphs represent the mean±s.e. of two experiments each with four animals. Statistics was performed by the two-tailed non-parametric Mann–Whitney test (*P≤0.05).
DISCUSSION
Our work using the E. falciformis infection of the mouse identifies a retinue of host proteins that are modulated by the parasite. In particular, we reveal several host proteins and pathways regulated by IFNγ signaling during infection. Quite notably, some of the host responses promote parasite growth and others inhibit it (Figure 7). For example, chemokine signaling leading to lymphocyte recruitment is detrimental for the parasite’s life cycle, whereas the kynurenine pathway is beneficial. In addition, the parasite appears to evade the IRG-mediated innate defense of the host cells. These results support a model with divergent roles of IFNγ-regulated host proteins during infection, some of which defend the insult and others are co-opted or subverted by the parasite. Further, a variety of transgenic mice devoid of the key differentially regulated factors are available, which can be employed to examine many other determinants of parasite infection (Figure 7). To this end, the monoxenous life cycle of E. falciformis in a well-investigated host offers an excellent model for in vivo parasite–host interactions.
Proposed in vivo model of Eimeria-mouse interactions. The model is based on gene expression analyses, experimental support for selected host proteins and proposed hypotheses. Only selected host pathways influencing the parasite development are indicated. The color-coded (red or green) proteins, induced or repressed in our analyses and for which transgenic animals are available, are also shown. The plus (+) and minus (−) signs in circles denote a positive or negative impact of a given pathway on the parasite’s in vivo life cycle. Figure was produced using Servier Medical Art (www.servier.com).
IFNγ is pivotal to control diverse infections including those caused by protozoan parasites. Its absence typically increases host susceptibility, enhancing the pathogen burden.50, 51, 52 Our results suggest an IFNγ-regulated innate defense in Eimeria-infected animals, which is orchestrated at various cellular levels. Of note is the induction of the tryptophan catabolism, which is usually deemed as an effector mechanism of the innate host defense to deprive and kill pathogens auxotrophic for this essential amino acid.24, 53, 54 In E. falciformis infection, however, inhibition of tryptophan catabolism by independent means impairs parasite growth, suggesting a supportive role of IFNγ-mediated induction of amino-acid metabolism for the in vivo parasite cycle. Indeed, we have shown that a major by-product metabolite of the kynurenine pathway, xanthurenic acid, is required by the parasite for its efficient development in mice.40 Conversely, it has also been shown that in vitro treatment of parasite cultures with IFNγ reduces Eimeria growth,41 the mechanism of which is still unclear. It is quite conceivable that our in vivo results reflect a balance between the two opposing phenomena (Figure 7).
In parallel to the parasite-promoting function of IFNγ signaling, our results show the presence of other IFNγ-regulated innate defense mechanisms in the mouse, involving several members of the GBP and IRG family. These proteins confer resistance to a variety of organisms in a largely pathogen-specific manner. For instance, IRGM1 deficiency causes an increased susceptibility to the protozoan parasites including T. gondii,48 L. major,17 and T. cruzi,16 and to the bacteria, Listeria monocytogenes18 and Mycobacterium tuberculosis.19 Interestingly, IRGM3−/− animals were more vulnerable to T. gondii15 and L. major,17 but not to T. cruzi55 and L. monocytogenes.18 GBP members are also known for their antimicrobial effect on bacteria as well as on parasites.20, 56, 57 For example, GBP1−/− mice infected with L. monocytogenes, M. bovis or T. gondii show an increased microbial burden,39, 57 and the animals harboring a collective deletion of GBP1–3, 5, and 7 on chromosome 3 succumb to parasite infection.21 The mechanism of their action has been extensively characterized during T. gondii infection. Many IRG and GBP family members assemble at the vacuole membrane, subsequently killing the parasite.14, 20 The virulent (type I) strains of T. gondii can inactivate the IRG proteins by phosphorylation and prevent their accumulation onto parasite vacuole.38 Likewise, IRGB6 failed to coat the Eimeria vacuole, implying either a lack of vacuolar propensity to bind IRG, or its active subversion by the parasite. The use of the indicated IRG (and GBP) knockout mice (Figure 7) should allow further research on the precise roles of these protein families during Eimeria infection.
In addition to innate defense, we observed the onset of an adaptive immunity, principally during late infection. The major chemokines CxCL9-11, likely secreted by intestinal epithelial cells, were induced within 24 h of infection. IFNγ signaling regulates the expression of all these chemokines in various models, including in Eimeria-infected animals.41, 58, 59, 60 Their signaling through the CxCr3 receptor present on T and B lymphocytes, macrophages, and NK cells results in the recruitment of immune cells to the site of infection.61, 62 Indeed, we detected a significant increase in the numbers of lymphocytes and macrophages to the Eimeria-infected caeca, and inhibition of CxCr3 in parasitized animals increased the parasite burden. The data suggest an important function of epithelial chemokines in the inception of the adaptive immune response and control of parasite infection. Although requiring a validation in CxCr3−/− mice, these findings are consistent with a role of the indicated chemokines in orchestrating a cell-mediated immune response against other parasites, T. gondii and T. cruzi.58, 60
In summary, we show the reciprocal consequences of host IFNγ signaling on the life cycle of a tissue-specific intracellular pathogen, while providing a glimpse of parasite–host interplay occurring in vivo. We also highlight potential host proteins bridging the innate and adaptive arms of the mouse immune response, which can be investigated by using the available transgenic mice (Figure 7). Last but not least, the work illustrates the value of ex vivo transcriptomics and Eimeria-mouse model to characterize host determinants of parasite development.
METHODS
Chemical and biological reagents. IFNγ-R−/− (C57BL/6) and IDO1−/− (BALB/c) mice strains were donated by Uwe Klemm (Max-Planck Institute for Infection Biology, Berlin) and Muriel Moser (Université Libre de Bruxelles, Belgium), respectively. The parental control (C57BL/6, BALB/c) and NMRI mice strains were purchased from the Charles River Laboratories (Sulzfeld, Germany). All animal experiments were executed according to the animal protection laws as directed by the overseeing authority, Landesamt fuer Gesundheit und Soziales (Berlin, Germany). The oocysts of E. falciformis were procured from Bayer (Leverkusen, Germany). The type I (RH) and type II (Pru) strains of T. gondii were donations from Carsten Lüder (Georg-August University, Germany) and Louis Weiss (Albert Einstein College of Medicine, USA). Anti-TgActin, anti-TgSag1, and anti-MmIRGB6 antibodies were donated by Dominique Soldati-Favre (University of Geneva, Switzerland), Jean-François Dubremetz (University of Montpellier, France), and Jonathan Howard (University of Cologne, Germany), respectively. YAMC cells were provided by Robert Whitehead (Vanderbilt University, USA63).
Propagation of E. falciformis in the mouse. The life cycle of E. falciformis was maintained by continuous passage in NMRI mice. In brief, oocysts were stored in potassium dichromate at 4 °C up to 3 months. For purification, oocysts were washed with water, sterilized, and floated with NaOCl.64 Oocysts were quantified by using a McMaster counting chamber (Frick, Switzerland). For gene expression analyses, 8- to 12-week-old mice were orally infected with 1,500 oocysts suspended in 100 μl phosphate-buffered saline (PBS). The IFNγ-R and IDO1 knockout animals and their parental controls were infected with 50 oocysts and assessed for their oocyst outputs, as described previously.40
Isolation of mouse caecum epithelium. Infected and uninfected caeca were washed in Ca++ and Mg++-free Hank's balanced salt solution, longitudinally dissected into sections (5 mm) in culture dishes, collected in polypropylene tubes, and incubated with 1 mM dithiothreitol in Hank's balanced salt solution on a horizontal rotation shaker (30 min, 37 °C). Supernatants were discarded and tissues were suspended in 1 mM EDTA/Hank's balanced salt solution. Samples were incubated for 30 min at 37 °C, and shaken rigorously every 10 min. Subsequent supernatants were passed through a cell strainer (70 μm) into fresh polypropylene tubes and centrifuged at 1,500 r.p.m. for 5 min. Pellets were resuspended in 1 ml of TRIzol reagent for gene expression analyses and stored at −80 °C until further use. To test the sample purity, cells were counted using a Casy counter (Innovatis, Bielefeld, Germany), and suspended in 0.2% BSA/5 mM EDTA/PBS for flow cytometry. Samples (106 cells per tube) were washed with 0.2% BSA/5 mM EDTA/PBS (1,300 r.p.m., 5 min, 4 °C), treated with anti-mouse FcγR-blocker (20 μg ml−1), and stained with fluorescent antibodies for 30 min on ice. Epithelial and immune cells were labeled with UEA1-FITC (1:80) and CD45-ef450 (1:200) antibodies, respectively. Subsequently, cells were fixed in 0.5% paraformaldehyde/PBS and subjected to flow cytometry (FlowJo Software, Tree Star, Ashland, OR).
RNA isolation, qPCR, and microarray. Total RNA isolated from infected or uninfected mouse caecum cells using a PureLink RNA kit (Life Technologies, Darmstadt, Germany) was subjected to gene expression analyses. Quantitative real-time PCR was done using SuperScript III Platinum SYBR Green with ROX (One-Step Kit; Life Technologies). PCR primers were designed based on the NCBI accession numbers (Table 3). The fold induction in qPCR studies was normalized to 18S rRNA. Gene expression was analyzed by dual-color dye-swap hybridizations using mouse whole-genome microarrays (Agilent Technologies, Boeblingen, Germany). All sample-processing steps were performed according to the manufacturer’s instructions. Briefly, total RNA was reverse-transcribed, amplified, and labeled using an oligo-dT-T7 promoter-primer and Cyanine 3-CTP or Cyanine 5-CTP (QuickAmp kit, Agilent Technologies). After precipitation, purification, and quantification, 1.25 μg of each labeled cRNA was fragmented and hybridized to microarrays (AMADID 014868) overnight followed by washing steps. The microarray images were recorded using a laser microarray scanner (G2565CA) at 5 μm resolution (Agilent Technologies).
Analyses of microarray data. Raw microarray image data were analyzed with the Agilent image analysis and feature extraction software using default settings (G2567AA). Dye ratios were calculated using the most conservative estimate between the universal and propagated error. The extracted MAGE-ML files were analyzed with the Rosetta Resolver Biosoftware (Boston, MA). Only anti-correlated genes of dye-reversal hybridizations with 1.5-fold cutoffs and an error-weighted P-value of <0.01 were considered as differentially regulated.
Alternatively, extracted features were subjected to R/bioconductor analyses (www.r-project.org; www.bioconductor.org). The package limma was used to subtract background intensities and resulting negative values were set to half the minimum of the positive corrected intensities.65 The “loess” and “Aquantile” methods were used to normalize within and between arrays, respectively. Probes were filtered according to Agilent quality flags using the package Agi4 × 44PreProcess and only non-control probes over the background level and with well-saturated signal were retained. Intensity values for genes were calculated as average of their probes. Symmetry of most gene probes around the zero horizontal line in MA intensity plots (log ratio/log average) indicated appropriate normalization over a large dynamic range of signal intensity (not shown). Statistical testing was performed with limma (dye-swapped samples as technical replicates), fitting a mixed effects linear model. Differential expression was estimated between infected and respective uninfected samples using empirical Bayes moderated t-statistics. A fold change of ≥1.5 and a P-value of <0.05 were used as thresholds for statistical significance. Annotations were obtained using the R-package mgug4122a.db (www.bioconductor.org/packages/2.13/data/annotation/html/mgug4122a.db.html).
The above two independent microarray analyses yielded consistent results for 99% of genes at 24 h post infection, and 85% of the genes at 144 h. All genes discussed in this work were prominently regulated in both approaches. In-depth analysis of low to modestly regulated genes was not undertaken in this work, which warrants further investigation using a higher number of biological replicates. The expression data were used as input for Ingenuity Pathway Analysis (http://www.ingenuity.com) and for interferome suite (http://www.interferome.org) to deduce the biological relevance of the genes modulated during infection. Microarray data have been deposited in the Gene Expression Omnibus (GEO accession number GSE49948).
Parasite and host cell culture. The two strains of T. gondii were maintained by serial passage in human foreskin fibroblasts as described elsewhere.66 To isolate E. falciformis sporozoites, purified oocysts were digested with 0.4% pepsin (pH 3, 37 °C, 1 h) and washed with PBS. The oocyst pellet was mixed with glass beads (0.5 mm) in a 1:1 ratio and briefly vortexed to release sporocysts. For excystation of sporozoites, sporocyst samples were incubated with (0.25%) trypsin and sodium tauroglycocholate (0.04%; MP Biomedicals, Eschwege, Germany) in DMEM supplemented with glutamine (20 mM), streptomycin (100 μg ml−1), and penicillin (100 U ml−1) at 37 °C for up to 2 h. Sporozoites were column purified by DE52 anion exchange chromatography.67 YAMC cells were cultured in RPMI medium containing FCS (5%), insulin (1 μg ml−1), α-thioglycerol (10 μM), hydrocortisone (1 μM), and IFNγ (5 U ml−1) in a humidified incubator (33 °C, 10% CO2). Cells were harvested by trypsin/EDTA treatment for propagation.
IRG loading onto parasite vacuole. YAMC cells, grown on glass coverslips, were stimulated with 1,000 U ml−1 IFNγ at least 24 h before infection with Eimeria sporozoites or Toxoplasma tachyzoites (multiplicity of infection, 0.5). Infection was performed in medium with 1% FCS for 1–2 h followed by immunostaining. Cells were fixed in 4% paraformaldehyde (10 min) followed by neutralization with 0.1 M glycine/PBS (5 min, room temperature). Permeabilization was achieved by 0.2% Triton X-100/PBS (20 min). Samples were blocked with 2% BSA in Triton X-100/PBS, and labeled with the primary antibodies (rabbit anti-MmIRGB6 and mouse anti-TgActin) suspended in the blocking solution (1 h, room temperature). They were washed three times in 0.2% Triton X-100/PBS, and treated with the secondary antibodies (Alexa488/594-conjugated IgG, 1:3,000, 45 min, room temperature). After washing, the coverslips were mounted in Fluoromount G/DAPI, and examined by fluorescence microscopy (Apotome, Carl-Zeiss, Germany).
A red/green invasion assay was performed in parallel to examine intracellular tachyzoites for each experiment. Briefly, cells were fixed and stained with anti-TgSag1 antibody before permeabilization. Incubation with rabbit anti-TgGap45 was carried out under permeabilized conditions (2% BSA/0.2% Triton X-100/PBS) for 1 h. Alexa488/594-conjugated secondary antibodies were used for fluorescent staining. Invasion rates of tachyzoites were determined by calculating the proportion of extracellular (Sag1-stained) to Gap45-stained parasites. Intracellular sporozoites of Eimeria were determined by differential interference contrast imaging. The amount of IRGB6-coated vacuoles was counted with respect to intracellular parasites.
Immunohistochemical analyses. The caeca of infected and uninfected mice were washed carefully and stored in 4% paraformaldehyde/PBS. Tissues were paraffin embedded and dissected into 2 μm sections. Samples were de-paraffinized in xylene, and re-hydrated in descending concentrations of ethanol followed by washing with water. Antigen retrieval was achieved by boiling the samples in citric buffer (pH 6.0) in a pressure cooker. Slides were rinsed with water and Tris-buffered saline. The primary rat anti-mouse antibody (MmIDO1-48, 1:200, Biolegend, Fell, Germany) was added for 30 min. Following three washes, the sections were incubated for 30 min with the secondary antibody (biotinylated rabbit anti-rat, 1:200, Dako, Hamburg, Germany). Streptavidin alkaline phosphatase (Dako) was applied onto the tissue sections for 30 min, and the reaction was visualized by incubating with Fast RED (Dako). Slides were rinsed with water and counterstained with hematoxylin. Images were taken using a LEICA DMIL microscope fitted with a DFC480 camera (Wetzlar, Germany), and processed with LEICA software (v2.8.1, Wetzlar, Germany).
Accession codes
References
Levine, N.D. The Protozoan Phylum Apicomplexa vols. 1 and 2 CRC Press: Boca Raton, FL, (1988).
Sheils, O, Finn, S & O'Leary, J. Nucleic acid microarrays: an overview. Curr. Diagn. Pathol. 9, 155–158 (2003).
Blader, I.J., Manger, I.D. & Boothroyd, J.C. Microarray analysis reveals previously unknown changes in Toxoplasma gondii-infected human cells. J. Biol. Chem. 276, 24223–24231 (2001).
Sexton, A.C. et al. Transcriptional profiling reveals suppressed erythropoiesis, up-regulated glycolysis, and interferon-associated responses in murine malaria. J. Infect. Dis. 189, 1245–1256 (2004).
Lovegrove, F.E., Gharib, S.A., Patel, S.N., Hawkes, C.A., Kain, K.C. & Liles, W.C. Expression microarray analysis implicates apoptosis and interferon-responsive mechanisms in susceptibility to experimental cerebral malaria. Am. J. Pathol. 171, 1894–1903 (2007).
Taubert, A, Wimmers, K, Ponsuksili, S, Jimenez, C.A., Zahner, H & Hermosilla, C. Microarray-based transcriptional profiling of Eimeria bovis-infected bovine endothelial host cells. Vet. Res. 41, 70 (2010).
Ellis, J, Sinclair, D, Morrison, D, Al-Qassab, S, Springett, K & Ivens, A. Microarray analyses of mouse responses to infection by Neospora caninum identifies disease associated cellular pathways in the host response. Mol. Biochem. Parasitol. 174, 117–127 (2010).
Albuquerque, S.S. et al. Host cell transcriptional profiling during malaria liver stage infection reveals a coordinated and sequential set of biological events. BMC Genomics 10, 270 (2009).
Miu, J, Hunt, N.H. & Ball, H.J. Predominance of interferon-related responses in the brain during murine malaria, as identified by microarray analysis. Infect. Immun. 76, 1812–1824 (2008).
Delahaye, N.F. et al. Gene expression analysis reveals early changes in several molecular pathways in cerebral malaria-susceptible mice versus cerebral malaria-resistant mice. BMC Genomics 8, 452 (2007).
Zhang, L. et al. Eimeria tenella: expression profiling of toll-like receptors and associated cytokines in the cecum of infected day-old and three-week old SPF chickens. Exp. Parasitol. 130, 442–448 (2012).
Min, W. et al. Profiling local gene expression changes associated with Eimeria maxima and Eimeria acervulina using cDNA microarray. Appl. Microbiol. Biotechnol. 62, 392–399 (2003).
Guo, A. et al. Transcriptome analysis in chicken cecal epithelia upon infection by Eimeria tenella in vivo. PLoS ONE 8, e64236 (2013).
Martens, S. et al. Disruption of Toxoplasma gondii parasitophorous vacuoles by the mouse p47-resistance GTPases. PLoS Pathog. 1, e24 (2005).
Taylor, G.A. et al. Pathogen-specific loss of host resistance in mice lacking the IFN-gamma-inducible gene IGTP. Proc. Natl. Acad. Sci. USA 97, 751–755 (2000).
Santiago, H.C. et al. Mice deficient in LRG-47 display enhanced susceptibility to Trypanosoma cruzi infection associated with defective hemopoiesis and intracellular control of parasite growth. J. Immunol. 175, 8165–8172 (2005).
Taylor, G.A., Feng, C.G. & Sher, A. p47 GTPases: regulators of immunity to intracellular pathogens. Nat. Rev. Immunol. 4, 100–109 (2004).
Collazo, C.M. et al. Inactivation of LRG-47 and IRG-47 reveals a family of interferon gamma-inducible genes with essential, pathogen-specific roles in resistance to infection. J. Exp. Med. 194, 181–188 (2001).
MacMicking, J.D., Taylor, G.A. & McKinney, J.D. Immune control of tuberculosis by IFN-gamma-inducible LRG-47. Science 302, 654–659 (2003).
Degrandi, D. et al. Extensive characterization of IFN-induced GTPases mGBP1 to mGBP10 involved in host defense. J. Immunol. 179, 7729–7740 (2007).
Yamamoto, M. et al. A cluster of interferon-gamma-inducible p65 GTPases plays a critical role in host defense against Toxoplasma gondii. Immunity 37, 302–313 (2012).
Adams, O, Besken, K, Oberdörfer, C, MacKenzie, C.R., Takikawa, O & Däubener, W. Role of indoleamine-2,3-dioxygenase in alpha/beta and gamma interferon-mediated antiviral effects against herpes simplex virus infections. J. Virol. 78, 2632–2636 (2004).
Bodaghi, B, Goureau, O, Zipeto, D, Laurent, L, Virelizier, J.L. & Michelson, S. Role of IFN-gamma-induced indoleamine 2,3 dioxygenase and inducible nitric oxide synthase in the replication of human cytomegalovirus in retinal pigment epithelial cells. J. Immunol. 162, 957–964 (1999).
Byrne, G.I., Lehmann, L.K. & Landry, G.J. Induction of tryptophan catabolism is the mechanism for gamma-interferon-mediated inhibition of intracellular Chlamydia psittaci replication in T24 cells. Infect. Immun. 53, 347–351 (1986).
Pfefferkorn, E.R., Eckel, M & Rebhun, S. Interferon-gamma suppresses the growth of Toxoplasma gondii in human fibroblasts through starvation for tryptophan. Mol. Biochem. Parasitol. 20, 215–224 (1986).
Hu, X & Ivashkiv, L.B. Cross-regulation of signaling pathways by interferon-gamma: implications for immune responses and autoimmune diseases. Immunity 31, 539–550 (2009).
Denkers, E.Y. & Gazzinelli, R.T. Regulation and function of T-cell-mediated immunity during Toxoplasma gondii infection. Clin. Microbiol. Rev. 11, 569–588 (1998).
Hou, B, Benson, A, Kuzmich, L, DeFranco, A.L. & Yarovinsky, F. Critical coordination of innate immune defense against Toxoplasma gondii by dendritic cells responding via their Toll-like receptors. Proc. Natl. Acad. Sci. USA 108, 278–283 (2011).
Ungar, B.L., Kao, T.C., Burris, J.A. & Finkelman, F.D. Cryptosporidium infection in an adult mouse model. Independent roles for IFN-gamma and CD4+ T lymphocytes in protective immunity. J. Immunol. 147, 1014–1022 (1991).
McDonald, V, Deer, R, Uni, S, Iseki, M & Bancroft, G.J. Immune responses to Cryptosporidium muris and Cryptosporidium parvum in adult immunocompetent or immunocompromised (nude and SCID) mice. Infect. Immun. 60, 3325–3331 (1992).
Doolan, D.L., Sedegah, M, Hedstrom, R.C., Hobart, P, Charoenvit, Y & Hoffman, S.L. Circumventing genetic restriction of protection against malaria with multigene DNA immunization: CD8+ cell-, interferon gamma-, and nitric oxide-dependent immunity. J. Exp. Med. 183, 1739–1746 (1996).
Good, M.F., Xu, H, Wykes, M & Engwerda, C.R. Development and regulation of cell-mediated immune responses to the blood stages of malaria: implications for vaccine research. Annu. Rev. Immunol. 23, 69–99 (2005).
Lillehoj, H.S. Role of T lymphocytes and cytokines in coccidiosis. Int. J. Parasitol. 28, 1071–1081 (1998).
Lillehoj, H.S. & Choi, K.D. Recombinant chicken interferon-gamma-mediated inhibition of Eimeria tenella development in vitro and reduction of oocyst production and body weight loss following Eimeria acervulina challenge infection. Avian Dis. 42, 307–314 (1998).
Rose, M.E., Smith, A.L. & Wakelin, D. Gamma interferon-mediated inhibition of Eimeria vermiformis growth in cultured fibroblasts and epithelial cells. Infect. Immun. 59, 580–586 (1991).
Khaminets, A. et al. Coordinated loading of IRG resistance GTPases on to the Toxoplasma gondii parasitophorous vacuole. Cell Microbiol. 12, 939–961 (2010).
Zhao, Y, Ferguson, D.J., Wilson, D.C., Howard, J.C., Sibley, L.D. & Yap, G.S. Virulent Toxoplasma gondii evade immunity-related GTPase-mediated parasite vacuole disruption within primed macrophages. J. Immunol. 182, 3775–3781 (2009).
Steinfeldt, T. et al. Phosphorylation of mouse immunity-related GTPase (IRG) resistance proteins is an evasion strategy for virulent Toxoplasma gondii. PLoS Biol. 8, e1000576 (2010).
Selleck, E.M. et al. Guanylate-binding protein 1 (Gbp1) contributes to cell-autonomous immunity against Toxoplasma gondii. PLoS Pathog. 9, e1003320 (2013).
Schmid, M, Lehmann, M.J., Lucius, R & Gupta, N. Apicomplexan parasite, Eimeria falciformis, co-opts host tryptophan catabolism for life cycle progression in mouse. J. Biol. Chem. 287, 20197–20207 (2012).
Stange, J. et al. IL-22 mediates host defense against an intestinal intracellular parasite in the absence of IFN-gamma at the cost of Th17-driven immunopathology. J. Immunol. 188, 2410–2418 (2012).
Pogonka, T. et al. CD8+ cells protect mice against reinfection with the intestinal parasite Eimeria falciformis. Microbes Infect 12, 218–226 (2010).
Munn, D.H., Shafizadeh, E, Attwood, J.T., Bondarev, I, Pashine, A & Mellor, A.L. Inhibition of T cell proliferation by macrophage tryptophan catabolism. J. Exp. Med. 189, 1363–1372 (1999).
Hwu, P, Du, M.X., Lapointe, R, Do, M, Taylor, M.W. & Young, H.A. Indoleamine 2,3-dioxygenase production by human dendritic cells results in the inhibition of T cell proliferation. J. Immunol. 164, 3596–3599 (2000).
Munn, D.H. et al. Prevention of allogeneic fetal rejection by tryptophan catabolism. Science 281, 1191–1193 (1998).
Pellicciari, R. et al. Modulators of the kynurenine pathway of tryptophan metabolism: synthesis and preliminary biological evaluation of (S)-4-(ethylsulfonyl)benzoylalanine, a potent and selective kynurenine aminotransferase II (KAT II) inhibitor. ChemMedChem 1, 528–531 (2006).
Röver, S, Cesura, A.M., Huguenin, P, Kettler, R & Szente, A. Synthesis and biochemical evaluation of N-(4-phenylthiazol-2-yl)benzenesulfonamides as high-affinity inhibitors of kynurenine 3-hydroxylase. J. Med. Chem. 40, 4378–4385 (1997).
Butcher, B.A. et al. p47 GTPases regulate Toxoplasma gondii survival in activated macrophages. Infect. Immun. 73, 3278–3286 (2005).
Clark-Lewis, I, Mattioli, I, Gong, J.H. & Loetscher, P. Structure-function relationship between the human chemokine receptor CXCR3 and its ligands. J. Biol. Chem. 278, 289–295 (2003).
Huang, S. et al. Immune response in mice that lack the interferon-gamma receptor. Science 259, 1742–1745 (1993).
Scharton-Kersten, T.M. et al. In the absence of endogenous IFN-gamma, mice develop unimpaired IL-12 responses to Toxoplasma gondii while failing to control acute infection. J. Immunol. 157, 4045–4054 (1996).
Su, Z & Stevenson, M.M. Central role of endogenous gamma interferon in protective immunity against blood-stage Plasmodium chabaudi AS infection. Infect. Immun. 68, 4399–4406 (2000).
Pfefferkorn, E.R. Interferon gamma blocks the growth of Toxoplasma gondii in human fibroblasts by inducing the host cells to degrade tryptophan. Proc. Natl. Acad. Sci. USA 81, 908–912 (1984).
MacKenzie, C.R., Hucke, C, Müller, D, Seidel, K, Takikawa, O & Däubener, W. Growth inhibition of multiresistant enterococci by interferon-gamma-activated human uro-epithelial cells. J. Med. Microbiol. 48, 935–941 (1999).
de Souza, A.P. et al. Absence of interferon-gamma-inducible gene IGTP does not significantly alter the development of chagasic cardiomyopathy in mice infected with Trypanosoma cruzi (Brazil strain). J. Parasitol. 89, 1237–1239 (2003).
Boehm, U. et al. Two families of GTPases dominate the complex cellular response to IFN-gamma. J. Immunol. 161, 6715–6723 (1998).
Kim, B.H., Shenoy, A.R., Kumar, P, Das, R, Tiwari, S & MacMicking, J.D. A family of IFN-gamma-inducible 65-kD GTPases protects against bacterial infection. Science 332, 717–721 (2011).
Hardison, J.L., Wrightsman, R.A., Carpenter, P.M., Lane, T.E. & Manning, J.E. The chemokines CXCL9 and CXCL10 promote a protective immune response but do not contribute to cardiac inflammation following infection with Trypanosoma cruzi. Infect. Immun. 74, 125–134 (2006).
Liu, Z. et al. Chemokine CXCL11 links microbial stimuli to intestinal inflammation. Clin. Exp. Immunol. 164, 396–406 (2011).
Khan, I.A. et al. IP-10 is critical for effector T cell trafficking and host survival in Toxoplasma gondii infection. Immunity 12, 483–494 (2000).
Loetscher, M. et al. Chemokine receptor specific for IP10 and mig: structure, function, and expression in activated T-lymphocytes. J. Exp. Med. 184, 963–969 (1996).
Qin, S. et al. The chemokine receptors CXCR3 and CCR5 mark subsets of T cells associated with certain inflammatory reactions. J. Clin. Investig. 101, 746–754 (1998).
Whitehead, R.H., VanEeden, P.E., Noble, M.D., Ataliotis, P & Jat, P.S. Establishment of conditionally immortalized epithelial cell lines from both colon and small intestine of adult H-2Kb-tsA58 transgenic mice. Proc. Natl. Acad. Sci. USA 90, 587–591 (1993).
Stockdale, P.G., Stockdale, M.J., Rickard, M.D. & Mitchell, G.F. Mouse strain variation and effects of oocyst dose in infection of mice with Eimeria falciformis, a coccidian parasite of the large intestine. Int. J. Parasitol. 15, 447–452 (1985).
Wettenhall, J.M. & Smyth, G.K. limmaGUI: a graphical user interface for linear modeling of microarray data. Bioinformatics 20, 3705–3706 (2004).
Gupta, N, Zahn, M.M., Coppens, I, Joiner, K.A. & Voelker, D.R. Selective disruption of phosphatidylcholine metabolism of the intracellular parasite Toxoplasma gondii arrests its growth. J. Biol. Chem. 280, 16345–16353 (2005).
Schmatz, D.M., Crane, M.S. & Murray, P.K. Purification of Eimeria sporozoites by DE-52 anion exchange chromatography. J. Protozool. 31, 181–183 (1984).
Acknowledgements
We thank Grit Meusel, Ina Wagner, Simone Spieckermann, and Ines Heyn for their research assistance. This work was supported by a graduate fellowship to M.S. from the Helmholtz Foundation, and research grants (SFB618/C7, GRK1121) to N.G. and R.L. from the German Research Foundation (DFG). Microarray data have been deposited in the NCBI Gene Expression Omnibus (GEO), and are accessible by an accession number GSE49948.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declared no conflict of interest.
Additional information
SUPPLEMENTARY MATERIAL is linked to the online version of the paper
Supplementary information
Rights and permissions
About this article
Cite this article
Schmid, M., Heitlinger, E., Spork, S. et al. Eimeria falciformis infection of the mouse caecum identifies opposing roles of IFNγ-regulated host pathways for the parasite development. Mucosal Immunol 7, 969–982 (2014). https://doi.org/10.1038/mi.2013.115
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mi.2013.115
This article is cited by
-
Eimeria proteins: order amidst disorder
Parasites & Vectors (2022)
-
Phosphatidylinositol synthesis, its selective salvage, and inter-regulation of anionic phospholipids in Toxoplasma gondii
Communications Biology (2020)
-
Effects of host fatty acid–binding protein 4 on Eimeria tenella sporozoites invasion of cells
Parasitology Research (2019)
-
Lipid analysis of Eimeria sporozoites reveals exclusive phospholipids, a phylogenetic mosaic of endogenous synthesis, and a host-independent lifestyle
Cell Discovery (2018)
-
Dual RNA-seq reveals no plastic transcriptional response of the coccidian parasite Eimeria falciformis to host immune defenses
BMC Genomics (2017)