Abstract
During colonization of germfree mice with the total fecal microbial community of their conventionally born and raised siblings (conventionalization), the intestinal mucosal immune system initiates and maintains a balanced immune response. However, the genetic regulation of these balanced, appropriate responses to the microbiota is obscure. Here, combined analysis of germfree and conventionalized mice revealed that the major molecular responses could be detected initiating at day 4 post conventionalization, with a strong induction of innate immune functions followed by stimulation of adaptive immune responses and development and expansion of adaptive immune cells at later stages of conventionalization. This study provides a comprehensive overview of mouse developmental and immune-related cellular pathways and processes that were co-mediated by the commensal microbiota and suggests which mechanisms were involved in this reprogramming. The dynamic, region-dependent mucosal responses to the colonizing microbiota revealed potential transcriptional signatures for the control of intestinal homeostasis in healthy mice, which may help to decipher the genetic basis of pathway dysregulation in human intestinal inflammatory diseases.
Similar content being viewed by others
Introduction
Mammals are germfree in utero and become colonized by microbes during and after birth following a dynamic, genetically controlled process that results in gut colonization by taxonomically diverse bacterial populations that establish a symbiotic relationship with the host.1 After colonization is completed, the intestine of conventionally raised mice is in continuous contact with a vast diversity of microbes, collectively termed gut microbiota. Despite the exposure to trillions of immunogenic-diverse bacteria, the intestinal mucosa maintains a state of homeostasis that involves tightly controlled immune responses. To achieve this, epithelial cells and immune cells of the lamina propria mount innate and adaptive immune responses that sustain tolerance to microbiota but at the same time will detect and kill invading pathogens.2 The gut microbiota has been proposed to have a crucial role in the establishment and maintenance of adaptive immunity and homeostasis,3 in which the complexity of the microbial community elicits a comparably complex immunological response in the host. Despite our knowledge on biological processes and signaling pathways that have roles in the mucosal immune system,4, 5, 6, 7 our understanding of the genetic regulation of homeostasis is still incomplete. In healthy animals, maintenance of homeostasis is a dynamic process where the composition of the gut microbiota and the presence or absence of even a single microbial species in the gut all contribute to appropriate, tolerant responses in the mucosa.8, 9, 10
In a hallmark study by Gaboriau-Routhiau et al.,9 it was reported that nearly 50% of the genes differentially expressed in the intestine of gnotobiotic mice regulated T-cell development in response to colonizing gut microbiota. The study of Gaboriau-Routhiau et al.9 aimed to unravel the mechanisms by which segmented filamentous bacteria induced mucosal adaptive immune responses, with the main focus on the terminal ileum. More recently, Larsson et al.11 provided a detailed description of the tissue-specific host transcriptional responses to the normal gut microbiota, with the main focus to identify the interaction between the host innate immune responses and microbial composition throughout the gut, by comparing germfree and conventional mice. In the current study, we present the time-resolved, genome-wide immune-related gene expression programs that are elicited in the mucosa of jejunum, ileum, and colon in germfree mice upon their conventionalization, with special attention to immune-related gene expression programs and to the validation of these programs by immunohistochemistry. Our findings show that conventionalization of germfree mice induced multigenic defense- and immune- related transcriptional responses that reflect the sequential activation of innate and adaptive immune responses, most pronounced processes associated with T-cell development and maturation. Moreover, this study enabled the identification of time-resolved transcriptional signatures of genes that are proposed to be involved in the regulation of the dynamic intestinal response to the microbiota and have a key role in the maintenance of mucosal homeostasis.
Results
Dynamic changes in intestinal physiology and morphometry during conventionalization
This study was aimed at identifying the temporal and spatial intestinal mucosal changes in germfree and conventionalized mice as measured in three independent experiments (for an experimental set up, see Supplementary Figure S1 online). As a typical hallmark of conventionalization, the cecal weight was 80% reduced upon conventionalization when compared with germfree mice (Figure 1a); this difference was detected from day 4 post conventionalization onward. Intestinal morphometric analysis revealed an initial significant increase (P<0.05) in the intestinal crypt depth in the small intestine and the colon of conventionalized mice four days post conventionalization, which was not yet visible on days 1 and 2 (Figure 1b). In the small intestine, crypt depth increased continuously during the first 16 days. At day 30, small intestinal crypt depth remained higher as compared with the germfree mice. By contrast, the colon crypt depth reached a maximum level from day 8 to day 16 but at day 30, had returned to crypt depths that were also measured in the germfree mice (Figure 1b). Concomitant to the lengthening of the crypts, the lamina propria in both the jejunum and ileum expanded by a global increase in connective tissue cells, with a simultaneous increase in the abundance of different types of immune cells (see Supplementary Figure S2A online).
Effect of microbial colonization on the intestinal physiology and morphology. (a) Total cecal weight was determined as a measure of bacterial colonization, and the inset shows a photograph of cecum at day 0 (left) and day 4 (right), respectively. (***P=0.001 compared with germfree). (b) Mean crypt depth measured from villi and crypts from the jejunum, ileum, and colon in germfree and conventionalized mice at different time points post conventionalization (n=4–6 mice/day). Results are presented as means ± SD. Significant differences between time points are indicated by distinctive characters above the measurement groups. (c) Immunohistochemical detection of Ki-67–positive cells using Mib-1 antibody in ileal tissues in (A) germfree, (B) day 4, and (C) day 30 post conventionalization. Arrows refer to the positive-stained cells (brown color) (n=4–6/day). (d) Representative high iron diamine (HID)-stained colon sections, showing the dynamics of mucin subtypes distribution in (A) germfree, (B) day 4, and (C) day 16 post conventionalization. Arrows refer to sialylated mucins stained blue and sulfated mucins stained brown/black (n=4–6/day).
Conventionalized mice at later time points (days 4–30) had higher number of Ki-67 (cellular marker for proliferation)–positive cells compared with germfree animals (Figure 1c). Ki-67–positive cells predominantly localized in the crypts, but were also seen in the inter-villus region and in the lamina propria of the small intestine starting at day 8. Ki-67–positive and –negative cells in colonic crypt epithelia revealed a maximal staining of positive cells on days 4 and 8, followed by a decline at days 16 and 30 (P<0.05; see Supplementary Figure S2B–C online).
High iron diamine-alcian blue stain, which detects the mucin load of goblet cells and discriminates between sialylated and sulfated mucins, showed that in the colon, day 4 was characterized by a transient domination of sialylated over sulfated mucin-containing goblet cells (Figure 1d). Taken together, these results clearly illustrate the region-specific transient and permanent changes in the intestinal morphology and cell proliferation, correlating with longer microbial colonization. These changes did become most pronounced from day 4 post conventionalization onward.
Establishment of the gut microbiota during conventionalization
To evaluate whether the above mentioned intestinal changes were accompanied by correlating changes in the abundance of microbial groups or just a consequence of increased numbers of bacteria without changing the ratios of specific bacterial groups, samples from jejunum, ileum, and colon were collected at days 1, 2, 4, 8, and 16 post conventionalization and were compared for 16S RNA gene diversity among each other and with the inoculum. Quantitative PCR detection of 16S rRNA gene copies in colon samples indicated that a full-sized microbial community was very rapidly established, i.e., already on day 1 post conventionalization the microbial community contained approximately 11.6±0.5 16S rRNA copies/g colon content (expressed in log10). This community size estimate did not significantly change during the experiment, indicating that the microbial community reached its climax size in a single day. Molecular fingerprinting of the composition of the colonizing microbiota was performed using MITChip analysis, a 16S rRNA-based phylogenetic array specifically designed to classify murine microbiota.12, 13 MITChip analysis revealed that the colon microbial diversity remained relatively low during days 1 and 2 and significantly increased (P=0.001) at later time points of conventionalization, reaching a stable diversity level on days 8 and 16. This diversity resembled that of the original inoculum (Figure 2a). Pearson correlation-based similarity analysis of MITChip profiles of the colon samples indicated that the similarity of the colon microbiota relative to the inoculum increased from approximately 60% during early days to 80% during later days of conventionalization (Figure 2b), indicating that the climax community at days 8 and 16 was indeed comparable with that of conventional mice (inoculum). This level of similarity (80%) corresponded to what was found when the similarity of the colon microbiota was assessed in individual mice at days 8 and 16 post conventionalization (see Supplementary Table S1 online). MITChip analysis also revealed that day 1 was characterized by a higher relative abundance of Gram-negative Bacteroidetes, whereas later stages of conventionalization (days 8 and 16) showed an expansion of the relative abundance of the Gram-positive Firmicutes (Figure 2c). The expansion of the Firmicutes phylum was particularly large for the members of Clostridium clusters IV and XIVa, while the initial (days 1 and 2) abundance of the bacilli declined upon prolonged conventionalization (see Supplementary Figure S3A online). Finally, multi-variate analysis by redundancy analysis of colon- and small intestine (jejunum and ileum)-derived microbiota profiles clearly established that each intestinal region did harbor different microbial consortia; especially the diversity of the small intestine community appeared to be significantly lower as that encountered in the colon (see Supplementary Figure S3B–C and S4 online). This outcome indicates that the colonization of the gut in the conventionalized mice was efficient and representative for normal colonization levels that are reached in conventionally raised mice.
Establishment of gut microbiota during conventionalization. (a) Diversity of the total colon microbiota at different time points post conventionalization, expressed as Simpson index of the hybridization profiles analyzed by the MITChip. (b) Pearson correlation similarity index of the MITChip profiles from colon samples at different time points post conventionalization, including the comparison to the inoculum. Significant differences between time points are indicated by distinctive characters above the measurement groups. (c) Dynamics of the relative contribution of different microbial groups (level 0, which is similar to phylum level phylogeny) to the overall microbiota in the colon of mice at different time points post conventionalization, and in comparison to the inoculum. d=day (n=5–6/day).
Induction of local antimicrobial defense and surface receptors at day 4 post conventionalization along the gastrointestinal (GI) tract
In order to investigate the pathways underlying the mucosal changes observed, tissue gene expression patterns of jejunum, ileum, and colon at all time points post conventionalization were compared with each other. The Short Time series Expression Miner (STEM) and GO-enriched bayesian clustering were used to identify genes with similar, time-dependent gene expression patterns over the 30-day timespan of conventionalization. STEM time-series analysis and GO-enriched bayesian clustering (for detailed description, see Supplementary Methods online) demonstrated that nearly 40% of the genes regulated in response to conventionalization were annotated with immune-related GO terms (P<0.001), in a time- and region-dependent manner (see Supplementary Figure S5, Supplementary Table S2, S3 online). Significant induction of expression of surface receptors involved in microbial recognition was detected at day 4 post conventionalization throughout the GI tract. These receptors included the lipopolysaccharide receptor Cd14, the intracellular signaling adaptor protein, Myd88 and the Toll-like receptors Tlr1, 2, 8, 9, and 12 but not Tlr4 or 5 (see Supplementary Figure S6 online). In parallel, the expression levels of several antimicrobial peptides, including lysozyme P (LyzP), regenerating islet-derived protein (Reg) IIIβ and -γ, resistin-like beta (Retnlβ), and phospholipase A2 (Pla2g2a) had significantly increased (Figure 3a). Notably, prolonged exposure to microbiota (30 days) retained increased expression levels of RegIIIβ and -γ in the small intestine, but returned to the germfree level in the colon. By contrast, the expression levels of Retnlβ and Pla2g2a returned to germfree levels in the small intestine, but remained high in the colon. Indeed, immunohistochemical (IHC) analysis verified the LyzP loading of secretory granules in the Paneth cells in the small intestine at day 4 (Figure 3b). The increased lysozyme loading of granules, indicative of activation of an innate immune response program, was in agreement with the coinciding increased expression of matrix metalloprotease 7 (Mmp7) (Figure 3a) that regulates the activity of defensins in intestinal mucosa via proteolytic processing of the defensin precursors.14 IHC analysis also confirmed the peak production of RegIIIγ at day 4 (Figure 3c). The gene expression and IHC data show that transient induction of innate immune factors was region dependent and was most pronounced after four days of conventionalization.
Dynamics of induction of innate immune molecules during conventionalization. (a) Gene expression levels of RegIIIγ, RegIIIβ, Retnlβ, and Pla2g2a, in jejunal, ileal, and colonic tissues, and Mmp7 in jejunal and ileal tissues from germfree and conventionalized mice at indicated days post conventionalization. Individual values determined in the animals and their medians are shown. Significant differences between time points are indicated by distinctive characters above the measurement groups (P<0.05; n=6–11/day) (b) Representative IHC of lysozyme-P in ileal tissues from (A) germfree, (B) day 1, (C) day 2, (D) day 4, (E) day 8, and (F) day 16 post conventionalization (n=4–6/day). (c) Representative immunohistochemistry of RegIIIγ in ileal tissues from (A) germfree, (B) day 4, and (C) day 30 post conventionalization. Arrows indicate positively stained cells (n=4–6/day). The corresponding transcriptome data for lyzP were not shown because it is not trivial to assign gene expression signals to this gene due to ambiguous gene identifiers in the array datasets. Instead, the expression level of matrix metalloprotease 7 (Mmp7) that regulates the activity of defensins in intestinal mucosa via proteolytic processing of the defensin precursors14 is presented.
Pro-inflammatory cytokine induction and antigen presentation at day 4 activates time- and region-dependent adaptive immune responses during later days of conventionalization
To further assess time- and region-dependent induction of innate and adaptive immune responses during conventionalization, the temporal expression profiles of specific cytokines were used as markers for the release of pro-inflammatory signals and attraction of immune cells in the gut tissues. Tumor necrosis factor alpha (Tnf-α) and interferon gamma (Ifn-γ) were expressed significantly higher throughout the gut upon conventionalization (Figure 4a). In the small intestine, their expression increased from day 4 onward, and peaked at day 16 post conventionalization, whereas in the colon, peak induction occurred at day 4, followed by a decline of expression at later time points and a gradual return to the levels seen in germfree mice (Figure 4a). To investigate whether elevated expression of pro-inflammatory cytokines coincided with the expected induction of surface expression of major histocompatibility complex (MHC) class I and II complexes, the dynamics of expression of the associated genes were investigated. Members of the MHC class I complex and their activators were induced from day 4 onward throughout the GI tract, whereas the induction of members of the MHC class II complex and their transactivator (Ciita) appeared to occur at later time points throughout the GI tract, mainly peaking at days 8 and 16 post conventionalization (Figure 4a,b).
Expression of TNF- α, Ifn-γ and major histocompatibility complex (MHC) class I and II complexes. (a) Jejunum, ileum, and colon gene expression levels of TNF- α and Ifn-γ, and MHC class I and II complex activators were analyzed in germfree and conventionalized mice at indicated days post conventionalization. Values are depicted as box and whisker diagrams (top-to-bottom, maximum value, upper quartile, median, lower quartile, and minimal value, respectively). Any data not included between the whiskers is plotted as an outlier with a dot. Significant differences between time points are indicated by distinctive characters above the measurement groups (P<0.05), (n=6–11/day). (b) Heat map generated from the significantly expressed MHC class I and II genes (P<0.05) between the germfree and conventionalized mice at the indicated time points (n=6–11/day).
Time- and region-specific adaptive immune system development
As anticipated, the increased expression of pro-inflammatory cytokines and MHC class I and II molecules elicited the induction of expression of genes required for immune (T) cell function and development on day 8 (colon) and day 16 (small intestine) post conventionalization throughout the intestinal tissues. The most prominent among these genes were the T-cell accessory molecules that participate in antigen response, inflammatory chemokine ligands (Cxcl9, 10, Ccl2, 3, and 5), and chemokine receptors (Cxcr3, Ccr2, and 5) (see Supplementary Figure S7 online). Increased expression of these genes coincided with the increased villus width and lamina propria cellularity in the small intestine that were observed in hematoxylin and eosin-stained mucosal tissue sections (see Supplementary Figure S2A online).
GO-enriched bayesian clustering was used to further detail the biological functions and signaling pathways involved in the time- and region-dependent events related to immune (T) cell activation and development. Detailed inspection of the gene set annotated with GO category “T cell activation” (see Supplementary Information online), allowed to further explore the tissue distribution of T cells. This gene set was upregulated from day 4 onward in ileum and from day 8 to 30 throughout the intestine (see Supplementary Figure S8 online) and included the surface markers of T-cell infiltration;15 Cd3ɛ, Cd4, and Cd8.
IHC was used to verify that the inferred gradual increase of T cells expressing the mentioned surface markers, with the largest numbers in the small intestine, did indeed occur. Microscopic inspection of sections hybridized with the appropriate antibodies showed increased numbers of cells positive for the T-cell maturation markers Cd3ɛ and Cd8 at day 8 onward and highlighted the prominent localization of Cd8-positive cells along the epithelial lining of the small intestine (Figure 5). Compared with Cd8+ and Cd3ɛ+ cells, cells positive for the Cd4 marker were observed at lower numbers in the lamina propria of the small intestine at day 16 post conventionalization (Figure 5). Notably, prolonged conventionalization (day 30) revealed that the increase in T-cell numbers (especially Cd8+) had continued in the small intestine but had already reached a more steady level in the colon between days 8 and 16. Altogether, the increased numbers of cells positive for typical markers of T-cell activation and maturation indicate activation and development of the adaptive arm of the immune system. Development of adaptive immunity appeared to have reached a climax level on day 8 in the colon and day 16 in the small intestine and was likely a consequence of the strong activation of the innate immune response that was apparent at day 4 post conventionalization.
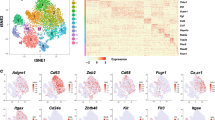
Regional variation of increasing T-cell numbers and maturation. Representative immunohistochemistry of (a) Cd8a, (b) Cd4 and (c) Cd3ɛ in ileal tissues from (A) germfree, (B) days 16 and (C) 30 post conventionalization, (n=4–6/day). Dot plots represent the expression levels of Cd8a, Cd4 and Cd3ɛ in jejunal, ileal, and colonic tissues in germfree and conventionalized mice at indicated days post conventionalization. Individual values and medians are shown. Significant differences between time points are indicated by distinctive characters above the measurement groups (P<0.05; n=6–11/day). All panels are shown at the same magnification; arrows indicate positively stained cells (brown color).
Temporal expression profile of the negative regulation of the activated immune response along the gut
So far, pro-inflammatory signals have been shown to activate innate and adaptive immunity in response to the microbiota. No signs of disease were noticed in the mice during conventionalization and no microscopic signs of damage to the intestine or any infiltration of immune cells were identified. The microbiota were therefore assumed to have induced a tolerant immune response, implying the co-induction of negative regulators together with the more pro-inflammatory molecules such as Tnf-α and Ifn-γ that were induced (see previous two sections). To investigate this, first the temporal expression profiles of immune-suppressive cytokines were analyzed along the gut. As expected, the induction of immune responses coincided with the elevation of expression of tolerance-associated molecules, starting on day 8 post conventionalization and continuing during later time points with a climax level on day 8 in the colon and day 16 in the small intestine. These molecules included Foxp3, the marker for regulatory T cells or Tregs,16 interleukin-10 (Il10), which enforces immune tolerance,17 Tbx21 (T-bet), a transcription factor that drives Th1 cell maturation,18 and Ctla4 (cytotoxic T-lymphocyte antigen 4), which transmits an inhibitory signal to activated T cells19 (Figure 6). Collectively, the gene expression profiles and IHC studies appear to correlate with region-specific induction of pro- and anti-inflammatory signals that together drive balanced, tolerant (adaptive) immune responses to the microbiota.
Increased expression of tolerance-associated functions at later time points of conventionalization. Jejunum, ileum, and colon gene expression levels of regulatory cytokines were analyzed in germfree and conventionalized mice at indicated days post conventionalization. Values are depicted as box and whisker diagram (top-to-bottom; maximum value, upper quartile, median, lower quartile, and minimal value, respectively). Any data not included between the whiskers is plotted as an outlier with a dot. Significant differences between time-points are indicated by distinctive characters above the measurement groups (P<0.05), (n=6–11/day).
Time-resolved transcriptome signatures for the dynamic, region-specific mucosal responses to the colonizing microbiota
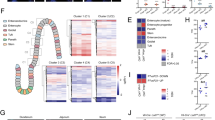
From the time-series transcriptome analysis, we hypothesized that there might be a core set of regulatory genes that could serve as transcriptional signatures for the re-establishment of mucosal homeostasis upon conventionalization along the gut. Therefore, all the genes associated with the temporal expression profiles identified by the time-series analysis, in each of the three intestinal regions (see Supplementary Table S2 online) were mined to search for potential transcriptional signatures using ingenuity pathway analysis (IPA; see Supplementary Methods online for detailed description).The resulting IPA-derived network (Figure 7) exemplified the strong impact of conventionalization on both innate and adaptive immune gene expression throughout the gut mucosa, and encompassed several core regulatory genes that are known to control the induction of innate and adaptive immune responses. The identified core gene set included the major gene categories that were strongly induced from day 4 post conventionalization onward, including nodes belonging to bacterial recognition (Cd14), pro-inflammatory cytokines (Tnf-α and Ifn-γ), chemokines (Ccl5, Ccr5, Cxcl9, Cxcr3, and Ccl8), and MHC Class I (Psmb8, 9, Tap1,2, and H2-Q). Moreover, MHC Class II molecules (Ciita, H2-Ab1, H2-DMa, and H2-DMb1), T-cell differentiation and maturation (Lck, Lat, and Zap70), cell surface markers (Cd3ɛ, Cd4, and Cd8), and B-cell differentiation (Ptprc encoding the Cd45 antigen) were among the identified gene categories. Notably, at 8, 16, and 30 days post conventionalisation, all the identified genes were induced (Figure 7b) in a region-specific manner, i.e., day 8 in the colon and day 16 in the small intestine.
Time-resolved transcriptome signatures for the dynamic, region-specific mucosal responses to the colonizing microbiota. (a) The ingenuity protein–protein interaction network derived by plotting STEM (Short Time series Expression Miner) output genes involved in the temporal expression profiles involved in immune response in the jejunum (J), ileum (I), and colon (C). Transcriptional data was projected onto the interaction map. Red arrows refer to genes associated with inflammatory bowel disease. (b) Heat map of the genes that constitute the core regulatory network (n=6–11).
In parallel, a protein–protein interaction map was generated from cluster-driven time-series analysis of GO categories using bayesian statistics (see Supplementary Figure S9 online). This network representing tissue responses to microbial colonization combined genes that belong to T-cell differentiation and maturation, again showing that, as in the IPA output above, T-cell selection,—induction and—differentiation pathways are among the most important induced mucosal pathways during mouse conventionalization. Also this core regulatory network, constructed using different approaches and statistical methods, contained identical major regulatory nodes as found in the IPA output, supporting the prominent roles of genes involved in T-cell differentiation and maturation in the tissue response to commensal microbial colonization.
Among the identified tissue transcriptome signatures, the human counterparts of 13 genes have known roles in inflammatory bowel disease (indicated with red arrows in Figure 7a). These findings support the biological relevance of the identified transcriptional signatures for mucosal control of homeostasis along the gut.
Discussion
It has been widely recognized that the interplay between gut microbiota and the host is crucial for the proper development of the (adaptive) immune system3 and that dysregulation of this interaction contributes to the development of inflammatory bowel disease symptoms in human.20 There is a clear requirement for tightly controlled genetic regulation of appropriate, tolerant responses to the microbiota.21, 22 This is also supported by the finding that mutated forms of genes involved in the regulation of basal immunological processes such as microbial uptake are strongly associated with the inflammatory bowel disease phenotypes.23
The present study implemented the genome-wide expression profiling of genes during microbial colonization of germfree mice, using a time-series design with six time points. We found that throughout the intestine, the largest proportion of differentially expressed genes was involved in the development of mucosal immune system. Our study corroborated several important findings from the studies by Gaboriau-Routhiau et al.9 and Larsson et al.11 These three studies consistently identify “immune response” as the largest category of genes that is regulated in response to microbial colonization. Our transcriptome analysis showed that there was a time- and region-dependent enrichment of genes involved in balanced innate and adaptive immune responses. These tolerant responses ensured that a novel state of homeostasis was reached within 30 days of conventionalization. Unlike days 1 and 2, which did not show any remarkable changes in the mucosal transcriptome or histological staining, day 4 post conventionalization consistently stood out in the transcriptome analyses and was characterized by drastic changes in gene transcription. For instance, gene expression could switch from induction to repression and vice versa; and some genes were no longer expressed, whereas others were expressed for the first time. At this time point, the activation of cascades of genes involved in innate immunity and initiation of adaptive immune (T) cell activation and maturation was most pronounced.
Strikingly, both transcriptome and IHC analysis for cytokines, chemokines, T-cell surface markers, immune cell transcription factors, and histological stainings of innate immune parameters showed that a novel homeostasis had been reached in the colon within 8 to 16 days, whereas establishment of homeostasis in the small intestine required 16 to 30 days of conventionalization, roughly double the amount of time. Remarkably, the largest shift in the microbiota composition coincided with the most comprehensive shift in the expression of mucosal genes that regulate the host immune response. Microbial profiling of the colon microbiota during conventionalization indicated that the microbial colonization proceeds via the rapid (one day) appearance of early colonizers, followed by the establishment of a stable community that resembles the microbiota of the conventional donor animals. As recently reported,22 our data support the notion that inflammatory tissue conditions were avoided by Tregs, inferred from increased expression of T-bet, Foxp3, and Il10, markers for tolerance-promoting Tregs that were induced especially from day 8 onward. Interestingly, these Tregs markers and other cytokine markers showed a tendency towards increasing expression in jejunum and ileum throughout the experiment. However, in the colon, expression of the same tolerance markers clearly peaked at day 8 and subsequently declined at days 16 and 30 post conventionalization, always remaining higher than the levels observed in the germfree state and during the first two days of conventionalization. Notably, the climax expression level of Tregs at day 8 post-conventionalization in the colon coincides with the colonization by Clostridium groups, which have recently been reported to stimulate the expression of colonic regulatory T cells.10
Similar differential expression patterns in small intestine vs. colon were also observed for six inflammatory chemokine ligands and the corresponding three receptors. We propose that the expression of these chemokines contributed to T-cell chemotaxis. IHC showed that at day 8, Cd8+ T cells were predominantly localized near the mucosal epithelia, which likely resulted from epithelial chemotactic chemokine secretion and expression of MHC class I molecules. This timing of Cd8+ T-cell accumulation in response to accumulation of Th1 chemokines in “danger zones” is in line with the results reported by Valbuena et al.24 during bacterial infection of mice. Moreover, the faster accumulation of Cd8+ T in the lamina propria of the colon at day 8, but at day 16 in the small intestine, illustrates an important location difference that is relevant in the context of establishing homeostasis.
No changes in the expression level of Il-17 were noted throughout the GI tract during the process of conventionalization (not shown), suggesting that the colonization of the C57BL/6 J mice with their normal fecal microbiota did not induce Th17 differentiation. This finding corroborates the results of Ivanov et al.25 who reported that colonization of germfree C57BL/6 mice purchased from the Jackson laboratories did not lead to Th17 differentiation in their lamina propria.
Epithelia contain, in addition to the common enterocytes that are mainly involved in metabolic functions, specialized Paneth cells that secrete high amounts of a broad range of antimicrobial peptides and goblet cells that secrete mucins. One of the broad spectrum antimicrobials secreted by Paneth cells, RegIIIγ, was induced in this study in agreement with Gaboriau-Routhiau et al.9 and Larsson et al.,11 together with the related RegIIIβ. The expression of these two peptides appeared to peak at day 4 post conventionalization, in particular in the ileum as supported by IHC analysis. This could reflect a pronounced induction of innate immune responses at day 4 post conventionalization, corroborated by peak expression levels of the genes encoding the innate immune molecules Retnlβ and Pla2g2a, and the coinciding increased lysozyme P load of secretory granules in Paneth cells. Although innate immunity was clearly induced in all the sampled regions of the gut, its dynamics over time was distinct per region. For example, RegIIIβ and RegIIIγ expression peaked at day 4 post conventionalization in both small intestine and colon, and high level expression was retained in the small intestine but declined to germfree levels in the colon at later time points. Conversely, expression of Retnlβ and Pla2g2a peaked also at day 4 but returned to germfree levels in the small intestine, whereas expression remained high in colon. These data suggest that RegIIIβ and RegIIIγ are important to keep microbes at bay in the small intestine,26 whereas this antimicrobial function is predominantly exerted by Retnlβ and Pla2g2a in the colon. This is in agreement with the presence of RegIII peptide-producing Paneth cells in the small intestine and their absence in the colon. By contrast, Pla2g2a and Retnlβ are secreted by goblet cells,27, 28 a cell type that is common in the colon. These results show that, in healthy germfree mice and during bacterial colonization, innate immune responses are the first line of defence against microbiota, and that this response displays regional (small intestine vs. colon) differences in terms of molecules and expression levels. Other responses of epithelia to increasing bacterial colonization were the increased proliferation of crypt epithelial cells and villus connective tissue cells, measured as Ki-67 expression starting at day 4 post conventionalization, and the transient lengthening of the crypts, measured as crypt depth, also starting at day 4. Our data corroborate results obtained by Cherbuy et al.,29 demonstrating the role of microbial colonization in maturation of epithelial cells in gnotobiotic animals.
The change in biochemistry of colon mucins at day 4 post conventionalization, characterized by a reduction in the amounts of sulfated, thus stronger antimicrobial mucins, compared with sialylated, less antimicrobial mucins,30, 31 could have led to a more intense contact between microbiota and epithelia. This could indeed be shown using the bacterial FISH EUB338 probe (see Supplementary Figure S10 online). It seems that the biochemical changes of the mucin barrier at day 4 post conventionalization may have allowed a more intense contact between the microbiota and the mucosa, which then primed innate immune responses that were followed by adaptive immune responses four days later.
The temporal and spatial analysis presented in this study provides a solid catalogue of genes, pathways, and histology of intestinal adaptations of germfree mice to microbial colonization, thereby providing an important resource that complements various other studies of mouse intestinal colonization by microbiota. Taken together, the data presented here show that a novel state of homeostasis was reached within 30 days following the conventionalization of germfree mice. Homeostasis appeared to be established earlier in the colon (days 8 and 16) as compared with the jejunum and ileum (days 16 and 30). We show that activation of the adaptive immune system mainly involved T cells, not B cells, both in the small intestine and in the colon. The extensive transcriptome datasets for jejunum, ileum, and colon identified a time-resolved transcriptional signature of genes that appear to regulate the major tissue transcriptome changes throughout the intestine during the 30-day conventionalization. The identified signatures included several genes of which the human orthologues are inflammatory bowel disease-associated genes that have also been discovered in genome-wide association studies, suggesting their relevance for the mucosal control of homeostasis, and supporting their importance in the dysregulation of immune-associated pathways in inflammatory bowel disease patients.
Methods
Animals, experimental design, and sampling. All procedures were carried out according to the European guidelines for the care and use of laboratory animals and with permission 78–122 of the French Veterinary Services. Germfree and conventionalized mice (male, C57 BL/6 J) were maintained in sterile conditions, on a commercial laboratory chow diet. Three independent biological experiments were performed using mice of different age. After 2 weeks of acclimatization and diet adaptation, a first set of germfree mice (n=3) were randomly assigned to sacrifice by oral anesthesia using isoflurane. The remaining germfree mice were conventionalized by oral gavage with 0.5 ml of mixed fecal suspension obtained from 0.2 g of freshly obtained fecal material of conventionally raised mice (C57 BL/6 J) diluted 100-folds in brain heart infusion broth. In the first two experiments; conventionalized mice were killed at days 1, 2, 4, 8, and 16 post conventionalization (n=3 per group per experiment). In the third experiment; conventionalized mice were killed at days 4 and 30 post conventionalization (n=4–5 per group). Small intestine (jejunum, and ileum), and colon from each mouse were removed. The two segments of the small intestine and the entire colon were then divided into 2 cm segments that were immediately stored in RNAlater at room temperature for 1 h before subsequent storage at −80 °C for RNA isolation, fixed overnight in 4% (wt/vol) paraformaldehyde or snap frozen and stored at −80 °C for IHC procedures. Luminal content from intestinal segments was removed by gentle squeezing, snap frozen, and stored at −80 °C for microbiota analysis (see Supplementary Figure S1 and Supplementary Methods online).
Histology and immunohistochemistry. In all, 4 μm-thick cross sections of the 2 cm intestinal segments fixed in 4% (wt/vol) paraformaldehyde and paraffin-embedded were stained with haematoxylin (Vector Laboratories, Burlingame, CA) and eosin (Sigma-Aldrich, Zwijndrecht, the Netherlands). To detect morphometric differences, 12–15 well-oriented villi and crypts were chosen per intestinal segment and measured. Mucin histochemistry was performed using high iron diamine-alcian blue as described.32 For Lysozyme-P detection, sections were incubated with anti-Lysozyme P (1:50 in PBS, DakoCytomation, Denmark). For Cd3ɛ and Ki-67 detection, sections were incubated with anti-Cd3ɛ (DAKO, Heverlee, Belgium) or anti-Ki-67 (NovoCastra Laboratories, Newcastle upon Tyne, UK), respectively. Expression of RegIIIγ was detected using a custom-made antibody (for detailed descriptions, see Supplementary Material online). For Cd4–8 detection, cryostat sections were incubated with anti-Cd4 and anti-Cd8 (DakoCytomation). Primary antibodies were detected using VECTASTAIN ABC Elite kit (Vector Laboratories), including biotinylated Donkey anti-rat serum (Sigma-Aldrich) using the manufacturer's instructions. For all stainings, nuclei were counterstained with haematoxylin (Vector Laboratories). Stained tissues were examined using a Nikon Microphot FXA microscope (for detailed descriptions, see Supplementary Methods online). All data were presented as means±s.d. for the number of animals indicated above. Comparisons of data were performed at each time point using one-way analysis of variance (ANOVA) followed by Tukey's Studentized range test (GLM, SPSS program, Chicago, IL). For all parameters P<0.05 was considered the level of significance.
Microbial profiling of intestinal luminal contents. Luminal contents from jejunum, ileum, colon, as well as inoculum were analyzed by Mouse Intestinal Tract Chip (MITChip), a diagnostic 16S rRNA arrays that consists of 3,580 unique probes especially designed to profile murine gut microbiota.12, 13
Quantification of total bacteria was performed using qPCR detection of 16 S rRNA-gene copies, while fluorescent in situ hybridization was used to detect bacteria from tissue samples (for detailed descriptions, see Supplementary Methods online).
Transcriptome analysis. High-quality total RNA was obtained from a 2 cm segment of jejunum, ileum, and colon by extraction with TRIzol reagent, followed by DNAse treatment and column purification. Samples were hybridized on Affymetrix GeneChip Mouse Gene 1.1 ST arrays. Quality control and statistical analysis were performed using Bioconductor packages integrated in an on-line pipeline33 (for detailed descriptions, see Supplementary Methods online). Complementary methods were used for the biological interpretation for the transcriptome data; gene clustering using Multi-experiment Viewer (MeV),34 overrepresentation analysis of GO terms using temporal and location comparative analysis using STEM,35 Bayesian clustering using Genomica, and construction of biological interaction networks using IPA (for detailed descriptions see Supplementary Methods online).
Accession numbers. The mouse microarray dataset is deposited in the Gene Expression Omnibus (GEO) with accession number (GSE32513).
Accession codes
References
Falk, P.G., Hooper, L.V., Midtvedt, T. & Gordon, J.I. Creating and maintaining the gastrointestinal ecosystem: what we know and need to know from gnotobiology. Microbiol. Mol. Biol. Rev. 62, 1157–1170 (1998).
Sansonetti, P.J. & Di Santo, J.P. Debugging how bacteria manipulate the immune response. Immunity 26, 149–161 (2007).
Lee, Y.K. & Mazmanian, S.K. Has the microbiota played a critical role in the evolution of the adaptive immune system? Science 330, 1768–1773 (2010).
Zhu, Y., Yao, S. & Chen, L. Cell surface signaling molecules in the control of immune responses: a tide model. Immunity 34, 466–478 (2011).
Chen, G., Shaw, M.H., Kim, Y.G. & Nunez, G. NOD-like receptors: role in innate immunity and inflammatory disease. Annu. Rev. Pathol. 4, 365–98 (2009).
Hayden, M.S. & Ghosh, S. Shared principles in NF-kappa B signaling. Cell 132, 344–362 (2008).
West, A.P., Koblansky, A.A. & Ghosh, S. Recognition and signaling by toll-like receptors. Annu. Rev. Cell Dev. Biol. 22, 409–37 (2006).
Kuchroo, V.K., Ohashi, P.S., Sartor, R.B. & Vinuesa, C.G. Dysregulation of immune homeostasis in autoimmune diseases. Nat. Med. 18, 42–47 (2012).
Gaboriau-Routhiau, V. et al. The Key role of segmented filamentous bacteria in the coordinated maturation of gut helper T cell responses. Immunity 31, 677–689 (2009).
Atarashi, K. et al. Induction of colonic regulatory T cells by indigenous clostridium species. Science 331, 337–341 (2011).
Larsson, E. et al. Analysis of gut microbial regulation of host gene expression along the length of the gut and regulation of gut microbial ecology through MyD88. Gut (2011)doi:10.1136/gutjnl-2011-301104.
Rajilic-Stojanovic, M. et al. Development and application of the human intestinal tract chip, a phylogenetic microarray: analysis of universally conserved phylotypes in the abundant microbiota of young and elderly adults. Environ. Microbiol. 11, 1736–1751 (2009).
Geurts, L. et al. Altered gut microbiota and endocannabinoid system tone in obese and diabetic leptin-resistant mice: impact on apelin regulation in adipose tissue. Front Microbiol. 2, 149 (2011).
Wilson, C.L. et al. Regulation of intestinal alpha-defensin activation by the metalloproteinase matrilysin in innate host defense. Science 286, 113–117 (1999).
DeJarnette, J.B. et al. Specific requirement for CD3 epsilon in T cell development. Proc. Natl. Acad. Sci USA 95, 14909–14914 (1998).
Hori, S., Nomura, T. & Sakaguchi, S. Control of regulatory T cell development by the transcription factor Foxp3. Science 299, 1057–1061 (2003).
Fujio, K., Okamura, T. & Yamamoto, K. The Family of IL-10-secreting CD4(+) T cells. Adv. Immunol. 105, 99–130 (2010).
Miller, S.A. & Weinmann, A.S. Molecular mechanisms by which T-bet regulates T-helper cell commitment. Immunol Rev 238, 233–246 (2010).
Magistrelli, G. et al. A soluble form of CTLA-4 generated by alternative splicing is expressed by nonstimulated human T cells. Eur. J. Immunol. 29, 3596–3602 (1999).
Abraham, C. & Cho, J.H. Mechanisms of disease. Inflammatory bowel disease. N. Engl. J. Med. 2066–2078 (2009).
Bouma, G. & Strober, W. The immunological and genetic basis of inflammatory bowel disease. Nat. Rev. Immunol. 3, 521–533 (2003).
Geuking, M.B. et al. Intestinal bacterial colonization induces mutualistic regulatory T cell responses. Immunity 34, 794–806 (2011).
Deretic, V. & Levine, B. Autophagy, immunity, and microbial adaptations. Cell Host Microbe. 5, 527–549 (2009).
Valbuena, G., Bradford, W. & Walker, D.H. Expression analysis of the T-cell-targeting chemokines CXCL9 and CXCL10 in mice and humans with endothelial infections caused by rickettsiae of the spotted fever group. Am. J. Pathol. 163, 1357–1369 (2003).
Ivanov, I.I. et al. Induction of intestinal Th17 cells by segmented filamentous bacteria. Cell 139, 485–498 (2009).
Vaishnava, S. et al. The antibacterial lectin RegIII gamma promotes the spatial segregation of microbiota and host in the intestine. Science 334, 255–258 (2011).
Fijneman, R.J.A. et al. Expression of Pla2g2a prevents carcinogenesis in Muc2-deficient mice. Cancer Sci. 99, 2113–2119 (2008).
Krimi, R.B. et al. Resistin-like molecule beta regulates intestinal mucous secretion and curtails TNBS-induced colitis in mice. Inflamm. Bowel Dis. 14, 931–941 (2008).
Cherbuy, C. et al. Microbiota matures colonic epithelium through a coordinated induction of cell cycle-related proteins in gnotobiotic rat. Am. J. Physiol. Gastrointest. Liver Physiol. 299, G348–G357 (2010).
Deplancke, B. & Gaskins, H.R. Microbial modulation of innate defense: goblet cells and the intestinal mucus layer. Am. J. Clin. Nutr. 73, 1131S–1141S (2001).
Linden, SK.SP., Karlsson, N.G., Korolik, V. & McGuckin, M.A. Mucins in the mucosal barrier to infection. Mucosal Immunol. 1, 183–197 (2008).
Bogomoletz, W.V., Williams, G.T. & Potet, F. High iron diamine-alcian blue and histochemistry of mucins in colic diseases-20 years later. Gastroenterol. Clin. Biol. 11, 865–868 (1987).
Lin, K. et al. MADMAX – Management and analysis database for multiple ∼omics experiments. J. Integr. Bioinform. 8, 160 (2011).
Saeed, A.I. et al. TM4 microarray software suite. Methods Enzymol. 411, 134–193 (2006).
Ernst, J. & Bar-Joseph, Z. STEM: a tool for the analysis of short time series gene expression data. BMC Bioinformatics 7, 191 (2006).
Acknowledgements
We thank the technical staff in the animal facilities in the lab of J Doré; (INRA, Jouy en Jossas) for assistance with animal sacrifice and sampling. R Raatgeep and CL Menckeberg (Department of Pediatrics, Erasmus Medical Center), A Taverne-Thiele and H Schipper (Cell biology and immunology, Wageningen University), S Brugman (Pediatric Gastroenterology, University Medical Center Utrecht) are acknowledged for their excellent assistance with immunohistochemical staining and data analyses. J Jansen, M Grootte-Bromhaar, M Boekschoten and P de Groot (Division for Human Nutrition, Wageningen University) for their technical support in microarray hybridization and microarray data-quality control and processing. L Loonen and J Wells (Host-Microbe Interactomics, Wageningen University) are thanked for providing the RegIIIγ antibody.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declared no conflict of interest.
Additional information
SUPPLEMENTARY MATERIAL is linked to the online version of the paper
Supplementary information
Supplementary Information 2
To view table content properly, download Genomica software from http://genomica.weizmann.ac.il/ and open the table there. (ZIP 876 kb)
Rights and permissions
About this article
Cite this article
El Aidy, S., van Baarlen, P., Derrien, M. et al. Temporal and spatial interplay of microbiota and intestinal mucosa drive establishment of immune homeostasis in conventionalized mice. Mucosal Immunol 5, 567–579 (2012). https://doi.org/10.1038/mi.2012.32
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mi.2012.32
This article is cited by
-
Selected commensals educate the intestinal vascular and immune system for immunocompetence
Microbiome (2022)
-
Catestatin selects for colonization of antimicrobial-resistant gut bacterial communities
The ISME Journal (2022)
-
Understanding the Connection Between the Gut–Brain Axis and Stress/Anxiety Disorders
Current Psychiatry Reports (2021)
-
Eggshell and environmental bacteria contribute to the intestinal microbiota of growing chickens
Journal of Animal Science and Biotechnology (2020)
-
Imprinting of the immune system by the microbiota early in life
Mucosal Immunology (2020)