Abstract
Cells of the mononuclear phagocyte system (MPS) are found in large numbers in every organ of the body, where they contribute to innate and acquired immunity and homeostasis. This review considers the locations of MPS cells, surface markers that distinguish subsets of monocytes and macrophages, the pathways of MPS differentiation, and the growth factors and transcription factors that guide them. Although the number of MPS sub-populations that can be defined is infinite, the features that unite the MPS remain compelling. Those features clearly include antigen-presenting dendritic cells within the MPS and argue against any basis for separating them from macrophages.
Similar content being viewed by others
Introduction
The concept of the mononuclear phagocyte system (MPS) is derived from research by van Furth and co-workers1, 2, 3, 4 in the late 1960s and early 1970s. MPS embraces a family of cells that includes committed precursors in the bone marrow, circulating blood monocytes and tissue macrophages, and dendritic cells (DCs) in every organ in the body (reviewed in ref. 5 and ref. 6). Numerous anti-macrophage antibodies were produced, with a consequent subdivision of macrophages into many sub-populations.7, 8 Many, but not all, tissue macrophage populations in the mouse were recognized by the anti-macrophage antibody, F4/80.5, 9, 10, 11, 12, 13, 14 The use of F4/80 led to the recognition that macrophages in tissues are numerous and they occupy a precise anatomical niche in relation to other tissue cell types.5 Figure 1 contains some examples of images from these studies. Macrophages have a particularly intimate relationship with epithelial and endothelial cells. In simple epithelia and throughout the capillary and lymphatic circulation, tissue macrophages spread along basement membranes; in stratified and pseudostratified epithelia such as skin, trachea, and cervix, they are integrated within the epithelium. Sinusoidal macrophages, such as those of liver, spleen, and some endocrine organs, have direct contact with the blood. But the separation by endothelium does not prevent pericapillary macrophages from extending processes into the lumen and sampling the blood contents.15, 16 The ability of macrophages to extend processes across epithelia and into lymphatic vessels has also been recognized.17, 18, 19 We commonly think of macrophages as cells of the immune system and forget their central function in many other aspects of embryonic development, homeostasis, and wound repair.5, 6, 20 As an example, the macrophages of the epidermis, known as Langerhans cells, form the center of the so-called epidermal proliferative units and control the proliferation and differentiation of keratinocytes.21, 22, 23 Those lining the surfaces of bone control osteoblast differentiation and calcification,24 and those in the embryo can control the development and nephron endowment in the kidney.25 The central importance of macrophages in development is highlighted by the many systems affected by macrophage depletion in the colony-stimulating factor-1 (CSF-1)-deficient op/op mouse, including somatic growth, development of the pancreas and nervous system, and male and female fertility.5 Resident macrophages become adapted to perform particular functions in different organs; as a result of this, brain macrophages (microglia) are very different from alveolar macrophages of the lung, Kupffer cells of the liver, or the largest tissue macrophage population of those lining the wall of the gut.
Common locations and morphologies of MPS cells as defined by immunolocalization of F4/80 antigen. These sections are from tissues perfusion-fixed with glutaraldehyde and paraffin-embedded, which preserves the ramified processes of tissues of MPS cells. (a) The F4/80+ macrophages in the walls of the afferent and efferent arterioles of the juxtaglomerular complex, where blood flow to the glomerulus glom is regulated. (b) The lamina propria of the intestine, demonstrating the remarkable density of macrophages in this region. Three distinct locations can be seen; cells spread in the plane of the epithelial basement membranes indicated by the long arrow, along the walls of vessels including major vessels indicated by the short arrow and monocyte-like cells indicated by the arrowhead, which are presumably recently arrived. (c) Graphical representation of the non-random distribution of macrophages in tissues; in this case microglia spread in two dimensions in the inner plexiform layer of the retina. (d) A section of bone showing the complete absence of F4/80 from osteoclasts Ocl; the large ramified macrophages that form the center of hemopoietic islands indicated by the long arrow and macrophages within the canopy layer on the bone surface indicated by the short arrow. (e) A major role of macrophages in development; in this case a macrophage that has migrated through retina to engulf a dying neuron in the developing retina. MPS, mononuclear phagocyte system. Original studies.5, 9, 10, 11, 12, 13, 14 Other images with detailed annotation can be seen at www.macrophages.com.
Of course, macrophages are essential for both innate and acquired immunity. A reasonable definition of a pathogen is a microorganism that evades constitutive killing by macrophages; one effector mechanism for the acquired immune system is to increase the microbicidal activity of macrophages, in the process increasing bystander damage and pathology in the host. Mononuclear phagocytes may also become adapted to present antigen to T lymphocytes and can control the precise nature of the T-cell response. A subset of MPS cells express the endocytic receptors, intracellular trafficking and processing pathways, and cell surface major histocompatibility complex (MHC) molecules required to present antigen to T lymphocytes. A subset of these cells is referred to as DCs; whether there is a genuine functional distinction between DCs and other MPS cells in terms of antigen-presenting activity has been debated elsewhere.5 Cells of the MPS may also act as powerful suppressors of T-cell activation and probably have a major function in the development of central and peripheral tolerance. In this review, we will consider evidence of similarities and differences between macrophage and DC differentiation; their relative importance in T-cell activation will not be considered.
Heterogeneity and Markers in the Mononuclear Phagocyte System
Although macrophages in tissues have many features in common, including extensive lysosomes, stellate morphology, and location relative to epithelia, they are nevertheless extremely heterogeneous in terms of function and surface marker expression. Our knowledge of this plasticity is most extensive for mouse.26 Two different transgenic lines have been used to delineate the MPS: the CX3CR1-EGFP27, 28 and csf1r transgenes; MacGreen/MacBlue.29, 30, 31 Both appear to highlight the large majority of tissue cells that we would regard as belonging to the MPS, although both also express in other cells. CX3CR1 is expressed in some endothelial and epithelial cells.27 Csf1r-EGFP is expressed in granulocytes.30 Granulocytes are excluded by definition from the MPS, being “polymorphonuclear” as opposed to “mononuclear,” although as noted below, they clearly derived from a common committed myeloid progenitor. The expression of csf1r-EGFP transgene in granulocytes is actually a reflection of the fact that csf1r, alongside many other macrophage-specific genes, is expressed at the mRNA level in these cells, but the proteins are not translated. In this and several other papers referenced therein, granulocytes could be converted to macrophages in culture.30 Whether this occurs in vivo, in the context of inflammation, remains to be proven or disproven. The advantage of fluorescent transgenes is that they permit ready isolation of tissue macrophage populations (see Figure 2). We used this feature to isolate and characterize the large macrophage population from developing mouse embryo, which have features in common with tumor-associated macrophages.25 The functions and ontogeny of macrophages in the embryo have been reviewed elsewhere.32 An interesting subset of these cells are derived from the yolk sac, do not pass through a monocyte stage, and are not affected by mutation of the macrophage-specific transcription factor, PU.1.33 Macrophages in different adult organs differ in their gene expression profiles; for example, csf1r-EGFP(+) cells isolated from the lung express genes such as tartrate-resistant acid phosphatase, matrix metalloproteinase 9, and cathepsin K in common with osteoclasts (unpublished). Another widely studied MPS reporter gene is a CD11c-GFP/YFP transgene.34 CD11c, along with several candidate endocytic receptors including DEC-205, DC-SIGN, DCL1, and DC-HIL/gpnmb that probably function in antigen uptake, has been advocated as a marker that distinguishes DC from macrophages.35, 36, 37 CD11c in humans was originally defined as a marker for the MPS38 and was subsequently shown to be an active complement receptor 4 that is induced during macrophage maturation.39 The original description of mouse CD11c40 showed that the antigenic epitope detected by a hamster monoclonal antibody was enriched on interdigitating cells in T-cell areas. The more recent studies of CD11c-YFP transgenic mice focused solely on the lymph node, where the transgene permits the study of antigen presentation and antigen-presenting cell (APC) dynamics by live imaging, but did not provide evidence for distinction between macrophages and DCs.34 Most other evidence suggests that CD11c is present on the large majority of macrophages associated with epithelia–mucosal surfaces. The lung contains two major macrophage populations, the alveolar and interstitial cells, that differ in many functions.41 CD11c appears to be expressed on both populations.42 The CD11c-positive cells of the lamina propria of the gut became especially interesting when it was shown, partly through the use of a CD11c-EGFP transgene, that they can extend processes between epithelial cells and sample the lumenal contents17, 18, 43, 44 and appear to specifically express tlr5.45 Vallon-Eberhard et al.46 isolated intestinal lamina propria cells from rag−/− mice and identified two major populations of myeloid cells, both expressing class II MHC and CD11c, but differing in the expression of CX3CR1 and CD14. Both populations expressed the macrophage marker F4/80. In fact, the large majority of lamina propria F4/80-positive/class II MHC-positive phagocytic macrophages are not active APCs, but are powerfully suppressive.47 CD11c also overlaps substantially, if not completely, with F4/80 and CX3CR1-EGFP in the kidney.9, 48 Similarly, in the brain, a subset of microglia,49, 50, 51 and in the liver, a subset of Kupffer cells,52 express CD11c antigen or the CD11c-EGFP/EYFP transgene, and in both locations CD11c is clearly expressed/induced on inflammatory macrophages.
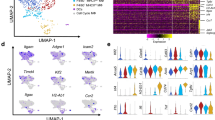
The uses of the MacGreen transgene. In the MacGreen transgene, the csf1r promoter drives the expression of EGFP. Panel a–d at left show EGFP fulorescence in spleen (a), liver (b) lung (c) and small intestine (d). Note the high frequency of positive cells that can be detected by fluorescence-activated cell sorting following enzymatic digestion of tissues. At the right, the remarkable frequency of positive cells in the embryo, especially in areas of high cell death, such as the interdigital area in the left panel and the yolk sac in the right panel. EGFP, enhanced green fluorescent protein.
Molecules expressed on the cell surface are of particular functional interest because they determine the ability of MPS cells to interact with pathogens and with other cell types to generate an appropriate innate and acquired immune response.26 With the possible exception of the CSF-1 receptor,53 which is cleaved selectively from the cell surface in response to Toll-like receptor signaling,54, 55 there are no markers that are expressed specifically and ubiquitously on all MPS cells. Surface markers of mouse MPS cells can be divided into two categories; those that are heterogeneously expressed on individual cells within any one location and those that are enriched on defined functional sub-populations of cells in specific organs or locations within organs.56 Considering the latter category, Taylor et al.26, highlight the distinct distributions of F4/80 and various scavenger receptors and C-type lectins between white pulp, red pulp, marginal zone, marginal zone metalophil, and tingible body macrophages in the spleen. Examples of location-specific expression appear to become less well defined with more detailed observations. For example, langerin (CD207) was originally considered as a marker for Langerhans cells of the skin and appears to be necessary and sufficient for the formation of the Birbeck granules, which are characteristic of these cells.57, 58 A langerin-DT transgene was created and used to demonstrate functions of Langerhans cells in contact sensitivity.59 Subsequently, we find that CD207 is more widely expressed outside the skin and draining lymph nodes,60, 61 and even within skin, on large populations of dermal cells that derive from blood monocytes rather than Langerhans cells.62, 63 In addition, to make matters worse, the distribution of langerin/CD207 appears to vary between mouse strains.64 Similarly, sialoadhesin, which was recently shown to be identical to the MOMA-1 antigen, was originally shown to be expressed most strongly on the marginal metalophilic macrophages of the spleen and the macrophages that form the centers of hematopoietic islands. However, it is also expressed on subcapsular sinus macrophages in lymph nodes, most macrophages of intestinal lamina propria, and Peyer's patch and subsets in central nervous system,65 and is widely expressed on macrophages in the lung,66 as well as uterus.67
Most studies of macrophage marker expression in tissues, or on isolated MPS cell populations, have used only one or two markers and seldom examined them on the same section. One exception was a thorough study of the large dermal macrophage population,68 which demonstrated that most coexpress F4/80, CD11b, and mMGL (a C-type lectin). Subsets expressed sialoadhesin, CD11c, class II MHC, and mSIGN-R1. A recent publication examined the expression of four markers, F4/80, CD68/macrosialin, CD11b, and CD11c in spleen, and normal and inflamed liver, by three-color immunofluorescence.52 Essentially, all combinations of single-, double-, triple-, or quadruple-positive cells were possible in different locations or circumstances. Studies on isolated myeloid cell populations or tissue sections from the skin, brain, lung, gut, ovary, testis, kidney, and lymph node/Peyer's patch8, 9, 10, 13, 41, 43, 47, 48, 51, 56, 60, 64, 67, 69, 70, 71, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 83, 84, 85, 86, 87, 88 reveal similar diversity. In any two-color immunofluorescence profile, one finds all possible null, single-, and double-positive populations. In many cases, the incidence of double-positive cells is no greater than that expected from random intersection of the two frequencies of single-positive cells. Many published studies refer to subsets of cells, e.g., CD11c (hi), as DCs and distinguish them from macrophages. But, it is not clear that there is any one marker, or even a combination of markers, that can provide this distinction. One interesting study examined such heterogeneity in the external muscle layer of the intestine. The authors observed a regular dense planar network of cells with dendritic–stellate morphology.60 These cells are evident in earlier studies of F4/8013 and class II MHC79 immunolocalization, and are obvious in the external muscle and/or mesothelial layers of many internal organs such as bladder, gut, uterus, peritoneal wall, and diaphragm of the MacGreen transgene unpublished. Although direct comparison is not possible and there is the formal possibility that there is a parallel unrelated network of stellate cells, the large majority of these cells must express csf1r and F4/80, and by inference would be considered MPS cells. On account of the regular two-dimensional array of these cells, one can assess the relative frequency of positive cells per unit area in a flat mount. Other macrophage markers, CD14 and Fc receptors, were detected at around 60–70% of the frequency of MHC-II-positive cells, where CD11c or DEC205 was detected at 20–30% frequency, and somewhat less expressed detectable langerin CD207. Since there was no two colours analysis, this study cannot assess the extent of overlap of these markers on individual cells.
If one considers the full set of surface markers that display heterogeneous expression in macrophages, which also include the macrophage scavenger receptors CD36, CD14, SIRPα, Toll-like receptors, other integrins, EGF-TM7 proteins, other Ig superfamily receptors (Siglecs), and multiple C-type lectins,26 the set of combinations and sub-populations is essentially infinite. This is especially true if one identifies subsets of cells on a fluorescence-activated cell sorting profile that express high or low levels of a marker. At least some of this heterogeneity arises because of the stochastic nature of transcriptional regulation, so that there is a genuinely random assortment of surface markers.89 One could take the view that this presents potential pathogens with a formidable arsenal of potential host defense in which every macrophage is unique, especially as we know that some receptors that have functional polymorphisms, such as Toll-like receptor 4, are expressed monoallelically in individual cells.90 In other cases, a particular combination of surface markers determines potential function (e.g., the antigen uptake, presenting, and costimulatory molecules such as CD80, CD86, and CD40) but even in this case, different combinations may provide different T-cell subsets with distinct signals. Surface markers such as the chemokine receptors and integrins must act in concert to determine recruitment and location in a tissue. As discussed below, monocyte subsets differ in their propensity to be recruited. There must be a degree of determination that ensures that certain sets of genes are coexpressed at the right time and place, and this in turn determines a cellular function. The appropriate timing is an important issue, because much of the heterogeneity we see in tissue reflects the fact that macrophages within the tissue are in different stages of their cycle of life and death, migration, development, and function, responding to an infinite combination of signals. Many apparent subsets, such as the CD8-positive and CD8-negative APC of spleen, are now known to be interconvertible and derived from a common progenitor.91 Similarly, CD11b+ and F4/80+ myeloid suppressor cells in spleen can be stimulated with granulocyte macrophage (GM)-CSF to become functional CD11c-positive APCs.92 So, surface marker expression cannot be taken as the sole indication of lineage, function, or destiny.
Differentiation in the Mononuclear Phagocyte System
Committed cells within the mononuclear phagocyte lineage progress through a series of defined morphologically distinct stages; a common myeloid progenitor shared with granulocytes giving rise to monoblasts, promonocytes, and thence monocytes, which migrate into tissues.5 The production of mononuclear phagocytes from progenitor cells is directed by CSFs, to some extent lineage restricted and hierarchical in their actions. They include macrophage CSF (CSF-1), granulocyte-macrophage CSF (GM-CSF), and fms-like tyrosine kinase 3 ligand. Natural mutations in the CSF-1 gene provided the evidence that CSF-1 is required for the production of a substantial subset of tissue macrophages in the mouse (op/op) and rat (tl/tl), at the same time highlighting the importance of tissue macrophages in many aspects of normal development.93, 94 Initially, analysis of the op/op mutation suggested that CSF-1 was not required for DC development, but subsequent knockout of the CSF-1 receptor gene in the mouse produced an even more penetrant phenotype,95 perhaps due in part to the existence of a second ligand for the CSF-1 receptor, interleukin (IL)-34.96 Both the CSF-1 and CSF-1R mutations do cause significant reductions in populations of CD11c-positive cells and their presumptive precursors.95, 97 Injection of CSF-1 into mice expanded the blood monocyte and tissue macrophage populations,98 and more recently was shown to expand circulating CD11b/CD11c-positive mononuclear cells in blood99 and the splenic population of CD11c-positive cells as well as plasmacytoid DCs (ref. 100; DA Hume, unpublished). A key discovery in the DC field was the observation that GM-CSF alone, or its combination with IL-4, can promote the expansion–differentiation of bone marrow and blood monocytes into cells with potent APC activity.101, 102, 103, 104, 105, 106, 107, 108 Interestingly, combined null mutations in CSF-1, GM-CSF, and G-CSF do not completely ablate macrophage numbers or their recruitment in mice.109 This may, in part, be attributable to substitution by IL-34 which acts on the CSF-IR.96 In addition, fms-like tyrosine kinase 3 ligand110 and vascular endothelial growth factor-A,111, 112 both of which act through tyrosine kinase receptors related to the CSF-1R, are partly responsible for age-dependent compensation of the osteopetrotic phenotype of CSF-1-deficient mice, and fms-like tyrosine kinase 3 ligand can expand circulating myeloid cell and CD11c-positive APC numbers.113
The cells of the MPS are produced from pluripotent progenitor cells in the bone marrow. These cells require combined stimulus from CSF-1 and factors including IL-1, IL-3, GM-CSF, and interferon-γ to produce colonies in semi-solid medium.114, 115 Early studies of these cells116 demonstrated that the colony-forming cells lacked surface CSF-1 receptors and the costimulatory factors probably act to induce the receptor on the cell surface. Nevertheless, in a detailed study of the CSF-1R locus, the promoter region was found to be in active chromatin and mRNA is expressed at low levels, in these early progenitors.117 Subsequent studies suggest that macrophage-specific transcription factor PU.1, which acts on elements in the promoter and a key enhancer in the first intron,118, 119 is involved in initiating chromatin rearrangement and commitment.120 PU.1 is not absolutely required for early macrophage differentiation,33 and the effect of the knockout depends upon mouse genetic background,121 perhaps because its actions can be compensated by other members of the Ets family.119, 122 The CSF-1R gene is the best understood MPS-specific gene, and its transcriptional regulation has been reviewed in detail elsewhere.93 Like many myeloid promoters, it has a novel proximal promoter architecture, lacking a TATA box or GC-rich elements, and instead has tandemly repeated purine-rich elements. Alongside of PU.1 and other Ets factors, it is responsive to runx1 AML1 and C/EBP factors as activators, and differentiation from primitive precursors apparently involves downregulation of repressors including c-myb, pax5, id2, and foxP1. Each of the regulators that acts upon the csf1r has been attributed functions in myeloid lineage commitment; this area has been reviewed recently.123 Alongside the genes that control what might be called the generic MPS program, there are, undoubtedly, factors that direct subsets of MPS cells to particular phenotypes. Some are surprisingly specialized. The macrophage-specific transcription factor, TFE-C, acts upon a small subset of target genes induced by IL-4, including the G-CSF receptor.124, 125 The related transcription factor, microphthalmia-associated transcription factor, is specifically required for the maturation of osteoclasts.126 The MafB transcription factor was attributed a role in the divergence of macrophage and DC phenotypes, but the knockout in mice had a surprisingly subtle phenotype, including selective loss of F4/80 and disordered cytoskeletal function.127, 128, 129 In fact, genome-wide mouse transcriptome analysis revealed that macrophages are among the most diverse single sources of mRNA130 and express some 60–70% of all predicted nuclear proteins in different states of activation.131 So, if there is a deterministic basis for the plasticity of MPS cells, the underlying control network is exceedingly complex.
Precursors of Tissue Mononuclear Phagocytes
There are a number of different issues surrounding the origins of heterogeneity in MPS, in particular the level of determination of fate during differentiation, the roles of local proliferation in tissues vs. monocyte recruitment, and the lineage relationship between macrophages and DCs. Inaba et al.106 reported that granulocytes, macrophages, and DCs can all be found within single colonies grown in GM-CSF in vitro. Nikolic et al.132 showed similarly that in liquid culture, precursors of progressive commitment retained the ability to produce macrophage or DC when switched between GM-CSF- and CSF-1-containing medium. However, this does not mean that the colony-forming cells are uniform and functionally equivalent. Witsell and Schook133 reported that individual colonies produced in GM-CSF or CSF-1 varied widely in their ability to respond to inflammatory stimuli and in the sets of cytokines they produced. The common lineage relationship between macrophages and DCs in vivo was confirmed with the identification of a common clonogenic progenitor cell in the bone marrow called the macrophage–DC progenitor. Grafting studies indicated that this cell is an immediate precursor of blood monocytes and of tissue macrophages and DCs in many tissues.91, 134 Macrophage–DC progenitors were not efficient at generating resident DC in lymphoid organs, but gave rise to so-called inflammatory or TipDC.91, 135, 136, 137 The conclusion that many tissue CD11c-positive cells arise from blood monocytes was supported in detailed studies of the origins of the Langerhans cells of the skin and their dependence upon CSF-1R signaling.138 A contrary view came from Onai et al.,139 who identified a population of cells that they claimed were both clonogenic and “DC-restricted”. The flaw in this study lies in the use of an antibody against the CSF-1R (CD115) to purify the population. The antibody used, AFS98, was originally isolated on the basis of its ability to block CSF-1 signaling, and we showed recently that CSF-1 does, indeed, compete directly with the antibody for binding to cells.30
In keeping with studies on macrophage–DC progenitor, in the steady state, the CD11c-positive cell populations of the spleen and probably the lymph node were not replenished directly from blood monocytes,91, 134, 140, 141 supporting the view that there is a proliferating local progenitor population. By contrast, monocytes were able to replenish the CD11c-positive populations of the intestine and lung under the same conditions. The presence of locally proliferating macrophage populations has been known for many years. In fact, van Furth and Diesselhoff-den Dulk142 recognized that around half of the splenic MPS cells are derived from local proliferation in the steady state. In studies of bone marrow chimeras, using a ROSA-26-lacZ reporter transgene, mature monocyte–macrophages engrafted into tissues such as the spleen, liver, and lung and formed clusters suggestive of continued local proliferation. Interestingly, engraftment and replacement of tissue macrophages, especially in the brain, was relatively slow.143, 144 This may indicate that resident macrophages occupying specific anatomical niches turn over very slowly. It could also argue in favor of local proliferation, but again earlier studies suggest that in the lung at least, there is little evidence of labeled thymidine uptake, and the half-lives of interstitial and alveolar macrophages in mice are in the order of a month.145
Monocyte Subsets
The issue of whether certain monocytes have a determined fate has also been clarified recently. On the basis of a model system, Randolph et al.146 initially suggested that monocyte fate (macrophage vs. DC) was determined following transendothelial migration. However, a major body of work now recognizes the existence within peripheral blood of at least two functional monocyte states, distinguished by size and the level of expression of markers such as F4/80, ly6C (Gr-1), CD11b, CD43, and the chemokine receptor, CX3CR1.147 They are believed to be the functional equivalent of human monocyte populations distinguished based upon expression of CD14 and CD16.147 The key observation in the field was based upon monocyte adoptive transfer experiments, in which ly6C(+) and CX3CR1(lo) monocytes were selectively recruited to the peritoneal cavity as thioglycollate-elicited exudate cells, where ly6C(−) and CX3CR1(hi) cells were more inclined to replenish resident tissue MPS populations.147 Selective recruitment of the ly6C(+) inflammatory monocyte probably relates to their selective expression of the chemokine receptor, CCR2, and the adhesion molecule, P-selectin. Sunderkotter et al.148 showed that during replenishment of blood monocyte pools following toxic liposome administration, the “inflammatory” ly6C(+) population appears first, and they argue that the two populations represent a differentiation sequence. Their studies of in vitro culture of bone marrow in GM-CSF or CSF-1 support this view,132 and in our hands, the expression of ly6C mRNA in CSF-1-stimulated bone marrow cultures is rapidly downregulated as cells become adherent (DA Hume and JA Hamilton, unpublished). As discussed in detail elsewhere136, the ly6C(+) cells have a very short half-life and can be depleted selectively without loss of the ly6C(−) population. So, even if they are able to differentiate, the normal fate of the “immature” cells may be to leave the circulation before they have an opportunity to become “mature” or “resident” type cells.
The definition of the ly6C(−) monocytes as “resident” was confounded by subsequent findings; the visualization of a subset of monocytes that patrol the endothelial surface.149 These cells were ly6C(−) and CX3CR1(hi), but in response to a microbial or other challenge they extravasated rapidly. Furthermore, these cells displayed a selective ability to produce inflammatory cytokines such as tumor necrosis factor-α. The latter finding was essentially confirmed by Burke et al.,150 who separated blood monocytes based upon the csf1r-EGFP transgene and CD43 expression. This study showed that both populations are depleted from blood in response to microbial or lipopolysaccharide challenge.
The “resident” vs. “inflammatory” definition must also take account of evidence that the monocyte subsets may have different propensities to give rise to particular resident MPS populations, including DCs. Jakubzick et al.83 claimed that the “inflammatory” monocytes repopulate a resident DC sub-population in the lung, distinguished mainly by the expression of another integrin, CD103. “Inflammatory” monocytes are certainly recognized as precursors of so-called inflammatory DCs (reviewed in ref. 136.) In rats, Yrlid et al.151 identified equivalent monocyte sub-populations, suggested that they can interconvert, and also provided evidence that the CX3CR1/CD43-high population can give rise to the so-called veiled cells, APCs of afferent lymph, following transplantation. These might be regarded as resident or inflammatory, depending upon whether the gut mucosa is considered a chronic inflammatory environment.
What is clearly the case is that blood monocytes are heterogeneous in terms of their expression of key molecules, chemokine receptors, and cell adhesion molecules that determine exactly how they will respond to stimuli that recruit them into tissues. What is not yet clear is the extent to which the monocyte subsets can be further subdivided in terms of their effector functions and fates, and whether distinct populations are recruited to distinct stimuli and distinct locations.
Conclusion
MPS is a complex system that can be viewed in different ways. It can be unified based upon a common origin of the cells from pluripotent progenitors, the coexpression of certain markers, dependence upon growth factors especially CSF-1, and common location and morphology of the cells of the system. It can be divided into essentially infinite subsets because of the plasticity of gene expression profiles and the incredible diversity of different stimuli that MPS cells can encounter and respond to (reviewed in refs. 5, 6). As in so many intellectual pursuits, whether one sees MPS subsets as useful constructs depends in some measure upon whether you are a lumper or a splitter,152 and arguments for splitting are generally more contentious. In facing this challenge, it is worth remembering that the MPS functions as a system within individual organs and indeed within the whole body; a diverse population of interacting cells. We may not gain insight or therapeutic benefit by focusing on arbitrarily defined sub-populations.
Disclosure
The authors declare no conflict of interest.
References
van Furth, R. Origin and kinetics of monocytes and macrophages. Semin. Hematol. 7, 125–141 (1970).
van Furth, R. & Cohn, Z.A. The origin and kinetics of mononuclear phagocytes. J. Exp. Med. 128, 415–435 (1968).
van Furth, R. et al. The mononuclear phagocyte system. A new classification of macrophages, monocytes, and their precursor cells. Bull. World Health Organ. 46, 845–852 (1972).
van Furth, R. & Diesselhoff-Den Dulk, M.M. The kinetics of promonocytes and monocytes in the bone marrow. J. Exp. Med. 132, 813–828 (1970).
Hume, D.A. The mononuclear phagocyte system. Curr. Opin. Immunol. 18, 49–53 (2006).
Hume, D.A. et al. The mononuclear phagocyte system revisited. J. Leukoc. Biol. 72, 621–627 (2002).
Gordon, S. The macrophage: past, present and future. Eur. J. Immunol. 37 (Suppl 1), S9–S17 (2007).
Gordon, S. & Taylor, P.R. Monocyte and macrophage heterogeneity. Nat. Rev. 5, 953–964 (2005).
Hume, D.A. & Gordon, S. Mononuclear phagocyte system of the mouse defined by immunohistochemical localization of antigen F4/80. Identification of resident macrophages in renal medullary and cortical interstitium and the juxtaglomerular complex. J. Exp. Med. 157, 1704–1709 (1983).
Hume, D.A., Halpin, D., Charlton, H. & Gordon, S. The mononuclear phagocyte system of the mouse defined by immunohistochemical localization of antigen F4/80: macrophages of endocrine organs. Proc. Natl. Acad. Sci. USA 81, 4174–4177 (1984).
Hume, D.A., Loutit, J.F. & Gordon, S. The mononuclear phagocyte system of the mouse defined by immunohistochemical localization of antigen F4/80: macrophages of bone and associated connective tissue. J. Cell Sci. 66, 189–194 (1984).
Hume, D.A., Perry, V.H. & Gordon, S. Immunohistochemical localization of a macrophage-specific antigen in developing mouse retina: phagocytosis of dying neurons and differentiation of microglial cells to form a regular array in the plexiform layers. J. Cell Biol. 97, 253–257 (1983).
Hume, D.A., Perry, V.H. & Gordon, S. The mononuclear phagocyte system of the mouse defined by immunohistochemical localisation of antigen F4/80: macrophages associated with epithelia. Anat. Rec. 210, 503–512 (1984).
Hume, D.A., Robinson, A.P., MacPherson, G.G. & Gordon, S. The mononuclear phagocyte system of the mouse defined by immunohistochemical localization of antigen F4/80. Relationship between macrophages, Langerhans cells, reticular cells, and dendritic cells in lymphoid and hematopoietic organs. J. Exp. Med. 158, 1522–1536 (1983).
Markham, N.P. & Florey, H.W. The effect on experimental tuberculosis of the intravenous injection of insoluble substances; experiments with carbon. Br. J. Exp. Pathol. 32, 25–33 (1951).
Sanders, A.G., Florey, H.W. & Wells, A.Q. The behaviour of intravenously injected particles of carbon and micrococcin in normal and tuberculous tissue. Br. J. Exp. Pathol. 32, 452–457 (1951).
Chieppa, M., Rescigno, M., Huang, A.Y. & Germain, R.N. Dynamic imaging of dendritic cell extension into the small bowel lumen in response to epithelial cell TLR engagement. J. Exp. Med. 203, 2841–2852 (2006).
Hapfelmeier, S. et al. Microbe sampling by mucosal dendritic cells is a discrete, MyD88-independent step in DeltainvG S. Typhimurium colitis. J. Exp. Med. 205, 437–450 (2008).
Martinez-Pomares, L. & Gordon, S. Antigen presentation the macrophage way. Cell 131, 641–643 (2007).
Martinez, F.O., Sica, A., Mantovani, A. & Locati, M. Macrophage activation and polarization. Front. Biosci. 13, 453–461 (2008).
Potten, C.S. Cell replacement in epidermis keratopoiesis via discrete units of proliferation. Int. Rev. Cytol. 69, 271–318 (1981).
Potten, C.S. & Allen, T.D. The fine structure and cell kinetics of mouse epidermis after wounding. J. Cell Sci. 17, 413–447 (1975).
Potten, C.S. & Allen, T.D. A model implicating the Langerhans cell in keratinocyte proliferation control. Differentiation 5, 43–47 (1976).
Chang, M.K. et al. Osteal tissue macrophages are intercalated throughout human and mouse bone lining tissues and regulate osteoblast function in vitro and in vivo. J. Immunol. 181, 1232–1244 (2008).
Rae, F. et al. Characterisation and trophic functions of murine embryonic macrophages based upon the use of a Csf1r-EGFP transgene reporter. Dev. Biol. 308, 232–246 (2007).
Taylor, P.R. et al. Macrophage receptors and immune recognition. Annu. Rev. Immunol. 23, 901–944 (2005).
Jung, S. et al. Analysis of fractalkine receptor CX3CR1 function by targeted deletion and green fluorescent protein reporter gene insertion. Mol. Cell. Biol. 20, 4106–4114 (2000).
Palframan, R.T. et al. Inflammatory chemokine transport and presentation in HEV: a remote control mechanism for monocyte recruitment to lymph nodes in inflamed tissues. J. Exp. Med. 194, 1361–1373 (2001).
Ovchinnikov, D.A. et al. Expression of Gal4-dependent transgenes in cells of the mononuclear phagocyte system labeled with enhanced cyan fluorescent protein using Csf1r-Gal4VP16/UAS-ECFP double-transgenic mice. J. Leukoc. Biol. 83, 430–433 (2008).
Sasmono, R.T. et al. Mouse neutrophilic granulocytes express mRNA encoding the macrophage colony-stimulating factor receptor CSF-1R as well as many other macrophage-specific transcripts and can transdifferentiate into macrophages in vitro in response to CSF-1. J. Leukoc. Biol. 82, 111–123 (2007).
Sasmono, R.T. et al. A macrophage colony-stimulating factor receptor-green fluorescent protein transgene is expressed throughout the mononuclear phagocyte system of the mouse. Blood 101, 1155–1163 (2003).
Lichanska, A.M. & Hume, D.A. Origins and functions of phagocytes in the embryo. Exp. Hematol. 28, 601–611 (2000).
Lichanska, A.M. et al. Differentiation of the mononuclear phagocyte system during mouse embryogenesis: the role of transcription factor PU.1. Blood 94, 127–138 (1999).
Lindquist, R.L. et al. Visualizing dendritic cell networks in vivo. Nat. Immunol. 5, 1243–1250 (2004).
Jiang, W. et al. The receptor DEC-205 expressed by dendritic cells and thymic epithelial cells is involved in antigen processing. Nature 375, 151–155 (1995).
Kato, M., Khan, S., d’Aniello, E., McDonald, K.J. & Hart, D.N. The novel endocytic and phagocytic C-Type lectin receptor DCL-1/CD302 on macrophages is colocalized with F-actin, suggesting a role in cell adhesion and migration. J. Immunol. 179, 6052–6063 (2007).
Ripoll, V.M., Irvine, K.M., Ravasi, T., Sweet, M.J. & Hume, D.A. Gpnmb is induced in macrophages by IFN-gamma and lipopolysaccharide and acts as a feedback regulator of proinflammatory responses. J. Immunol. 178, 6557–6566 (2007).
Hogg, N., Takacs, L., Palmer, D.G., Selvendran, Y. & Allen, C. The p150,95 molecule is a marker of human mononuclear phagocytes: comparison with expression of class II molecules. Eur. J. Immunol. 16, 240–248 (1986).
Myones, B.L., Dalzell, J.G., Hogg, N. & Ross, G.D. Neutrophil and monocyte cell surface p150,95 has iC3b-receptor CR4 activity resembling CR3. J. Clin. Invest. 82, 640–651 (1988).
Metlay, J.P. et al. The distinct leukocyte integrins of mouse spleen dendritic cells as identified with new hamster monoclonal antibodies. J. Exp. Med. 171, 1753–1771 (1990).
Franke-Ullmann, G. et al. Characterization of murine lung interstitial macrophages in comparison with alveolar macrophages in vitro. J. Immunol. 157, 3097–3104 (1996).
van Rijt, L.S. et al. In vivo depletion of lung CD11c+ dendritic cells during allergen challenge abrogates the characteristic features of asthma. J. Exp. Med. 201, 981–991 (2005).
Niess, J.H. & Reinecker, H.C. Lamina propria dendritic cells in the physiology and pathology of the gastrointestinal tract. Curr. Opin. Gastroenterol. 21, 687–691 (2005).
Niess, J.H. & Reinecker, H.C. Dendritic cells in the recognition of intestinal microbiota. Cell. Microbiol. 8, 558–564 (2006).
Uematsu, S. et al. Detection of pathogenic intestinal bacteria by Toll-like receptor 5 on intestinal CD11c+ lamina propria cells. Nat. Immunol. 7, 868–874 (2006).
Vallon-Eberhard, A., Landsman, L., Yogev, N., Verrier, B. & Jung, S. Transepithelial pathogen uptake into the small intestinal lamina propria. J. Immunol. 176, 2465–2469 (2006).
Pavli, P., Woodhams, C.E., Doe, W.F. & Hume, D.A. Isolation and characterization of antigen-presenting dendritic cells from the mouse intestinal lamina propria. Immunology 70, 40–47 (1990).
Soos, T.J. et al. CX3CR1+ interstitial dendritic cells form a contiguous network throughout the entire kidney. Kidney Int. 70, 591–596 (2006).
Templeton, S.P., Kim, T.S., O’Malley, K. & Perlman, S. Maturation and localization of macrophages and microglia during infection with a neurotropic murine coronavirus. Brain Pathol. 18, 40–51 (2008).
Ifergan, I. et al. The blood–brain barrier induces differentiation of migrating monocytes into Th17-polarizing dendritic cells. Brain 131, 785–799 (2008).
Bulloch, K. et al. CD11c/EYFP transgene illuminates a discrete network of dendritic cells within the embryonic, neonatal, adult, and injured mouse brain. J. Comp. Neurol. 508, 687–710 (2008).
Lloyd, C.M., Phillips, A.R., Cooper, G.J. & Dunbar, P.R. Three-colour fluorescence immunohistochemistry reveals the diversity of cells staining for macrophage markers in murine spleen and liver. J. Immunol. Methods 334, 70–81 (2008).
Chan, J., Leenen, P.J., Bertoncello, I., Nishikawa, S.I. & Hamilton, J.A. Macrophage lineage cells in inflammation: characterization by colony-stimulating factor-1 CSF-1 receptor c-Fms, ER-MP58, and ER-MP20 Ly-6C expression. Blood 92, 1423–1431 (1998).
Irvine, K.M. et al. A CSF-1 receptor kinase inhibitor targets effector functions and inhibits pro-inflammatory cytokine production from murine macrophage populations. FASEB J. 20, 1921–1923 (2006).
Sester, D.P. et al. Bacterial/CpG DNA down-modulates colony stimulating factor-1 receptor surface expression on murine bone marrow-derived macrophages with concomitant growth arrest and factor-independent survival. J. Immunol. 163, 6541–6550 (1999).
Leenen, P.J., de Bruijn, M.F., Voerman, J.S., Campbell, P.A. & van Ewijk, W. Markers of mouse macrophage development detected by monoclonal antibodies. J. Immunol. Methods 174, 5–19 (1994).
Kissenpfennig, A. et al. Disruption of the langerin/CD207 gene abolishes Birbeck granules without a marked loss of Langerhans cell function. Mol. Cell. Biol. 25, 88–99 (2005).
Valladeau, J. et al. Langerin, a novel C-type lectin specific to Langerhans cells, is an endocytic receptor that induces the formation of Birbeck granules. Immunity 12, 71–81 (2000).
Bennett, C.L. et al. Inducible ablation of mouse Langerhans cells diminishes but fails to abrogate contact hypersensitivity. J. Cell Biol. 169, 569–576 (2005).
Flores-Langarica, A. et al. Network of dendritic cells within the muscular layer of the mouse intestine. Proc. Natl. Acad. Sci. USA 102, 19039–19044 (2005).
Sung, S.S. et al. A major lung CD103 alphaE-beta7 integrin-positive epithelial dendritic cell population expressing Langerin and tight junction proteins. J. Immunol. 176, 2161–2172 (2006).
Bursch, L.S. et al. Identification of a novel population of Langerin+ dendritic cells. J. Exp. Med. 204, 3147–3156 (2007).
Ginhoux, F. et al. Blood-derived dermal langerin+ dendritic cells survey the skin in the steady state. J. Exp. Med. 204, 3133–3146 (2007).
Flacher, V. et al. Expression of langerin/CD207 reveals dendritic cell heterogeneity between inbred mouse strains. Immunology 123, 339–347 (2008).
Oetke, C., Kraal, G. & Crocker, P.R. The antigen recognized by MOMA-I is sialoadhesin. Immunol. Lett. 106, 96–98 (2006).
Breel, M., Van der Ende, M., Sminia, T. & Kraal, G. Subpopulations of lymphoid and non-lymphoid cells in bronchus-associated lymphoid tissue BALT of the mouse. Immunology 63, 657–662 (1988).
Keenihan, S.N. & Robertson, S.A. Diversity in phenotype and steroid hormone dependence in dendritic cells and macrophages in the mouse uterus. Biol. Reprod. 70, 1562–1572 (2004).
Dupasquier, M., Stoitzner, P., van Oudenaren, A., Romani, N. & Leenen, P.J. Macrophages and dendritic cells constitute a major subpopulation of cells in the mouse dermis. J. Invest. Dermatol. 123, 876–879 (2004).
Anjuere, F. et al. Definition of dendritic cell subpopulations present in the spleen, Peyer's patches, lymph nodes, and skin of the mouse. Blood 93, 590–598 (1999).
Bryniarski, K., Szczepanik, M., Maresz, K., Ptak, M. & Ptak, W. Subpopulations of mouse testicular macrophages and their immunoregulatory function. Am. J. Reprod. Immunol. 52, 27–35 (2004).
Cumberbatch, M. & Kimber, I. Phenotypic characteristics of antigen-bearing cells in the draining lymph nodes of contact sensitized mice. Immunology 71, 404–410 (1990).
Denning, T.L., Wang, Y.C., Patel, S.R., Williams, I.R. & Pulendran, B. Lamina propria macrophages and dendritic cells differentially induce regulatory and interleukin 17-producing T cell responses. Nat. Immunol. 8, 1086–1094 (2007).
Dudziak, D. et al. Differential antigen processing by dendritic cell subsets in vivo. Science (NY) 315, 107–111 (2007).
Duraiswamy, N., Tse, Y., Hammerberg, C., Kang, S. & Cooper, K.D. Distinction of class II MHC+ Langerhans cell-like interstitial dendritic antigen-presenting cells in murine dermis from dermal macrophages. J. Invest. Dermatol. 103, 678–683 (1994).
Geijtenbeek, T.B. et al. Marginal zone macrophages express a murine homologue of DC-SIGN that captures blood-borne antigens in vivo. Blood 100, 2908–2916 (2002).
Haines, K.A., Flotte, T.J., Springer, T.A., Gigli, I. & Thorbecke, G.J. Staining of Langerhans cells with monoclonal antibodies to macrophages and lymphoid cells. Proc. Natl. Acad. Sci. USA 80, 3448–3451 (1983).
Hamrah, P. & Dana, M.R. Corneal antigen-presenting cells. Chem. Immunol. Allergy 92, 58–70 (2007).
Heath, W.R. et al. Cross-presentation, dendritic cell subsets, and the generation of immunity to cellular antigens. Immunol. Rev. 199, 9–26 (2004).
Hume, D.A. Immunohistochemical analysis of murine mononuclear phagocytes that express class II major histocompatibility antigens. Immunobiology 170, 381–389 (1985).
Hutter, C. & Poulter, L.W. The balance of macrophage subsets may be customised at mucosal surfaces. FEMS Microbiol. Immunol. 5, 309–315 (1992).
Inaba, K. et al. Tissue distribution of the DEC-205 protein that is detected by the monoclonal antibody NLDC-145. I. Expression on dendritic cells and other subsets of mouse leukocytes. Cell. Immunol. 163, 148–156 (1995).
Inaba, K. et al. The tissue distribution of the B7-2 costimulator in mice: abundant expression on dendritic cells in situ and during maturation in vitro. J. Exp. Med. 180, 1849–1860 (1994).
Jakubzick, C. et al. Blood monocyte subsets differentially give rise to CD103+ and CD103− pulmonary dendritic cell populations. J. Immunol. 180, 3019–3027 (2008).
Julia, V. et al. A restricted subset of dendritic cells captures airborne antigens and remains able to activate specific T cells long after antigen exposure. Immunity 16, 271–283 (2002).
Jung, S. et al. In vivo depletion of CD11c+ dendritic cells abrogates priming of CD8+ T cells by exogenous cell-associated antigens. Immunity 17, 211–220 (2002).
Kelsall, B.L. & Strober, W. Distinct populations of dendritic cells are present in the subepithelial dome and T cell regions of the murine Peyer's patch. J. Exp. Med. 183, 237–247 (1996).
von Garnier, C. et al. Anatomical location determines the distribution and function of dendritic cells and other APCs in the respiratory tract. J. Immunol. 175, 1609–1618 (2005).
Witmer-Pack, M.D., Swiggard, W.J., Mirza, A., Inaba, K. & Steinman, R.M. Tissue distribution of the DEC-205 protein that is detected by the monoclonal antibody NLDC-145. II. Expression in situ in lymphoid and nonlymphoid tissues. Cell. Immunol. 163, 157–162 (1995).
Ravasi, T. et al. Generation of diversity in the innate immune system: macrophage heterogeneity arises from gene-autonomous transcriptional probability of individual inducible genes. J. Immunol. 168, 44–50 (2002).
Gimelbrant, A.A., Ensminger, A.W., Qi, P., Zucker, J. & Chess, A. Monoallelic expression and asynchronous replication of p120 catenin in mouse and human cells. J. Biol. Chem. 280, 1354–1359 (2005).
Fogg, D.K. et al. A clonogenic bone marrow progenitor specific for macrophages and dendritic cells. Science (NY) 311, 83–87 (2006).
Li, Q., Pan, P.Y., Gu, P., Xu, D. & Chen, S.H. Role of immature myeloid Gr-1+ cells in the development of antitumor immunity. Cancer Res. 64, 1130–1139 (2004).
Bonifer, C. & Hume, D.A. The transcriptional regulation of the colony-stimulating factor 1 receptor csf1r gene during hematopoiesis. Front Biosci. 13, 549–560 (2008).
Chitu, V. & Stanley, E.R. Colony-stimulating factor-1 in immunity and inflammation. Curr. Opin. Immunol. 18, 39–48 (2006).
Dai, X.M. et al. Targeted disruption of the mouse colony-stimulating factor 1 receptor gene results in osteopetrosis, mononuclear phagocyte deficiency, increased primitive progenitor cell frequencies, and reproductive defects. Blood 99, 111–120 (2002).
Lin, H. et al. Discovery of a cytokine and its receptor by functional screening of the extracellular proteome. Science (NY) 320, 807–811 (2008).
MacDonald, K.P. et al. The colony-stimulating factor 1 receptor is expressed on dendritic cells during differentiation and regulates their expansion. J. Immunol. 175, 1399–1405 (2005).
Hume, D.A., Pavli, P., Donahue, R.E. & Fidler, I.J. The effect of human recombinant macrophage colony-stimulating factor CSF-1 on the murine mononuclear phagocyte system in vivo. J. Immunol. 141, 3405–3409 (1988).
Misawa, E., Sakurai, T., Yamada, M., Tamura, Y. & Motoyoshi, K. Administration of macrophage colony-stimulating factor mobilized both CD11b+CD11c+ cells and NK1.1+ cells into peripheral blood. Int. Immunopharmacol. 4, 791–803 (2004).
Fancke, B., Suter, M., Hochrein, H. & O’Keeffe, M. M-CSF: a novel plasmacytoid and conventional dendritic cell poietin. Blood 111, 150–159 (2008).
Morrissey, P.J., Bressler, L., Park, L.S., Alpert, A. & Gillis, S. Granulocyte-macrophage colony-stimulating factor augments the primary antibody response by enhancing the function of antigen-presenting cells. J. Immunol. 139, 1113–1119 (1987).
Falk, L.A., Wahl, L.M. & Vogel, S.N. Analysis of Ia antigen expression in macrophages derived from bone marrow cells cultured in granulocyte-macrophage colony-stimulating factor or macrophage colony-stimulating factor. J. Immunol. 140, 2652–2660 (1988).
Fischer, H.G., Frosch, S., Reske, K. & Reske-Kunz, A.B. Granulocyte-macrophage colony-stimulating factor activates macrophages derived from bone marrow cultures to synthesis of MHC class II molecules and to augmented antigen presentation function. J. Immunol. 141, 3882–3888 (1988).
MacPherson, G.G. Properties of lymph-borne veiled dendritic cells in culture. I. Modulation of phenotype, survival and function: partial dependence on GM-CSF. Immunology 68, 102–107 (1989).
Inaba, K. et al. Generation of large numbers of dendritic cells from mouse bone marrow cultures supplemented with granulocyte/macrophage colony-stimulating factor. J. Exp. Med. 176, 1693–1702 (1992).
Inaba, K. et al. Granulocytes, macrophages, and dendritic cells arise from a common major histocompatibility complex class II-negative progenitor in mouse bone marrow. Proc. Natl. Acad. Sci. USA 90, 3038–3042 (1993).
Sallusto, F. & Lanzavecchia, A. Efficient presentation of soluble antigen by cultured human dendritic cells is maintained by granulocyte/macrophage colony-stimulating factor plus interleukin 4 and downregulated by tumor necrosis factor alpha. J. Exp. Med. 179, 1109–1118 (1994).
Kiertscher, S.M. & Roth, M.D. Human CD14+ leukocytes acquire the phenotype and function of antigen-presenting dendritic cells when cultured in GM-CSF and IL-4. J. Leukoc. Biol. 59, 208–218 (1996).
Hibbs, M.L. et al. Mice lacking three myeloid colony-stimulating factors G-CSF, GM-CSF, and M-CSF still produce macrophages and granulocytes and mount an inflammatory response in a sterile model of peritonitis. J. Immunol. 178, 6435–6443 (2007).
Lean, J.M., Fuller, K. & Chambers, T.J. FLT3 ligand can substitute for macrophage colony-stimulating factor in support of osteoclast differentiation and function. Blood 98, 2707–2713 (2001).
Niida, S. et al. Vascular endothelial growth factor can substitute for macrophage colony-stimulating factor in the support of osteoclastic bone resorption. J. Exp. Med. 190, 293–298 (1999).
Niida, S. et al. VEGF receptor 1 signaling is essential for osteoclast development and bone marrow formation in colony-stimulating factor 1-deficient mice. Proc. Natl. Acad. Sci. USA 102, 14016–14021 (2005).
Karsunky, H., Merad, M., Cozzio, A., Weissman, I.L. & Manz, M.G. Flt3 ligand regulates dendritic cell development from Flt3+ lymphoid and myeloid-committed progenitors to Flt3+ dendritic cells in vivo. J. Exp. Med. 198, 305–313 (2003).
Breen, F.N., Hume, D.A. & Weidemann, M.J. The effects of interleukin 3 IL-3 on cells responsive to macrophage colony-stimulating factor CSF-1 in liquid murine bone marrow culture. Br. J. Haematol. 74, 138–145 (1990).
Breen, F.N., Hume, D.A. & Weidemann, M.J. Interactions among granulocyte-macrophage colony-stimulating factor, macrophage colony-stimulating factor, and IFN-gamma lead to enhanced proliferation of murine macrophage progenitor cells. J. Immunol. 147, 1542–1547 (1991).
Sudo, T. et al. Functional hierarchy of c-kit and c-fms in intramarrow production of CFU-M. Oncogene 11, 2469–2476 (1995).
Tagoh, H. et al. Transcription factor complex formation and chromatin fine structure alterations at the murine c-fms CSF-1 receptor locus during maturation of myeloid precursor cells. Genes Dev. 16, 1721–1737 (2002).
Himes, S.R. et al. A highly conserved c-fms gene intronic element controls macrophage-specific and regulated expression. J. Leukoc. Biol. 70, 812–820 (2001).
Ross, I.L., Yue, X., Ostrowski, M.C. & Hume, D.A. Interaction between PU.1 and another Ets family transcription factor promotes macrophage-specific Basal transcription initiation. J. Biol. Chem. 273, 6662–6669 (1998).
Krysinska, H. et al. A two-step, PU.1-dependent mechanism for developmentally regulated chromatin remodeling and transcription of the c-fms gene. Mol. Cell. Biol. 27, 878–887 (2007).
Luchin, A. et al. Genetic and physical interactions between microphthalmia transcription factor and PU.1 are necessary for osteoclast gene expression and differentiation. J. Biol. Chem. 276, 36703–36710 (2001).
Hoogenkamp, M. et al. The Pu.1 locus is differentially regulated at the level of chromatin structure and noncoding transcription by alternate mechanisms at distinct developmental stages of hematopoiesis. Mol. Cell. Biol. 27, 7425–7438 (2007).
Rosenbauer, F. & Tenen, D.G. Transcription factors in myeloid development: balancing differentiation with transformation. Nat. Rev. 7, 105–117 (2007).
Rehli, M., Lichanska, A., Cassady, A.I., Ostrowski, M.C. & Hume, D.A. TFEC is a macrophage-restricted member of the microphthalmia-TFE subfamily of basic helix–loop–helix leucine zipper transcription factors. J. Immunol. 162, 1559–1565 (1999).
Rehli, M. et al. Transcription factor Tfec contributes to the IL-4-inducible expression of a small group of genes in mouse macrophages including the granulocyte colony-stimulating factor receptor. J. Immunol. 174, 7111–7122 (2005).
Meadows, N.A. et al. The expression of Clcn7 and Ostm1 in osteoclasts is coregulated by microphthalmia transcription factor. J. Biol. Chem. 282, 1891–1904 (2007).
Aziz, A. et al. Development of macrophages with altered actin organization in the absence of MafB. Mol. Cell. Biol. 26, 6808–6818 (2006).
Bakri, Y. et al. Balance of MafB and PU.1 specifies alternative macrophage or dendritic cell fate. Blood 105, 2707–2716 (2005).
Moriguchi, T. et al. MafB is essential for renal development and F4/80 expression in macrophages. Mol. Cell. Biol. 26, 5715–5727 (2006).
Wells, C.A. et al. Continued discovery of transcriptional units expressed in cells of the mouse mononuclear phagocyte lineage. Genome Res. 13, 1360–1365 (2003).
Nilsson, R. et al. Transcriptional network dynamics in macrophage activation. Genomics 88, 133–142 (2006).
Nikolic, T., de Bruijn, M.F., Lutz, M.B. & Leenen, P.J. Developmental stages of myeloid dendritic cells in mouse bone marrow. Int. Immunol. 15, 515–524 (2003).
Witsell, A.L. & Schook, L.B. Macrophage heterogeneity occurs through a developmental mechanism. Proc. Natl. Acad. Sci. USA 88, 1963–1967 (1991).
Varol, C. et al. Monocytes give rise to mucosal, but not splenic, conventional dendritic cells. J. Exp. Med. 204, 171–180 (2007).
Geissmann, F. The origin of dendritic cells. Nat. Immunol. 8, 558–560 (2007).
Geissmann, F. et al. Blood monocytes: distinct subsets, how they relate to dendritic cells, and their possible roles in the regulation of T-cell responses. Immunol. Cell Biol. 86, 398–408 (2008).
Serbina, N.V., Salazar-Mather, T.P., Biron, C.A., Kuziel, W.A. & Pamer, E.G. TNF/iNOS-producing dendritic cells mediate innate immune defense against bacterial infection. Immunity 19, 59–70 (2003).
Ginhoux, F. et al. Langerhans cells arise from monocytes in vivo. Nat. Immunol. 7, 265–273 (2006).
Onai, N. et al. Identification of clonogenic common Flt3+M-CSFR+ plasmacytoid and conventional dendritic cell progenitors in mouse bone marrow. Nat. Immunol. 8, 1207–1216 (2007).
Liu, K. et al. Origin of dendritic cells in peripheral lymphoid organs of mice. Nat. Immunol. 8, 578–583 (2007).
Waskow, C. et al. The receptor tyrosine kinase Flt3 is required for dendritic cell development in peripheral lymphoid tissues. Nat. Immunol. 9, 676–683 (2008).
van Furth, R. & Diesselhoff-den Dulk, M.M. Dual origin of mouse spleen macrophages. J. Exp. Med. 160, 1273–1283 (1984).
Kennedy, D.W. & Abkowitz, J.L. Kinetics of central nervous system microglial and macrophage engraftment: analysis using a transgenic bone marrow transplantation model. Blood 90, 986–993 (1997).
Kennedy, D.W. & Abkowitz, J.L. Mature monocytic cells enter tissues and engraft. Proc. Natl. Acad. Sci. USA 95, 14944–14949 (1998).
van oud Alblas, A.B. & van Furth, R Origin, kinetics, and characteristics of pulmonary macrophages in the normal steady state. J. Exp. Med. 149, 1504–1518 (1979).
Randolph, G.J., Beaulieu, S., Lebecque, S., Steinman, R.M. & Muller, W.A. Differentiation of monocytes into dendritic cells in a model of transendothelial trafficking. Science (NY) 282, 480–483 (1998).
Geissmann, F., Jung, S. & Littman, D.R. Blood monocytes consist of two principal subsets with distinct migratory properties. Immunity 19, 71–82 (2003).
Sunderkotter, C. et al. Subpopulations of mouse blood monocytes differ in maturation stage and inflammatory response. J. Immunol. 172, 4410–4417 (2004).
Auffray, C. et al. Monitoring of blood vessels and tissues by a population of monocytes with patrolling behavior. Science (NY) 317, 666–670 (2007).
Burke, B. et al. Increased TNF expression in CD43. Murine blood monocytes. Immunol. Lett. 118, 142–147 (2008).
Yrlid, U., Jenkins, C.D. & MacPherson, G.G. Relationships between distinct blood monocyte subsets and migrating intestinal lymph dendritic cells in vivo under steady-state conditions. J. Immunol. 176, 4155–4162 (2006).
Ledger, W.J. & Witkin, S.S. Are you a lumper or a splitter? Am. J. Obstet. Gynecol. 195, 1205–1209 (2006).
Author information
Authors and Affiliations
Corresponding author
PowerPoint slides
Rights and permissions
About this article
Cite this article
Hume, D. Differentiation and heterogeneity in the mononuclear phagocyte system. Mucosal Immunol 1, 432–441 (2008). https://doi.org/10.1038/mi.2008.36
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mi.2008.36
This article is cited by
-
CX3CR1 deficiency leads to impairment of immune surveillance in the epididymis
Cellular and Molecular Life Sciences (2023)
-
Immunogenicity Challenges Associated with Subcutaneous Delivery of Therapeutic Proteins
BioDrugs (2021)
-
Human microbiota-transplanted C57BL/6 mice and offspring display reduced establishment of key bacteria and reduced immune stimulation compared to mouse microbiota-transplantation
Scientific Reports (2020)
-
Epigenetic regulation of macrophages: from homeostasis maintenance to host defense
Cellular & Molecular Immunology (2020)
-
Quantitative biokinetics over a 28 day period of freshly generated, pristine, 20 nm titanium dioxide nanoparticle aerosols in healthy adult rats after a single two-hour inhalation exposure
Particle and Fibre Toxicology (2019)