Abstract
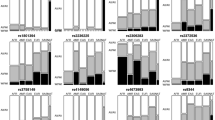
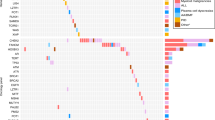
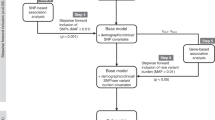
The causes of individual relapses in children with acute lymphoblastic leukemia (ALL) remain incompletely understood. We evaluated the contribution of germline genetic factors to relapse in 2225 children treated on Children’s Oncology Group trial AALL0232. We identified 302 germline single-nucleotide polymorphisms (SNPs) associated with relapse after adjusting for treatment and ancestry and 715 additional SNPs associated with relapse in an ancestry-specific manner. We tested for replication of these relapse-associated SNPs in external data sets of antileukemic drug pharmacokinetics and pharmacodynamics and an independent clinical cohort. 224 SNPs were associated with rapid drug clearance or drug resistance, and 32 were replicated in the independent cohort. The adverse risk associated with black and Hispanic ancestries was attenuated by addition of the 4 SNPs most strongly associated with relapse in these populations (for blacks: model without SNPs hazard ratio (HR)=2.32, P=2.27 × 10−4, model with SNPs HR=1.07, P=0.79; for Hispanics: model without SNPs HR=1.7, P=8.23 × 10−5, model with SNPs HR=1.31, P=0.065). Relapse SNPs associated with asparaginase resistance or allergy were overrepresented among SNPs associated with relapse in the more asparaginase intensive treatment arm (20/54 in Capizzi-methorexate arm vs 8/54 in high-dose methotrexate arm, P=0.015). Inherited genetic variation contributes to race-specific and treatment-specific relapse risk.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hunger SP, Lu X, Devidas M, Camitta BM, Gaynon PS, Winick NJ et al. Improved survival for children and adolescents with acute lymphoblastic leukemia between 1990 and 2005: a report from the children's oncology group. J Clin Oncol 2012; 30: 1663–1669.
Schrappe M, Moricke A, Reiter A, Henze G, Welte K, Gadner H et al. Key treatment questions in childhood acute lymphoblastic leukemia: results in 5 consecutive trials performed by the ALL-BFM study group from 1981 to 2000. Klin Padiatr 2013; 225 (Suppl 1): S62–S72.
Pui CH, Pei D, Sandlund JT, Ribeiro RC, Rubnitz JE, Raimondi SC et al. Long-term results of St Jude Total Therapy Studies 11, 12, 13A, 13B, and 14 for childhood acute lymphoblastic leukemia. Leukemia 2010; 24: 371–382.
Place AE, Stevenson KE, Vrooman LM, Harris MH, Hunt SK, O'Brien JE et al. Intravenous pegylated asparaginase versus intramuscular native Escherichia coli L-asparaginase in newly diagnosed childhood acute lymphoblastic leukaemia (DFCI 05-001): a randomised, open-label phase 3 trial. Lancet Oncol 2015; 16: 1677–1690.
Pui CH, Campana D, Pei D, Bowman WP, Sandlund JT, Kaste SC et al. Treating childhood acute lymphoblastic leukemia without cranial irradiation. N Engl J Med 2009; 360: 2730–2741.
Conter V, Valsecchi MG, Parasole R, Putti MC, Locatelli F, Barisone E et al. Childhood high-risk acute lymphoblastic leukemia in first remission: results after chemotherapy or transplant from the AIEOP ALL 2000 study. Blood 2014; 123: 1470–1478.
Stary J, Zimmermann M, Campbell M, Castillo L, Dibar E, Donska S et al. Intensive chemotherapy for childhood acute lymphoblastic leukemia: results of the randomized intercontinental trial ALL IC-BFM 2002. J Clin Oncol 2014; 32: 174–184.
Larsen EC, Devidas M, Chen S, Salzer WL, Raetz EA, Loh ML et al. Dexamethasone and high-dose methotrexate improve outcome for children and young adults with high-risk b-acute lymphoblastic leukemia: A Report From Children's Oncology Group Study AALL0232. J Clin Oncol 2016; 34: 2380–2388.
Bowman WP, Larsen EL, Devidas M, Linda SB, Blach L, Carroll AJ et al. Augmented therapy improves outcome for pediatric high risk acute lymphocytic leukemia: results of Children's Oncology Group trial P9906. Pediatr Blood Cancer 2011; 57: 569–577.
Marshall GM, Dalla Pozza L, Sutton R, Ng A, de Groot-Kruseman HA, van der Velden VH et al. High-risk childhood acute lymphoblastic leukemia in first remission treated with novel intensive chemotherapy and allogeneic transplantation. Leukemia 2013; 27: 1497–1503.
Diouf B, Crews KR, Lew G, Pei D, Cheng C, Bao J et al. Association of an inherited genetic variant with vincristine-related peripheral neuropathy in children with acute lymphoblastic leukemia. JAMA 2015; 313: 815–823.
Gregers J, Green H, Christensen IJ, Dalhoff K, Schroeder H, Carlsen N et al. Polymorphisms in the ABCB1 gene and effect on outcome and toxicity in childhood acute lymphoblastic leukemia. Pharmacogenomics J 2015; 15: 372–379.
Moriyama T, Nishii R, Perez-Andreu V, Yang W, Klussmann FA, Zhao X et al. NUDT15 polymorphisms alter thiopurine metabolism and hematopoietic toxicity. Nat Genet 2016; 48: 367–373.
Ramsey LB, Janke LJ, Edick MJ, Cheng C, Williams RT, Sherr CJ et al. Host thiopurine methyltransferase status affects mercaptopurine antileukemic effectiveness in a murine model. Pharmacogenet Genomics 2014; 24: 263–271.
Kawedia JD, Liu C, Pei D, Cheng C, Fernandez CA, Howard SC et al. Dexamethasone exposure and asparaginase antibodies affect relapse risk in acute lymphoblastic leukemia. Blood 2012; 119: 1658–1664.
Fernandez CA, Smith C, Yang W, Mullighan CG, Qu C, Larsen E et al. Genome-wide analysis links NFATC2 with asparaginase hypersensitivity. Blood 2015; 126: 69–75.
Abrahao R, Lichtensztajn DY, Ribeiro RC, Marina NM, Keogh RH, Marcos-Gragera R et al. Racial/ethnic and socioeconomic disparities in survival among children with acute lymphoblastic leukemia in California, 1988-2011: A population-based observational study. Pediatr Blood Cancer 2015; 62: 1819–1825.
Goggins WB, Lo FF . Racial and ethnic disparities in survival of US children with acute lymphoblastic leukemia: evidence from the SEER database 1988-2008. Cancer Causes Control 2012; 23: 737–743.
Kadan-Lottick NS, Ness KK, Bhatia S, Gurney JG . Survival variability by race and ethnicity in childhood acute lymphoblastic leukemia. JAMA 2003; 290: 2008–2014.
Bhatia S, Sather HN, Heerema NA, Trigg ME, Gaynon PS, Robison LL . Racial and ethnic differences in survival of children with acute lymphoblastic leukemia. Blood 2002; 100: 1957–1964.
Kahn JM, Keegan TH, Tao L, Abrahao R, Bleyer A, Viny AD . Racial disparities in the survival of American children, adolescents, and young adults with acute lymphoblastic leukemia, acute myelogenous leukemia, and Hodgkin lymphoma. Cancer 2016; 122: 2723–2730.
Bhatia S, Landier W, Hageman L, Kim H, Chen Y, Crews KR et al. 6MP adherence in a multiracial cohort of children with acute lymphoblastic leukemia: a Children's Oncology Group study. Blood 2014; 124: 2345–2353.
Pui CH, Sandlund JT, Pei D, Rivera GK, Howard SC, Ribeiro RC et al. Results of therapy for acute lymphoblastic leukemia in black and white children. JAMA 2003; 290: 2001–2007.
Bhatia S, Landier W, Hageman L, Chen Y, Kim H, Sun CL et al. Systemic Exposure to Thiopurines and Risk of Relapse in Children With Acute Lymphoblastic Leukemia: A Children's Oncology Group Study. JAMA oncology 2015; 1: 287–295.
Pollock BH, DeBaun MR, Camitta BM, Shuster JJ, Ravindranath Y, Pullen DJ et al. Racial differences in the survival of childhood B-precursor acute lymphoblastic leukemia: a Pediatric Oncology Group Study. J Clin Oncol 2000; 18: 813–823.
Perez-Andreu V, Roberts KG, Xu H, Smith C, Zhang H, Yang W et al. A genome-wide association study of susceptibility to acute lymphoblastic leukemia in adolescents and young adults. Blood 2015; 125: 680–686.
Xu H, Yang W, Perez-Andreu V, Devidas M, Fan Y, Cheng C et al. Novel susceptibility variants at 10p12.31-12.2 for childhood acute lymphoblastic leukemia in ethnically diverse populations. J Natl Cancer Inst 2013; 105: 733–742.
Yang JJ, Cheng C, Devidas M, Cao X, Fan Y, Campana D et al. Ancestry and pharmacogenomics of relapse in acute lymphoblastic leukemia. Nat Genet 2011; 43: 237–241.
Pritchard JK, Stephens M, Donnelly P . Inference of population structure using multilocus genotype data. Genetics 2000; 155: 945–959.
Mao X, Bigham AW, Mei R, Gutierrez G, Weiss KM, Brutsaert TD et al. A genomewide admixture mapping panel for Hispanic/Latino populations. Am J Hum Genet 2007; 80: 1171–1178.
Cheng C, Pounds SB, Boyett JM, Pei D, Kuo ML, Roussel MF . Statistical significance threshold criteria for analysis of microarray gene expression data. Stat Appl Genet Mol Biol 2004; 3: Article36.
Borowitz MJ, Devidas M, Hunger SP, Bowman WP, Carroll AJ, Carroll WL et al. Clinical significance of minimal residual disease in childhood acute lymphoblastic leukemia and its relationship to other prognostic factors: A Children's Oncology Group study. Blood 2008; 111: 5477–5485.
Borowitz MJ, Wood BL, Devidas M, Loh ML, Raetz EA, Salzer WL et al. Prognostic significance of minimal residual disease in high risk B-ALL: a report from Children's Oncology Group study AALL0232. Blood 2015; 126: 964–971.
Pui CH, Pei D, Coustan-Smith E, Jeha S, Cheng C, Bowman WP et al. Clinical utility of sequential minimal residual disease measurements in the context of risk-based therapy in childhood acute lymphoblastic leukaemia: a prospective study. Lancet Oncol 2015; 16: 465–474.
Rocconi RP, Lankes HA, Brady WE, Goodfellow PJ, Ramirez NC, Alvarez RD et al. The role of racial genetic admixture with endometrial cancer outcomes: An NRG Oncology/Gynecologic Oncology Group study. Gynecol Oncol 2016; 140: 264–269.
Hernandez-Suarez G, Sanabria MC, Serrano M, Herran OF, Perez J, Plata JL et al. Genetic ancestry is associated with colorectal adenomas and adenocarcinomas in Latino populations. Eur J Hum Genet 2014; 22: 1208–1216.
Torgerson DG, Capurso D, Ampleford EJ, Li X, Moore WC, Gignoux CR et al. Genome-wide ancestry association testing identifies a common European variant on 6q14.1 as a risk factor for asthma in African American subjects. J Allergy Clin Immunol 2012; 130: 622–9 e9.
Corvol H, De Giacomo A, Eng C, Seibold M, Ziv E, Chapela R et al. Genetic ancestry modifies pharmacogenetic gene-gene interaction for asthma. Pharmacogenet Genomics 2009; 19: 489–496.
Alarcon-Riquelme ME, Ziegler JT, Molineros J, Howard TD, Moreno-Estrada A, Sanchez-Rodriguez E et al. Genome-Wide Association Study in an Amerindian ancestry population reveals novel systemic lupus erythematosus risk loci and the role of European admixture. Arthritis Rheumatol 2016; 68: 932–943.
Molineros JE, Maiti AK, Sun C, Looger LL, Han S, Kim-Howard X et al. Admixture mapping in lupus identifies multiple functional variants within IFIH1 associated with apoptosis, inflammation, and autoantibody production. PLoS Genet 2013; 9: e1003222.
Niewold TB, Kelly JA, Kariuki SN, Franek BS, Kumar AA, Kaufman KM et al. IRF5 haplotypes demonstrate diverse serological associations which predict serum interferon alpha activity and explain the majority of the genetic association with systemic lupus erythematosus. Ann Rheum Dis 2012; 71: 463–468.
Kopp JB, Smith MW, Nelson GW, Johnson RC, Freedman BI, Bowden DW et al. MYH9 is a major-effect risk gene for focal segmental glomerulosclerosis. Nat Genet 2008; 40: 1175–1184.
Yang JJ, Cheng C, Devidas M, Cao X, Campana D, Yang W et al. Genome-wide association study identifies germline polymorphisms associated with relapse of childhood acute lymphoblastic leukemia. Blood 2012; 120: 4197–4204.
Yang JJ, Cheng C, Yang W, Pei D, Cao X, Fan Y et al. Genome-wide interrogation of germline genetic variation associated with treatment response in childhood acute lymphoblastic leukemia. JAMA 2009; 301: 393–403.
Evans WE, Crom WR, Stewart CF, Bowman WP, Chen CH, Abromowitch M et al. Methotrexate systemic clearance influences probability of relapse in children with standard-risk acute lymphocytic leukaemia. Lancet 1984; 1: 359–362.
Mikkelsen TS, Sparreboom A, Cheng C, Zhou Y, Boyett JM, Raimondi SC et al. Shortening infusion time for high-dose methotrexate alters antileukemic effects: a randomized prospective clinical trial. J Clin Oncol 2011; 29: 1771–1778.
Han X, Zheng T, Foss FM, Lan Q, Holford TR, Rothman N et al. Genetic polymorphisms in the metabolic pathway and non-Hodgkin lymphoma survival. Am J Hematol 2010; 85: 51–56.
Haque AK, Au W, Cajas-Salazar N, Khan S, Ginzel AW, Jones DV et al. CYP2E1 polymorphism, cigarette smoking, p53 expression, and survival in non-small cell lung cancer: a long term follow-up study. Appl Immunohistochem Mol Morphol 2004; 12: 315–322.
Roadmap Epigenomics C, Kundaje A, Meuleman W, Ernst J, Bilenky M, Yen A et al. Integrative analysis of 111 reference human epigenomes. Nature 2015; 518: 317–330.
Lin CW, Chang YL, Chang YC, Lin JC, Chen CC, Pan SH et al. MicroRNA-135b promotes lung cancer metastasis by regulating multiple targets in the Hippo pathway and LZTS1. Nat Commun 2013; 4: 1877.
Onken MD, Worley LA, Harbour JW . A metastasis modifier locus on human chromosome 8p in uveal melanoma identified by integrative genomic analysis. Clin Cancer Res 2008; 14: 3737–3745.
Wang XX, Zhu ZM, Su D, Lei T, Wu X, Fan Y et al. Down-regulation of leucine zipper putative tumor suppressor 1 is associated with poor prognosis, increased cell motility and invasion, and epithelial-to-mesenchymal transition characteristics in human breast carcinoma. Hum Pathol 2011; 42: 1410–1419.
Acknowledgements
The work was supported by the National Institutes of Health (grant numbers GM 92666, GM 115279, CA142665, CA 21765, CA 176063, CA 36401, CA 156449, CA98543 and CA 180886 (COG Chair's grant), CA98413 and CA180899 (COG Statistical Center), CA114766 (COG Specimen Banking), U01-HG04603, RC2- GM092618, R01-LM010685, 5T32-GM007569); Leukemia Lymphoma Society (grant number 6168); and by the American Lebanese Syrian Associated Charities. The funders had no role in the design and conduct of the study; collection, management, analysis, and interpretation of the data; preparation, review, or approval of the manuscript; and decision to submit the manuscript for publication.
Author contributions
MVR and JJY contributed to the conception and design of the study. EL, LBR, CAF, JRM, SWP, RJA, ELL, BD, SJ, C-HP, EAR, NJW, WLC, SPH, MLL, MD, WEE, JJY and MVR contributed to the provision of study materials, patient recruitment, or acquisition of data. SEK, CC, XC and MVR contributed to data analysis and interpretation. All authors contributed to the drafting and reviewing of the manuscript and gave their final approval to submit for publication.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Leukemia website
Supplementary information
Rights and permissions
About this article
Cite this article
Karol, S., Larsen, E., Cheng, C. et al. Genetics of ancestry-specific risk for relapse in acute lymphoblastic leukemia. Leukemia 31, 1325–1332 (2017). https://doi.org/10.1038/leu.2017.24
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2017.24
This article is cited by
-
Advances in germline predisposition to acute leukaemias and myeloid neoplasms
Nature Reviews Cancer (2021)
-
Identification of four novel associations for B-cell acute lymphoblastic leukaemia risk
Nature Communications (2019)
-
Early vs. late MRD response- and risk-based treatment intensification of childhood acute lymphoblastic leukemia: a prospective pilot study from Saudi Arabia
Experimental Hematology & Oncology (2018)